

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130806.4 - phase: 0
(359 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
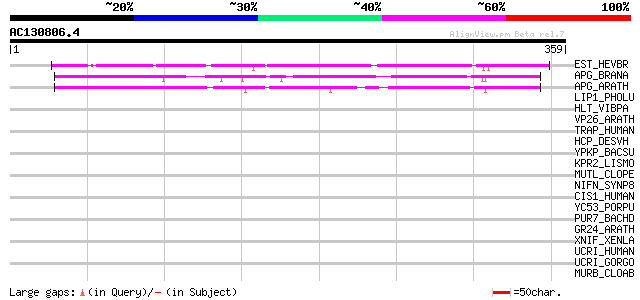
Score E
Sequences producing significant alignments: (bits) Value
EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early nodule... 168 2e-41
APG_BRANA (P40603) Anter-specific proline-rich protein APG (Prot... 96 1e-19
APG_ARATH (P40602) Anter-specific proline-rich protein APG precu... 95 2e-19
LIP1_PHOLU (P40601) Lipase 1 precursor (EC 3.1.1.3) (Triacylglyc... 34 0.50
HLT_VIBPA (Q99289) Thermolabile hemolysin precursor (TL) (Lecith... 34 0.65
VP26_ARATH (Q9T091) Vacuolar protein sorting 26 homolog (VPS26 p... 33 1.5
TRAP_HUMAN (Q9Y4A5) Transformation/transcription domain-associat... 32 1.9
HCP_DESVH (P31101) Hydroxylamine reductase (EC 1.7.-.-) (Hybrid-... 32 2.5
YPKP_BACSU (P54174) Hypothetical protein ypkP 32 3.2
KPR2_LISMO (Q8Y9L8) Ribose-phosphate pyrophosphokinase 2 (EC 2.7... 32 3.2
MUTL_CLOPE (Q8XL86) DNA mismatch repair protein mutL 31 4.2
NIFN_SYNP8 (O07356) Nitrogenase iron-molybdenum cofactor biosynt... 31 5.5
CIS1_HUMAN (Q96P20) Cold autoinflammatory syndrome 1 protein (Cr... 31 5.5
YC53_PORPU (P51202) Hypothetical 28.1 kDa protein ycf53 (ORF238) 30 7.2
PUR7_BACHD (Q9KF60) Phosphoribosylaminoimidazole-succinocarboxam... 30 7.2
GR24_ARATH (O81776) Glutamate receptor 2.4 precursor (Ligand-gat... 30 7.2
XNIF_XENLA (P35617) Low molecular weight neuronal intermediate f... 30 9.4
UCRI_HUMAN (P47985) Ubiquinol-cytochrome c reductase iron-sulfur... 30 9.4
UCRI_GORGO (Q69BK4) Ubiquinol-cytochrome c reductase iron-sulfur... 30 9.4
MURB_CLOAB (Q97LP4) UDP-N-acetylenolpyruvoylglucosamine reductas... 30 9.4
>EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early
nodule-specific protein homolog) (Latex allergen Hev b
13)
Length = 391
Score = 168 bits (426), Expect = 2e-41
Identities = 117/352 (33%), Positives = 177/352 (50%), Gaps = 42/352 (11%)
Query: 28 YEAFFNFGDSISDTGNAASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPF 87
+ A FNFGDS SDTG A+ F P+ NP PYG ++F +GR S+GRLIIDFIAE++ LP+
Sbjct: 32 FPAIFNFGDSNSDTGGKAAAFYPL-NP-PYGETFFHRSTGRYSDGRLIIDFIAESFNLPY 89
Query: 88 LPAYENKSIDQDIKKGVNFAFAGATV-LNVEYYVKNGLPLPDTNNSLSIQLGWFKNIKPL 146
L Y + S+ + K G +FA AG+T+ L +G P L +Q F+ P
Sbjct: 90 LSPYLS-SLGSNFKHGADFATAGSTIKLPTTIIPAHGGFSP---FYLDVQYSQFRQFIPR 145
Query: 147 LCKSKEDCNI---------YFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVPFMVEAIT 197
+E I YF+K+L+ +IG ND+ + + TV E+ VP +V + +
Sbjct: 146 SQFIRETGGIFAELVPEEYYFEKALYTF-DIGQNDLTEGFLNLTVEEVNATVPDLVNSFS 204
Query: 198 NTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNGQL 257
+ + GA + P+GC + + T +K D GC AYN + ++FN +L
Sbjct: 205 ANVKKIYDLGARTFWIHNTGPIGCLSFILTYFPWAEK---DSAGCAKAYNEVAQHFNHKL 261
Query: 258 KNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------G 306
K + LR+ P ++ D Y+ L+ P+++GF+ I CCG
Sbjct: 262 KEIVAQLRKDLPLATFVHVDIYSVKYSLFSEPEKHGFEFPLI--TCCGYGGKYNFSVTAP 319
Query: 307 CG---------SLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNP 349
CG ++ C+ PS R+NWDG H+TEAA + + G FS+P
Sbjct: 320 CGDTVTADDGTKIVVGSCACPSVRVNWDGAHYTEAANEYFFDQISTGAFSDP 371
>APG_BRANA (P40603) Anter-specific proline-rich protein APG (Protein
CEX) (Fragment)
Length = 449
Score = 95.9 bits (237), Expect = 1e-19
Identities = 96/348 (27%), Positives = 155/348 (43%), Gaps = 63/348 (18%)
Query: 30 AFFNFGDSISDTGNAASIFLPMP-NPIPYGSSY-FKHPSGRMSNGRLIIDFIAEAYGLP- 86
A F FGDSI DTGN ++ + N PYG + +GR SNGR+ D+I++ G+
Sbjct: 125 AVFFFGDSIFDTGNNNNLDTKLKCNYRPYGMDFPMGVATGRFSNGRVASDYISKYLGVKE 184
Query: 87 FLPAYENKSIDQ-------DIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSI---- 135
+PAY +K + Q D+ GV+FA GA L P T+ S +
Sbjct: 185 IVPAYVDKKLQQNNELQQSDLLTGVSFASGGAGYL------------PQTSESWKVTTML 232
Query: 136 -QLGWFKNIKPLLCK---SKEDCNIYFKKSLFIVGEIGGNDIM--------KHMKHKTVI 183
QL +F++ K + K K+ I K + +V G ND++ +H+K+
Sbjct: 233 DQLTYFQDYKKRMKKLVGKKKTKKIVSKGAAIVVA--GSNDLIYTYFGNGAQHLKN---- 286
Query: 184 ELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCL 243
++ M ++ + L GA + V G P+GC+ + ED
Sbjct: 287 DVDSFTTMMADSAASFVLQLYGYGARRIGVIGTPPIGCTPSQRVKKKKICNEDL------ 340
Query: 244 IAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKAC 303
N + FN +L + L + P I+Y D Y+ ++ ++P+ YGF++ I K C
Sbjct: 341 ---NYAAQLFNSKLVIILGQLSKTLPNSTIVYGDIYSIFSKMLESPEDYGFEE--IKKPC 395
Query: 304 C------GG--CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVE 343
C GG C S+ S + WDG H ++ AY++ + LV+
Sbjct: 396 CKIGLTKGGVFCKERTLKNMSNASSYLFWDGLHPSQRAYEISNRKLVK 443
>APG_ARATH (P40602) Anter-specific proline-rich protein APG
precursor
Length = 534
Score = 95.1 bits (235), Expect = 2e-19
Identities = 92/339 (27%), Positives = 148/339 (43%), Gaps = 41/339 (12%)
Query: 30 AFFNFGDSISDTGNAASIFLPMP-NPIPYGSSY-FKHPSGRMSNGRLIIDFIAEAYGLP- 86
A F FGDS+ DTGN ++ + N PYG + F+ +GR SNG + D++A+ G+
Sbjct: 204 AVFFFGDSVFDTGNNNNLETKIKSNYRPYGMDFKFRVATGRFSNGMVASDYLAKYMGVKE 263
Query: 87 FLPAYENKSID-QDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWFKNIKP 145
+PAY + I D+ GV+FA GA N +P+ D L+ + + +
Sbjct: 264 IVPAYLDPKIQPNDLLTGVSFASGGAGYNPTTSEAANAIPMLD---QLTYFQDYIEKVNR 320
Query: 146 LLCKSK--------EDCNIYFKKSLFIVGEIGG-NDIMKHMKHKTVIELREIVPFMVEAI 196
L+ + K E N K + IV +GG ND++ L+ + I
Sbjct: 321 LVRQEKSQYKLAGLEKTNQLISKGVAIV--VGGSNDLIITYFGSGAQRLKNDIDSYTTII 378
Query: 197 TNTTNVLIEE----GAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEY 252
++ + + GA + V G P+GC + KK+ +E N +
Sbjct: 379 ADSAASFVLQLYGYGARRIGVIGTPPLGCVPSQ----RLKKKKICNE-----ELNYASQL 429
Query: 253 FNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCGG------ 306
FN +L + L + P +Y D Y ++ +TP YGF++ K CC
Sbjct: 430 FNSKLLLILGQLSKTLPNSTFVYMDIYTIISQMLETPAAYGFEETK--KPCCKTGLLSAG 487
Query: 307 --CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVE 343
C + +C + S + WDG H T+ AYK I K L++
Sbjct: 488 ALCKKSTSKICPNTSSYLFWDGVHPTQRAYKTINKVLIK 526
>LIP1_PHOLU (P40601) Lipase 1 precursor (EC 3.1.1.3)
(Triacylglycerol lipase)
Length = 645
Score = 34.3 bits (77), Expect = 0.50
Identities = 44/197 (22%), Positives = 81/197 (40%), Gaps = 48/197 (24%)
Query: 18 NVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIID 77
+ +SNA+ +Y + FGDS+SD GN + N + +L D
Sbjct: 17 SAISNAH--AYNNLYVFGDSLSDGGNNGRYTVDGING---------------TESKLYND 59
Query: 78 FIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQL 137
FIA+ G+ + K G N+A GAT + L + +N+ +
Sbjct: 60 FIAQQLGIELV---------NSKKGGTNYAAGGATAV---------ADLNNKHNTQDQVM 101
Query: 138 GWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVPFMVEAIT 197
G+ + ++ D N + V IGGND+ +++ + ++I+ A +
Sbjct: 102 GYLAS-----HSNRADHNGMY------VHWIGGNDVDAALRNPA--DAQKIITESAMAAS 148
Query: 198 NTTNVLIEEGAVELVVP 214
+ + L+ GA ++VP
Sbjct: 149 SQVHALLNAGAGLVIVP 165
>HLT_VIBPA (Q99289) Thermolabile hemolysin precursor (TL)
(Lecithin-dependent haemolysin) (LDH) (Atypical
phospholipase) (Phospholipase A2) (Lysophospholipase)
Length = 418
Score = 33.9 bits (76), Expect = 0.65
Identities = 74/318 (23%), Positives = 116/318 (36%), Gaps = 69/318 (21%)
Query: 35 GDSISDTGNAASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENK 94
GDS+SDTGN IF P +S+F G SNG + ++IA+A LP
Sbjct: 151 GDSLSDTGN---IFNASQWRFPNPNSWF---LGHFSNGFVWTEYIAKAKNLPL------- 197
Query: 95 SIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDC 154
N+A GA N +Y G+ +S L + K L K+ +
Sbjct: 198 ---------YNWAVGGAAGEN-QYIALTGV-----GEQVSSYLTYAK-----LAKNYKPA 237
Query: 155 NIYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVP 214
N F E G ND M + + + E+ EA+ T + GA ++
Sbjct: 238 NTLFTL------EFGLNDFMNYNR-----GVPEVKADYAEALIRLT----DAGAKNFMLM 282
Query: 215 GNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKII 274
P A F + +E+ D+ + N E+ Q + K I
Sbjct: 283 -TLPDATKAPQF---KYSTQEEIDKIRAKVLEMN--EFIKAQ------AMYYKAQGYNIT 330
Query: 275 YFDYYNDAKRLYQTPQQYGF--DKDAIFKACCGGCGSLIAT-------VCSDPSKRINWD 325
FD + + L P+++GF D + T S K + WD
Sbjct: 331 LFDTHALFETLTSAPEEHGFVNASDPCLDINRSSSVDYMYTHALRSECAASGAEKFVFWD 390
Query: 326 GPHFTEAAYKLIAKGLVE 343
H T A ++ +A+ ++E
Sbjct: 391 VTHPTTATHRYVAEKMLE 408
>VP26_ARATH (Q9T091) Vacuolar protein sorting 26 homolog (VPS26
protein homolog)
Length = 303
Score = 32.7 bits (73), Expect = 1.5
Identities = 15/39 (38%), Positives = 24/39 (61%)
Query: 100 IKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLG 138
+K V +AG+ + E V+N PLPD NNS+ +++G
Sbjct: 131 LKVTVTRGYAGSILEYQELVVRNYAPLPDINNSIKMEVG 169
>TRAP_HUMAN (Q9Y4A5) Transformation/transcription domain-associated
protein (350/400 kDa PCAF-associated factor)
(PAF350/400) (STAF40) (Tra1 homolog)
Length = 3859
Score = 32.3 bits (72), Expect = 1.9
Identities = 41/148 (27%), Positives = 65/148 (43%), Gaps = 27/148 (18%)
Query: 87 FLPAYENKSIDQDIKKGVNFA-------FAGATVLNVEYYVKNGLPLPDTNNSLSIQLGW 139
F+P + K D+ I G + A +T+ ++ ++V+ LPL D SL++QL +
Sbjct: 347 FIPCMD-KLFDESILIGSGYTARETLRPLAYSTLADLVHHVRQHLPLSDL--SLAVQL-F 402
Query: 140 FKNI---------KPLLCK---SKEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIELRE 187
KNI + + CK + DC KS G G D++ M V++
Sbjct: 403 AKNIDDESLPSSIQTMSCKLLLNLVDC--IRSKSEQESGN--GRDVLMRMLEVFVLKFHT 458
Query: 188 IVPFMVEAITNTTNVLIEEGAVELVVPG 215
I + + AI E GAVE +PG
Sbjct: 459 IARYQLSAIFKKCKPQSELGAVEAALPG 486
>HCP_DESVH (P31101) Hydroxylamine reductase (EC 1.7.-.-)
(Hybrid-cluster protein) (Prismane protein) (HCP)
Length = 553
Score = 32.0 bits (71), Expect = 2.5
Identities = 50/224 (22%), Positives = 90/224 (39%), Gaps = 23/224 (10%)
Query: 118 YYVKNGL----PLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI 173
Y KNG PLP+ + + K + + E+ ++ + L I+G G +
Sbjct: 102 YKAKNGKDFSEPLPEAATWTGDSTAFAEKAKSVGILATENEDVRSLRELLIIGLKG---V 158
Query: 174 MKHMKHKTVIELR--EIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNS 231
+ +H V+ R EI FM+EA+ +TT L + V LV+ + A+ N+
Sbjct: 159 AAYAEHAAVLGFRKTEIDEFMLEALASTTKDLSVDEMVALVMKAGGMAVTTMALLDEANT 218
Query: 232 NK--KEDYDEFGCLIAYNNLIEYFNGQLKNSIETLRQK-------HPEVKIIYFDYYNDA 282
+ + + N I LK+ E L+Q + +++ +YY
Sbjct: 219 TTYGNPEITQVNIGVGKNPGILISGHDLKDMAELLKQTEGTGVDVYTHGEMLPANYYPAF 278
Query: 283 KRLYQTPQQYG---FDKDAIFKACCGGCGSLIATVCSDPSKRIN 323
K+ YG + ++ F++ G L+ T C P K+ N
Sbjct: 279 KKYPHFVGNYGGSWWQQNPEFESFNGPI--LLTTNCLVPLKKEN 320
>YPKP_BACSU (P54174) Hypothetical protein ypkP
Length = 206
Score = 31.6 bits (70), Expect = 3.2
Identities = 30/126 (23%), Positives = 56/126 (43%), Gaps = 27/126 (21%)
Query: 47 IFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYG---------LPFLPAYENK--- 94
+ + P PIP+G + HP R++ LI +I E G LPF+ + ++
Sbjct: 60 VTIHQPEPIPHGPVLYVHPRLRLAELALIAGYIEEPAGFIANPKVFRLPFIGQWLDRMDV 119
Query: 95 -----------SIDQDIKKGVNFAFA-GATVLNVEYYVKNGLPL--PDTNNSLSIQLG-W 139
+ + ++KG + + ++ VE + LPL +T + ++Q G +
Sbjct: 120 ISDGDSGKVYEDVTKQLEKGQSLILSLDGSIDPVELAARYHLPLVMVETKGTDNMQNGSF 179
Query: 140 FKNIKP 145
FK +KP
Sbjct: 180 FKRLKP 185
>KPR2_LISMO (Q8Y9L8) Ribose-phosphate pyrophosphokinase 2 (EC
2.7.6.1) (RPPK 2) (Phosphoribosyl pyrophosphate
synthetase 2) (P-Rib-PP synthetase 2) (PRPP synthetase
2)
Length = 311
Score = 31.6 bits (70), Expect = 3.2
Identities = 20/63 (31%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Query: 173 IMKHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSN 232
++ +K K I + +I+ V A T + ++L+E+GAVE++ + A L NSN
Sbjct: 206 VIGDVKGKVAIVVDDIIDTGVRA-TTSADILLEKGAVEVIACATHSVMAGNATERLQNSN 264
Query: 233 KKE 235
KE
Sbjct: 265 IKE 267
>MUTL_CLOPE (Q8XL86) DNA mismatch repair protein mutL
Length = 674
Score = 31.2 bits (69), Expect = 4.2
Identities = 29/110 (26%), Positives = 43/110 (38%), Gaps = 14/110 (12%)
Query: 177 MKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKED 236
M T IE+R++ F V A E A+ + + F L N+NKK
Sbjct: 139 MNRGTQIEVRDLF-FNVPARKKFLKTTARESALINDLVNRISLANPDVSFKLFNNNKKI- 196
Query: 237 YDEFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLY 286
L Y NG+L + I T+ K +IYF+ + D +Y
Sbjct: 197 ------------LNTYGNGKLIDVIRTIYGKSTAENLIYFEEHKDTASVY 234
>NIFN_SYNP8 (O07356) Nitrogenase iron-molybdenum cofactor
biosynthesis protein nifN
Length = 461
Score = 30.8 bits (68), Expect = 5.5
Identities = 23/91 (25%), Positives = 39/91 (42%), Gaps = 17/91 (18%)
Query: 182 VIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFG 241
V RE +P A+T + +L + VE A+ T+V+ NK E
Sbjct: 53 VRHFRESIPLSTTAMTEVSTILGGQDHVE------------QAILTIVDKNKPEIIG--- 97
Query: 242 CLIAYNNLIEYFNGQLKNSIETLRQKHPEVK 272
+ L E ++ ++ +RQKHP++K
Sbjct: 98 --LLTTGLTETRGDDMEGILKDIRQKHPQLK 126
>CIS1_HUMAN (Q96P20) Cold autoinflammatory syndrome 1 protein
(Cryopyrin) (NACHT-, LRR- and PYD-containing protein 3)
(PYRIN-containing APAF1-like protein 1)
(Angiotensin/vasopressin receptor AII/AVP-like)
Length = 1034
Score = 30.8 bits (68), Expect = 5.5
Identities = 13/35 (37%), Positives = 25/35 (71%), Gaps = 1/35 (2%)
Query: 242 CLIAYNNLIE-YFNGQLKNSIETLRQKHPEVKIIY 275
CL+ L E YFN + K+++ETL+++ PE+ +++
Sbjct: 996 CLLQNLGLSEMYFNYETKSALETLQEEKPELTVVF 1030
>YC53_PORPU (P51202) Hypothetical 28.1 kDa protein ycf53 (ORF238)
Length = 238
Score = 30.4 bits (67), Expect = 7.2
Identities = 33/119 (27%), Positives = 47/119 (38%), Gaps = 21/119 (17%)
Query: 143 IKPLLCKSKEDCNIY--FKKSLFIVGEIGGNDIMKHMKHKTVIELREIVP---------F 191
I+ L K KE N KK L I+ I ND ++ K + R P
Sbjct: 5 IQVQLSKLKETSNSENAIKKQLEIIETIEYNDYLQLKKLAEIFHYRITKPEYISNCVDGL 64
Query: 192 MVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLI 250
+ E + N+ N I + +L G P+ SNK+ DY + L+ NNLI
Sbjct: 65 IYEKLLNSQNSEIIKFTSKLCPKGIVPL----------KSNKQMDYQDLQILLVQNNLI 113
>PUR7_BACHD (Q9KF60) Phosphoribosylaminoimidazole-succinocarboxamide
synthase (EC 6.3.2.6) (SAICAR synthetase)
Length = 237
Score = 30.4 bits (67), Expect = 7.2
Identities = 21/61 (34%), Positives = 29/61 (47%), Gaps = 8/61 (13%)
Query: 238 DEFGCL-IAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYND-------AKRLYQTP 289
DE G L + Y N FNG+ K SIE + + E+ + F Y + KRL +T
Sbjct: 19 DEEGLLWVEYKNSATAFNGEKKASIEGKARLNNEISSLIFSYLKEQGVDSHFVKRLSETE 78
Query: 290 Q 290
Q
Sbjct: 79 Q 79
>GR24_ARATH (O81776) Glutamate receptor 2.4 precursor (Ligand-gated
ion channel 2.4)
Length = 896
Score = 30.4 bits (67), Expect = 7.2
Identities = 27/130 (20%), Positives = 56/130 (42%), Gaps = 6/130 (4%)
Query: 16 LQNVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPIPYGSSYFKHPS-GRMSNGRL 74
++NV++ P++Y+ + ++G S +P +P K PS G +S +
Sbjct: 664 IKNVLAKGGPVAYQRDSFVLGKLRESGFPESRLVPFTSPEKCEELLNKGPSKGGVSAAFM 723
Query: 75 IIDFIAEAYGLPFLPAYENKSIDQDIKK-GVNFAFAGATVLNVEYYVKNGLPLPDTNNSL 133
+ ++ G + Y+ + D+ G F V +V + L + ++N +
Sbjct: 724 EVPYVRVFLG-QYCKKYKMVEVPFDVDGFGFVFPIGSPLVADVSRAI---LKVAESNKAT 779
Query: 134 SIQLGWFKNI 143
++ WFKNI
Sbjct: 780 QLETAWFKNI 789
>XNIF_XENLA (P35617) Low molecular weight neuronal intermediate
filament (XNIF)
Length = 470
Score = 30.0 bits (66), Expect = 9.4
Identities = 21/74 (28%), Positives = 34/74 (45%), Gaps = 4/74 (5%)
Query: 228 LVNSNKKEDY----DEFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAK 283
+V +N+KE D F I + +E N L++ + LRQKH E + Y + +
Sbjct: 83 IVRTNEKEQLQGLNDRFVTYIEKVHHLEQQNKLLESEVTLLRQKHSEPSRLSHIYEQEIR 142
Query: 284 RLYQTPQQYGFDKD 297
L ++ DKD
Sbjct: 143 ELRSKLEEQEQDKD 156
>UCRI_HUMAN (P47985) Ubiquinol-cytochrome c reductase iron-sulfur
subunit, mitochondrial precursor (EC 1.10.2.2) (Rieske
iron-sulfur protein) (RISP)
Length = 274
Score = 30.0 bits (66), Expect = 9.4
Identities = 34/147 (23%), Positives = 61/147 (41%), Gaps = 20/147 (13%)
Query: 45 ASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGV 104
AS+ L +P + Y + K P DF +E L L + ++ + +KG
Sbjct: 66 ASVGLNVPASVCYSHTDIKVP-----------DF-SEYRRLEVLDSTKSSRESSEARKGF 113
Query: 105 NFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFK---KS 161
++ G T + V Y KN + ++ S S + I+ L E N+ FK K
Sbjct: 114 SYLVTGVTTVGVAYAAKNAVTQFVSSMSASADVLALAKIEIKLSDIPEGKNMAFKWRGKP 173
Query: 162 LFIVGEIGGNDIMKHMKHKTVIELREI 188
LF+ + K ++ + +EL ++
Sbjct: 174 LFV-----RHRTQKEIEQEAAVELSQL 195
>UCRI_GORGO (Q69BK4) Ubiquinol-cytochrome c reductase iron-sulfur
subunit, mitochondrial precursor (EC 1.10.2.2) (Rieske
iron-sulfur protein) (RISP)
Length = 274
Score = 30.0 bits (66), Expect = 9.4
Identities = 34/147 (23%), Positives = 61/147 (41%), Gaps = 20/147 (13%)
Query: 45 ASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGV 104
AS+ L +P + Y + K P DF +E L L + ++ + +KG
Sbjct: 66 ASVGLNVPASVCYSHTDIKVP-----------DF-SEYRRLEVLDSTKSSRESSEARKGF 113
Query: 105 NFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFK---KS 161
++ G T + V Y KN + ++ S S + I+ L E N+ FK K
Sbjct: 114 SYLVTGVTTVGVAYAAKNAVTQFVSSMSASADVLALAKIEIKLSDIPEGKNMAFKWRGKP 173
Query: 162 LFIVGEIGGNDIMKHMKHKTVIELREI 188
LF+ + K ++ + +EL ++
Sbjct: 174 LFV-----RHRTQKEIEQEAAVELSQL 195
>MURB_CLOAB (Q97LP4) UDP-N-acetylenolpyruvoylglucosamine reductase
(EC 1.1.1.158) (UDP-N-acetylmuramate dehydrogenase)
Length = 305
Score = 30.0 bits (66), Expect = 9.4
Identities = 37/149 (24%), Positives = 59/149 (38%), Gaps = 33/149 (22%)
Query: 175 KHMKHKTVIELREIVPFMVEAITNTTNVLIEEG-----AVELVVPGNFPMG-------CS 222
K+ + +IEL E I N +N+L+ +G A++L+ +G C
Sbjct: 46 KYEEINRIIELCEKYDVNYYIIGNGSNLLVRDGGLRGVAIKLLKLNKLQIGNNKIIAGCG 105
Query: 223 AAMFTLVNSNKKEDYD----EFGCLI------AYNNLIEYFNGQLKNSIETLRQKHPEVK 272
+ L S K D EF C I A +NG++ N +E+
Sbjct: 106 VPLGYL--SRKARDKSLTGLEFACGIPGSVGGAVAMNAGAYNGEISNVVES--------- 154
Query: 273 IIYFDYYNDAKRLYQTPQQYGFDKDAIFK 301
++ D KRLY+ Q+G+ AI K
Sbjct: 155 VLVIDNKGKMKRLYRDELQFGYRSSAILK 183
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.140 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,900,878
Number of Sequences: 164201
Number of extensions: 2099206
Number of successful extensions: 4778
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 4761
Number of HSP's gapped (non-prelim): 21
length of query: 359
length of database: 59,974,054
effective HSP length: 111
effective length of query: 248
effective length of database: 41,747,743
effective search space: 10353440264
effective search space used: 10353440264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC130806.4


