

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133139.3 - phase: 0
(185 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
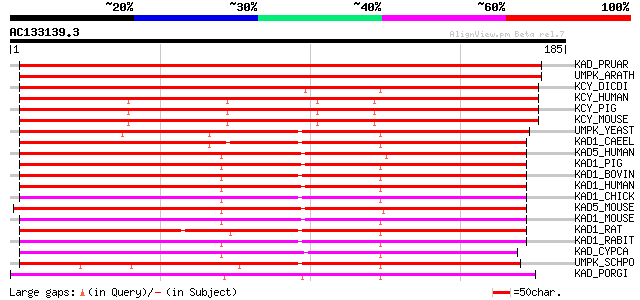
Score E
Sequences producing significant alignments: (bits) Value
KAD_PRUAR (O24464) Adenylate kinase (EC 2.7.4.3) (ATP-AMP transp... 269 2e-72
UMPK_ARATH (O04905) Uridylate kinase (EC 2.7.4.-) (UK) (Uridine ... 216 2e-56
KCY_DICDI (P20425) Cytidylate kinase (EC 2.7.4.14) (Deoxycytidyl... 157 1e-38
KCY_HUMAN (P30085) UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kina... 150 1e-36
KCY_PIG (Q29561) UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kinase... 149 3e-36
KCY_MOUSE (Q9DBP5) UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kina... 149 4e-36
UMPK_YEAST (P15700) Uridylate kinase (EC 2.7.4.-) (UK) (Uridine ... 145 7e-35
KAD1_CAEEL (Q20140) Probable adenylate kinase isoenzyme F38B2.4 ... 141 7e-34
KAD5_HUMAN (Q9Y6K8) Adenylate kinase isoenzyme 5 (EC 2.7.4.3) (A... 141 1e-33
KAD1_PIG (P00571) Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (ATP... 140 1e-33
KAD1_BOVIN (P00570) Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (A... 137 1e-32
KAD1_HUMAN (P00568) Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (A... 137 1e-32
KAD1_CHICK (P05081) Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (A... 136 3e-32
KAD5_MOUSE (Q920P5) Adenylate kinase isoenzyme 5 (EC 2.7.4.3) (A... 135 5e-32
KAD1_MOUSE (Q9R0Y5) Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (A... 135 7e-32
KAD1_RAT (P39069) Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (ATP... 134 9e-32
KAD1_RABIT (P00569) Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (A... 134 2e-31
KAD_CYPCA (P12115) Adenylate kinase (EC 2.7.4.3) (ATP-AMP transp... 129 5e-30
UMPK_SCHPO (O59771) Probable uridylate kinase (EC 2.7.4.-) (UK) ... 127 1e-29
KAD_PORGI (Q7MW54) Adenylate kinase (EC 2.7.4.3) (ATP-AMP transp... 120 1e-27
>KAD_PRUAR (O24464) Adenylate kinase (EC 2.7.4.3) (ATP-AMP
transphosphorylase)
Length = 231
Score = 269 bits (688), Expect = 2e-72
Identities = 128/174 (73%), Positives = 152/174 (86%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCA+IVE FGF H+SAGDLLR+ + S S YG++IL TIREG+IVPS VTV
Sbjct: 54 GGPGSGKGTQCAKIVEAFGFTHVSAGDLLRREIASGSAYGSVILSTIREGKIVPSQVTVE 113
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
LI +EM+ DN KFLIDGFPRSEENR AFE G EPD VL+FDCPE+EMVKRVL+RNQG
Sbjct: 114 LIQKEMESSDNYKFLIDGFPRSEENRKAFEQTIGAEPDVVLFFDCPEQEMVKRVLNRNQG 173
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVFA 177
R+DDNIDTIKKRL++F+ LN PVI++Y++RG+LH+INAVGT DEIFE+VRP+FA
Sbjct: 174 RVDDNIDTIKKRLEIFDELNWPVINYYSQRGKLHKINAVGTVDEIFEKVRPIFA 227
>UMPK_ARATH (O04905) Uridylate kinase (EC 2.7.4.-) (UK) (Uridine
monophosphate kinase) (UMP kinase) (UMP/CMP kinase)
Length = 202
Score = 216 bits (551), Expect = 2e-56
Identities = 105/174 (60%), Positives = 133/174 (76%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCA IVE +G+ HLSAGDLLR + S SE G MI I+EG+IVPS VT++
Sbjct: 21 GGPGSGKGTQCAYIVEHYGYTHLSAGDLLRAEIKSGSENGTMIQNMIKEGKIVPSEVTIK 80
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
L+ + +Q N KFLIDGFPR+EENR AFE +T EP FVL+FDCPEEEM KR+L RNQG
Sbjct: 81 LLQKAIQENGNDKFLIDGFPRNEENRAAFEKVTEIEPKFVLFFDCPEEEMEKRLLGRNQG 140
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVFA 177
R DDNI+TI+KR KVF +LPVI +Y +G++ +INA + +FE+V+ +F+
Sbjct: 141 REDDNIETIRKRFKVFLESSLPVIHYYEAKGKVRKINAAKPIEAVFEEVKAIFS 194
>KCY_DICDI (P20425) Cytidylate kinase (EC 2.7.4.14) (Deoxycytidylate
kinase) (UMP-CMP kinase)
Length = 194
Score = 157 bits (398), Expect = 1e-38
Identities = 85/177 (48%), Positives = 111/177 (62%), Gaps = 4/177 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCA IV FG+ HLSAGDLLR+ S S+ G MI I+ G IVPS VTV+
Sbjct: 13 GGPGSGKGTQCANIVRDFGWVHLSAGDLLRQEQQSGSKDGEMIATMIKNGEIVPSIVTVK 72
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITG--TEPDFVLYFDCPEEEMVKRVLSRN 121
L+ + + FL+DGFPR+EEN ++E + FVL+FDCPEE M +R+L R
Sbjct: 73 LLKNAIDANQGKNFLVDGFPRNEENNNSWEENMKDFVDTKFVLFFDCPEEVMTQRLLKRG 132
Query: 122 Q--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVF 176
+ GR DDNI++IKKR F VIDHY + ++ I A +E++ V +F
Sbjct: 133 ESSGRSDDNIESIKKRFNTFNVQTKLVIDHYNKFDKVKIIPANRDVNEVYNDVENLF 189
>KCY_HUMAN (P30085) UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kinase)
(Deoxycytidylate kinase) (Cytidine monophosphate kinase)
Length = 196
Score = 150 bits (380), Expect = 1e-36
Identities = 80/183 (43%), Positives = 119/183 (64%), Gaps = 10/183 (5%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVS-DSEYGAMILETIREGRIVPSAVTV 62
GGPG+GKGTQCARIVE +G+ HLSAG+LLR + DS+YG +I + I+EG+IVP +T+
Sbjct: 10 GGPGAGKGTQCARIVEKYGYTHLSAGELLRDERKNPDSQYGELIEKYIKEGKIVPVEITI 69
Query: 63 RLILREMQY-----GDNRKFLIDGFPRSEENRIAFEHITGTEPD--FVLYFDCPEEEMVK 115
L+ REM KFLIDGFPR+++N + + D FVL+FDC E ++
Sbjct: 70 SLLKREMDQTMAANAQKNKFLIDGFPRNQDNLQGWNKTMDGKADVSFVLFFDCNNEICIE 129
Query: 116 RVLSR--NQGRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVR 173
R L R + GR DDN ++++KR++ + P+ID Y G++ +I+A + DE+F++V
Sbjct: 130 RCLERGKSSGRSDDNRESLEKRIQTYLQSTKPIIDLYEEMGKVKKIDASKSVDEVFDEVV 189
Query: 174 PVF 176
+F
Sbjct: 190 QIF 192
>KCY_PIG (Q29561) UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kinase)
(Deoxycytidylate kinase)
Length = 196
Score = 149 bits (377), Expect = 3e-36
Identities = 79/183 (43%), Positives = 119/183 (64%), Gaps = 10/183 (5%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVS-DSEYGAMILETIREGRIVPSAVTV 62
GGPG+GKGTQCARIVE +G+ HLSAG+LLR + DS+YG +I + I++G+IVP +T+
Sbjct: 10 GGPGAGKGTQCARIVEKYGYTHLSAGELLRDERKNPDSQYGELIEKYIKDGKIVPVEITI 69
Query: 63 RLILREMQY-----GDNRKFLIDGFPRSEENRIAFEHITGTEPD--FVLYFDCPEEEMVK 115
L+ REM KFLIDGFPR+++N + + D FVL+FDC E ++
Sbjct: 70 SLLRREMDQTMAANAQKNKFLIDGFPRNQDNLQGWNKTMDGKADVSFVLFFDCNNEICIE 129
Query: 116 RVLSR--NQGRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVR 173
R L R + GR DDN ++++KR++ + P+ID Y G++ +I+A + DE+F++V
Sbjct: 130 RCLERGKSSGRSDDNRESLEKRIQTYLQSTKPIIDLYEEMGKVKKIDASKSVDEVFDEVV 189
Query: 174 PVF 176
+F
Sbjct: 190 KIF 192
>KCY_MOUSE (Q9DBP5) UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kinase)
(Deoxycytidylate kinase) (Cytidine monophosphate kinase)
Length = 196
Score = 149 bits (376), Expect = 4e-36
Identities = 80/183 (43%), Positives = 118/183 (63%), Gaps = 10/183 (5%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVS-DSEYGAMILETIREGRIVPSAVTV 62
GGPG+GKGTQCARIVE +G+ HLSAG+LLR + DS+YG +I + I+EG+IVP +T+
Sbjct: 10 GGPGAGKGTQCARIVEKYGYTHLSAGELLRDERKNPDSQYGELIEKYIKEGKIVPVEITI 69
Query: 63 RLILREMQY-----GDNRKFLIDGFPRSEENRIAFEHITGTEPD--FVLYFDCPEEEMVK 115
L+ REM KFLIDGFPR+++N + + D FVL+FDC E ++
Sbjct: 70 SLLKREMDQTMAANAQKNKFLIDGFPRNQDNLQGWNKTMDGKADVSFVLFFDCNNEICIE 129
Query: 116 RVLSR--NQGRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVR 173
R L R + GR DDN ++++KR++ + P+ID Y G++ +I+A + DE+F +V
Sbjct: 130 RCLERGKSSGRSDDNRESLEKRIQTYLESTKPIIDLYEEMGKVKKIDASKSVDEVFGEVV 189
Query: 174 PVF 176
+F
Sbjct: 190 KIF 192
>UMPK_YEAST (P15700) Uridylate kinase (EC 2.7.4.-) (UK) (Uridine
monophosphate kinase) (UMP kinase)
Length = 204
Score = 145 bits (365), Expect = 7e-35
Identities = 73/176 (41%), Positives = 117/176 (66%), Gaps = 7/176 (3%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAM-VSDSEYGAMILETIREGRIVPSAVTV 62
GGPG+GKGTQC ++V+ + F HLSAGDLLR + S+YG +I I+EG+IVP +T+
Sbjct: 23 GGPGAGKGTQCEKLVKDYSFVHLSAGDLLRAEQGRAGSQYGELIKNCIKEGQIVPQEITL 82
Query: 63 RLI---LREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLS 119
L+ + + + KFLIDGFPR + I+FE E F+L+FDCPE+ M++R+L
Sbjct: 83 ALLRNAISDNVKANKHKFLIDGFPRKMDQAISFERDI-VESKFILFFDCPEDIMLERLLE 141
Query: 120 RNQ--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVR 173
R + GR DDNI++IKKR F+ ++PVI+++ + ++ R+ + +++++ V+
Sbjct: 142 RGKTSGRSDDNIESIKKRFNTFKETSMPVIEYFETKSKVVRVRCDRSVEDVYKDVQ 197
>KAD1_CAEEL (Q20140) Probable adenylate kinase isoenzyme F38B2.4 (EC
2.7.4.3) (ATP-AMP transphosphorylase)
Length = 210
Score = 141 bits (356), Expect = 7e-34
Identities = 83/175 (47%), Positives = 107/175 (60%), Gaps = 8/175 (4%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IV +G HLS+GDLLR + S S GA + + G +VP V +
Sbjct: 27 GGPGSGKGTQCDKIVAKYGLTHLSSGDLLRDEVKSGSPRGAQLTAIMESGALVPLEVVLD 86
Query: 64 LI----LREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLS 119
L+ L+ ++ G ++ FLIDG+PR FE E VL+FD EE +VKR+L
Sbjct: 87 LVKEAMLKAIEKG-SKGFLIDGYPREVAQGQQFESEI-QEAKLVLFFDVAEETLVKRLLH 144
Query: 120 RNQ--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
R Q GR DDN DTIKKRL F PV+D+Y +G+L RINA G+ D+IF V
Sbjct: 145 RAQTSGRADDNADTIKKRLHTFVTSTAPVVDYYESKGKLVRINAEGSVDDIFAVV 199
>KAD5_HUMAN (Q9Y6K8) Adenylate kinase isoenzyme 5 (EC 2.7.4.3)
(ATP-AMP transphosphorylase)
Length = 198
Score = 141 bits (355), Expect = 1e-33
Identities = 72/173 (41%), Positives = 107/173 (61%), Gaps = 5/173 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC ++VE +GF HLS G+LLR+ + S+SE +I + + G +VPS + +
Sbjct: 18 GGPGSGKGTQCEKLVEKYGFTHLSTGELLREELASESERSKLIRDIMERGDLVPSGIVLE 77
Query: 64 LILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRN 121
L+ M GD R FLIDG+PR + F G +P V+ DC + M R+L R+
Sbjct: 78 LLKEAMVASLGDTRGFLIDGYPREVKQGEEFGRRIG-DPQLVICMDCSADTMTNRLLQRS 136
Query: 122 QGR--IDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
+ +DD TI KRL+ + ++PVI +Y + +LH+INA GT +++F Q+
Sbjct: 137 RSSLPVDDTTKTIAKRLEAYYRASIPVIAYYETKTQLHKINAEGTPEDVFLQL 189
>KAD1_PIG (P00571) Adenylate kinase isoenzyme 1 (EC 2.7.4.3)
(ATP-AMP transphosphorylase) (AK1) (Myokinase)
Length = 194
Score = 140 bits (354), Expect = 1e-33
Identities = 75/173 (43%), Positives = 108/173 (62%), Gaps = 5/173 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IV+ +G+ HLS GDLLR + S S G M+ E + +G++VP +
Sbjct: 15 GGPGSGKGTQCEKIVQKYGYTHLSTGDLLRAEVSSGSARGKMLSEIMEKGQLVPLETVLD 74
Query: 64 LILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRN 121
++ M + ++ FLIDG+PR + FE G +P +LY D E M KR+L R
Sbjct: 75 MLRDAMVAKVDTSKGFLIDGYPREVKQGEEFERKIG-QPTLLLYVDAGPETMTKRLLKRG 133
Query: 122 Q--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
+ GR+DDN +TIKKRL+ + PVI Y +RG + ++NA G+ D++F QV
Sbjct: 134 ETSGRVDDNEETIKKRLETYYKATEPVIAFYEKRGIVRKVNAEGSVDDVFSQV 186
>KAD1_BOVIN (P00570) Adenylate kinase isoenzyme 1 (EC 2.7.4.3)
(ATP-AMP transphosphorylase) (AK1) (Myokinase)
Length = 194
Score = 137 bits (346), Expect = 1e-32
Identities = 74/173 (42%), Positives = 106/173 (60%), Gaps = 5/173 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IV+ +G+ HLS GDLLR + S S G M+ E + +G++VP +
Sbjct: 15 GGPGSGKGTQCEKIVQKYGYTHLSTGDLLRAEVSSGSARGKMLSEIMEKGQLVPLETVLD 74
Query: 64 LILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRN 121
++ M + ++ FLIDG+PR + FE +P +LY D E M KR+L R
Sbjct: 75 MLRDAMVAKVNTSKGFLIDGYPRQVQQGEEFERRI-AQPTLLLYVDAGPETMQKRLLKRG 133
Query: 122 Q--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
+ GR+DDN +TIKKRL+ + PVI Y +RG + ++NA G+ D +F QV
Sbjct: 134 ETSGRVDDNEETIKKRLETYYKATEPVIAFYEKRGIVRKVNAEGSVDNVFSQV 186
>KAD1_HUMAN (P00568) Adenylate kinase isoenzyme 1 (EC 2.7.4.3)
(ATP-AMP transphosphorylase) (AK1) (Myokinase)
Length = 194
Score = 137 bits (345), Expect = 1e-32
Identities = 73/173 (42%), Positives = 106/173 (61%), Gaps = 5/173 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IV+ +G+ HLS GDLLR + S S G + E + +G++VP +
Sbjct: 15 GGPGSGKGTQCEKIVQKYGYTHLSTGDLLRSEVSSGSARGKKLSEIMEKGQLVPLETVLD 74
Query: 64 LILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRN 121
++ M + ++ FLIDG+PR + FE G +P +LY D E M +R+L R
Sbjct: 75 MLRDAMVAKVNTSKGFLIDGYPREVQQGEEFERRIG-QPTLLLYVDAGPETMTQRLLKRG 133
Query: 122 Q--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
+ GR+DDN +TIKKRL+ + PVI Y +RG + ++NA G+ D +F QV
Sbjct: 134 ETSGRVDDNEETIKKRLETYYKATEPVIAFYEKRGIVRKVNAEGSVDSVFSQV 186
>KAD1_CHICK (P05081) Adenylate kinase isoenzyme 1 (EC 2.7.4.3)
(ATP-AMP transphosphorylase) (AK1) (Myokinase)
Length = 194
Score = 136 bits (342), Expect = 3e-32
Identities = 76/173 (43%), Positives = 105/173 (59%), Gaps = 5/173 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IV +G+ HLS GDLLR + S SE G + + +G +VP +
Sbjct: 16 GGPGSGKGTQCEKIVHKYGYTHLSTGDLLRAEVSSGSERGKKLQAIMEKGELVPLDTVLD 75
Query: 64 LILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRN 121
++ M + ++ FLIDG+PR + FE P +LY D +E MVKR+L R
Sbjct: 76 MLRDAMLAKADTSKGFLIDGYPREVKQGEEFEKKI-APPTLLLYVDAGKETMVKRLLKRG 134
Query: 122 Q--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
+ GR+DDN +TIKKRL+ + PVI Y RG + ++NA GT DE+F+QV
Sbjct: 135 ETSGRVDDNEETIKKRLETYYKATEPVIAFYKGRGIVRQLNAEGTVDEVFQQV 187
>KAD5_MOUSE (Q920P5) Adenylate kinase isoenzyme 5 (EC 2.7.4.3)
(ATP-AMP transphosphorylase)
Length = 193
Score = 135 bits (340), Expect = 5e-32
Identities = 69/175 (39%), Positives = 107/175 (60%), Gaps = 5/175 (2%)
Query: 2 LSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVT 61
L GGPGSGKGTQC ++ E +GF HLS G+LLR+ + S+SE +I + + G +VPS V
Sbjct: 12 LMGGPGSGKGTQCEKLAEKYGFTHLSTGELLRQELTSESERSKLIRDIMERGDLVPSGVV 71
Query: 62 VRLILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLS 119
+ L+ M G+ + FLIDG+PR + F G +P V+ DC + M R+L
Sbjct: 72 LELLKEAMVASLGNTKGFLIDGYPREVKQGEKFGRRIG-DPHLVICMDCSADTMTNRLLQ 130
Query: 120 RNQG--RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
R+Q R +D +I KRL+ + ++PV+ +Y R+ +L ++NA GT +++F Q+
Sbjct: 131 RSQSSQRGEDGAKSIAKRLEAYHRASIPVVTYYERKTQLRKVNAEGTPEQVFLQL 185
>KAD1_MOUSE (Q9R0Y5) Adenylate kinase isoenzyme 1 (EC 2.7.4.3)
(ATP-AMP transphosphorylase) (AK1) (Myokinase)
Length = 194
Score = 135 bits (339), Expect = 7e-32
Identities = 73/173 (42%), Positives = 104/173 (59%), Gaps = 5/173 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IV+ +G+ HLS GDLLR + S SE G + + +G +VP +
Sbjct: 15 GGPGSGKGTQCEKIVQKYGYTHLSTGDLLRAEVSSGSERGKKLSAIMEKGELVPLDTVLD 74
Query: 64 LILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRN 121
++ M + + FLIDG+PR + FE G +P +LY D E M +R+L R
Sbjct: 75 MLRDAMLAKVDSSNGFLIDGYPREVKQGEEFEQKIG-QPTLLLYVDAGAETMTQRLLKRG 133
Query: 122 Q--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
+ GR+DDN +TIKKRL+ + PVI Y +RG + ++NA GT D +F +V
Sbjct: 134 ETSGRVDDNEETIKKRLETYYNATEPVISFYDKRGIVRKVNAEGTVDTVFSEV 186
>KAD1_RAT (P39069) Adenylate kinase isoenzyme 1 (EC 2.7.4.3)
(ATP-AMP transphosphorylase) (AK1) (Myokinase)
Length = 194
Score = 134 bits (338), Expect = 9e-32
Identities = 75/174 (43%), Positives = 105/174 (60%), Gaps = 7/174 (4%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IV+ +G+ HLS GDLLR + S S G M+ + +G +VP TV
Sbjct: 15 GGPGSGKGTQCEKIVQKYGYTHLSTGDLLRAEVSSGSSRGKMLSSIMEKGELVP-LETVL 73
Query: 64 LILREMQYG---DNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSR 120
+LR+ + + FLIDG+PR + FE +P +LY D E M +R+L R
Sbjct: 74 DMLRDAMFAKVDSSNGFLIDGYPREVKQGEEFERKI-AQPTLLLYVDAGPETMTQRLLKR 132
Query: 121 NQ--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
+ GR+DDN +TIKKRL+ + PVI Y +RG + ++NA G+ D +F QV
Sbjct: 133 GETSGRVDDNEETIKKRLETYYKATEPVISFYDKRGIVRKVNAEGSVDTVFSQV 186
>KAD1_RABIT (P00569) Adenylate kinase isoenzyme 1 (EC 2.7.4.3)
(ATP-AMP transphosphorylase) (AK1) (Myokinase)
Length = 194
Score = 134 bits (336), Expect = 2e-31
Identities = 73/173 (42%), Positives = 104/173 (59%), Gaps = 5/173 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IV +G+ HLS GDLLR + S S G + E + +G++VP +
Sbjct: 15 GGPGSGKGTQCEKIVHKYGYTHLSTGDLLRAEVSSGSARGKKLSEIMEKGQLVPLETVLD 74
Query: 64 LILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRN 121
++ M + ++ FLIDG+PR + FE +P +LY D E M KR+L R
Sbjct: 75 MLRDAMVAKADTSKGFLIDGYPRQVQQGEEFERRI-AQPTLLLYVDAGPETMQKRLLKRG 133
Query: 122 Q--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQV 172
+ GR+DDN +TIKKRL+ + PVI Y +RG + ++NA G+ D +F QV
Sbjct: 134 ETSGRVDDNEETIKKRLETYYKATEPVIAFYEKRGIVRKVNAEGSVDNVFSQV 186
>KAD_CYPCA (P12115) Adenylate kinase (EC 2.7.4.3) (ATP-AMP
transphosphorylase)
Length = 193
Score = 129 bits (323), Expect = 5e-30
Identities = 73/170 (42%), Positives = 103/170 (59%), Gaps = 5/170 (2%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IVE +G+ HLS+GDLLR + S SE G + +++G +VP +
Sbjct: 14 GGPGSGKGTQCEKIVEKYGYTHLSSGDLLRAEVASGSERGKQLQAIMQKGELVPLDTVLD 73
Query: 64 LILREM--QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRN 121
+I M + ++ +LIDG+PR + FE G P +LY D E MV+R++ R
Sbjct: 74 MIKDAMIAKADVSKGYLIDGYPREVKQGEEFEKKIGA-PALLLYIDAKAETMVQRLMKRG 132
Query: 122 Q--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIF 169
Q GR DDN +TIKKRL ++ PVI +Y RG + +IN+ DE+F
Sbjct: 133 QTSGRSDDNEETIKKRLDLYYKATEPVIAYYETRGIVRKINSELPVDEVF 182
>UMPK_SCHPO (O59771) Probable uridylate kinase (EC 2.7.4.-) (UK)
(Uridine monophosphate kinase) (UMP kinase)
Length = 191
Score = 127 bits (320), Expect = 1e-29
Identities = 74/174 (42%), Positives = 107/174 (60%), Gaps = 8/174 (4%)
Query: 4 GGPGSGKGTQCARIVETFG-FKHLSAGDLLRKAMVSD-SEYGAMILETIREGRIVPSAVT 61
GGPG+GKGTQC R+ E F F H+SAGD LR+ S+YG +I E I++G+IVP +T
Sbjct: 9 GGPGAGKGTQCDRLAEKFDKFVHISAGDCLREEQNRPGSKYGNLIKEYIKDGKIVPMEIT 68
Query: 62 VRLILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVL 118
+ L+ +M+ ++ KFLIDGFPR + FE F LYF C +E M+KR++
Sbjct: 69 ISLLETKMKECHDKGIDKFLIDGFPREMDQCEGFEKSV-CPAKFALYFRCGQETMLKRLI 127
Query: 119 SRNQ--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFE 170
R + GR DDNI++IKKR + ++PV+++ + RL I+A D +FE
Sbjct: 128 HRGKTSGRSDDNIESIKKRFVTYTKASMPVVEYLKSQNRLITIDAEQDPDAVFE 181
>KAD_PORGI (Q7MW54) Adenylate kinase (EC 2.7.4.3) (ATP-AMP
transphosphorylase)
Length = 194
Score = 120 bits (302), Expect = 1e-27
Identities = 65/181 (35%), Positives = 105/181 (57%), Gaps = 6/181 (3%)
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
++ G PGSGKGTQ ++ +GF+H+S G+LLR + + +E G I EG +VP ++
Sbjct: 5 LIFGAPGSGKGTQSEELIRRYGFRHISTGELLRAEIKAQTELGQAAAGYINEGHLVPDSL 64
Query: 61 TVRLILREMQ-YGDNRKFLIDGFPRSEENRIAFEHIT---GTEPDFVLYFDCPEEEMVKR 116
V ++ + + D + DGFPR+ A E + G + D VL PEE +++R
Sbjct: 65 IVDMMEKLISTLVDTEGIIFDGFPRTIPQAEAMETMLAHHGWKVDIVLNLQVPEEMLIER 124
Query: 117 VLSRNQ--GRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRP 174
+L+R + GR DDNI+TI+KRL V+ P++D + R+ LH + GT +EI ++ P
Sbjct: 125 LLNRGKVSGRSDDNIETIRKRLDVYANETAPLVDFFTRKNVLHNVVGTGTIEEIALRIAP 184
Query: 175 V 175
+
Sbjct: 185 I 185
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.142 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,261,529
Number of Sequences: 164201
Number of extensions: 914156
Number of successful extensions: 3133
Number of sequences better than 10.0: 262
Number of HSP's better than 10.0 without gapping: 223
Number of HSP's successfully gapped in prelim test: 39
Number of HSP's that attempted gapping in prelim test: 2534
Number of HSP's gapped (non-prelim): 319
length of query: 185
length of database: 59,974,054
effective HSP length: 104
effective length of query: 81
effective length of database: 42,897,150
effective search space: 3474669150
effective search space used: 3474669150
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC133139.3


