

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130806.3 - phase: 0
(320 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
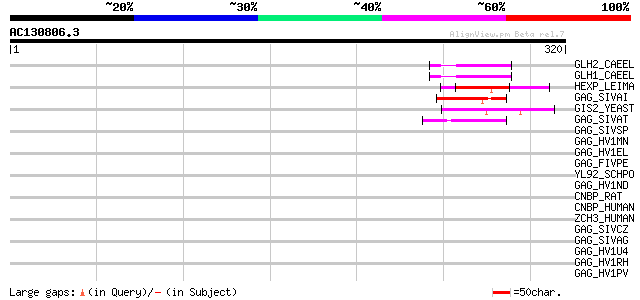
Score E
Sequences producing significant alignments: (bits) Value
GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-... 49 1e-05
GLH1_CAEEL (P34689) ATP-dependent RNA helicase glh-1 (EC 3.6.1.-... 49 1e-05
HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding p... 46 1e-04
GAG_SIVAI (Q02843) Gag polyprotein [Contains: Core protein p17; ... 46 1e-04
GIS2_YEAST (P53849) Zinc-finger protein GIS2 44 4e-04
GAG_SIVAT (P05892) Gag polyprotein [Contains: Core protein p17; ... 44 5e-04
GAG_SIVSP (P19504) Gag polyprotein [Contains: Core protein p17; ... 43 0.001
GAG_HV1MN (P05888) Gag polyprotein [Contains: Core protein p17 (... 43 0.001
GAG_HV1EL (P04592) Gag polyprotein [Contains: Core protein p17 (... 43 0.001
GAG_FIVPE (P16087) Gag polyprotein [Contains: Core protein p15; ... 43 0.001
YL92_SCHPO (Q9HFF2) Hypothetical protein C683.02c in chromosome I 42 0.002
GAG_HV1ND (P18800) Gag polyprotein [Contains: Core protein p17 (... 42 0.002
CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP) (... 42 0.002
CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)... 42 0.002
ZCH3_HUMAN (Q9NUD5) Zinc finger CCHC domain containing protein 3 42 0.002
GAG_SIVCZ (P17282) Gag polyprotein [Contains: Core protein p18; ... 42 0.002
GAG_SIVAG (P27978) Gag polyprotein [Contains: Core protein p17; ... 42 0.002
GAG_HV1U4 (P24736) Gag polyprotein [Contains: Core protein p17 (... 42 0.002
GAG_HV1RH (P05890) Gag polyprotein [Contains: Core protein p17 (... 42 0.002
GAG_HV1PV (P03350) Gag polyprotein [Contains: Core protein p17 (... 42 0.002
>GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-)
(Germline helicase-2)
Length = 974
Score = 49.3 bits (116), Expect = 1e-05
Identities = 21/47 (44%), Positives = 28/47 (58%), Gaps = 8/47 (17%)
Query: 243 EDRRPKKKDALAEIICFNCGEKGHKSNACPSEIKRCFRCGKKGHKSN 289
E+R P K CFNC +GH+S CP + CF CG++GH+SN
Sbjct: 448 EERGPMK--------CFNCKGEGHRSAECPEPPRGCFNCGEQGHRSN 486
Score = 39.7 bits (91), Expect = 0.010
Identities = 16/36 (44%), Positives = 21/36 (57%), Gaps = 5/36 (13%)
Query: 258 CFNCGEKGHKSNACPS-----EIKRCFRCGKKGHKS 288
CFNC + GH+SN CP E + C+ C + GH S
Sbjct: 373 CFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNS 408
Score = 39.7 bits (91), Expect = 0.010
Identities = 16/36 (44%), Positives = 21/36 (57%), Gaps = 5/36 (13%)
Query: 258 CFNCGEKGHKSNACPS-----EIKRCFRCGKKGHKS 288
CFNC + GH+SN CP E + C+ C + GH S
Sbjct: 259 CFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNS 294
Score = 38.1 bits (87), Expect = 0.029
Identities = 13/19 (68%), Positives = 16/19 (83%)
Query: 258 CFNCGEKGHKSNACPSEIK 276
CFNCGE+GH+SN CP+ K
Sbjct: 475 CFNCGEQGHRSNECPNPAK 493
Score = 36.2 bits (82), Expect = 0.11
Identities = 30/114 (26%), Positives = 40/114 (34%), Gaps = 44/114 (38%)
Query: 247 PKKKDALAEIICFNCGEKGHKSNACPSEIK------------------------------ 276
P+ K +C+NC + GH S CP E K
Sbjct: 387 PEPKKEREPRVCYNCQQPGHNSRDCPEERKPREGRNGFTSGFGGGNDGGFGGGNAEGFGN 446
Query: 277 -------RCFRCGKKGHKSNAEII-----CFNIQHEGRASHRGRSPAN-RSSAE 317
+CF C +GH+S AE CFN +G S+ +PA R AE
Sbjct: 447 NEERGPMKCFNCKGEGHRS-AECPEPPRGCFNCGEQGHRSNECPNPAKPREGAE 499
Score = 32.0 bits (71), Expect = 2.1
Identities = 12/30 (40%), Positives = 16/30 (53%)
Query: 247 PKKKDALAEIICFNCGEKGHKSNACPSEIK 276
P+ K +C+NC + GH S CP E K
Sbjct: 273 PEPKKEREPRVCYNCQQPGHNSRDCPEERK 302
>GLH1_CAEEL (P34689) ATP-dependent RNA helicase glh-1 (EC 3.6.1.-)
(Germline helicase-1)
Length = 763
Score = 49.3 bits (116), Expect = 1e-05
Identities = 21/47 (44%), Positives = 28/47 (58%), Gaps = 8/47 (17%)
Query: 243 EDRRPKKKDALAEIICFNCGEKGHKSNACPSEIKRCFRCGKKGHKSN 289
E+R P K CFNC +GH+S CP + CF CG++GH+SN
Sbjct: 237 EERGPMK--------CFNCKGEGHRSAECPEPPRGCFNCGEQGHRSN 275
Score = 38.1 bits (87), Expect = 0.029
Identities = 13/19 (68%), Positives = 16/19 (83%)
Query: 258 CFNCGEKGHKSNACPSEIK 276
CFNCGE+GH+SN CP+ K
Sbjct: 264 CFNCGEQGHRSNECPNPAK 282
Score = 37.4 bits (85), Expect = 0.050
Identities = 15/36 (41%), Positives = 21/36 (57%), Gaps = 5/36 (13%)
Query: 258 CFNCGEKGHKSNACPS-----EIKRCFRCGKKGHKS 288
CFNC + GH+S+ CP E + C+ C + GH S
Sbjct: 160 CFNCQQPGHRSSDCPEPRKEREPRVCYNCQQPGHTS 195
>HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding
protein)
Length = 271
Score = 46.2 bits (108), Expect = 1e-04
Identities = 19/38 (50%), Positives = 23/38 (60%), Gaps = 7/38 (18%)
Query: 258 CFNCGEKGHKSNACPSEIK-------RCFRCGKKGHKS 288
CF CGE+GH S CP+E + CFRCG+ GH S
Sbjct: 45 CFRCGEEGHMSRECPNEARSGAAGAMTCFRCGEAGHMS 82
Score = 45.4 bits (106), Expect = 2e-04
Identities = 24/70 (34%), Positives = 31/70 (44%), Gaps = 7/70 (10%)
Query: 249 KKDALAEIICFNCGEKGHKSNACPSEIK-------RCFRCGKKGHKSNAEIICFNIQHEG 301
+ A + CF CGE GH S CP+ K C++CG++GH S G
Sbjct: 63 RSGAAGAMTCFRCGEAGHMSRDCPNSAKPGAAKGFECYKCGQEGHLSRDCPSSQGGSRGG 122
Query: 302 RASHRGRSPA 311
RGRS A
Sbjct: 123 YGQKRGRSGA 132
Score = 41.6 bits (96), Expect = 0.003
Identities = 17/37 (45%), Positives = 21/37 (55%), Gaps = 6/37 (16%)
Query: 258 CFNCGEKGHKSNACPSE------IKRCFRCGKKGHKS 288
C+ CGE GH S CPS + C++CGK GH S
Sbjct: 198 CYKCGESGHMSRECPSAGSTGSGDRACYKCGKPGHIS 234
Score = 41.2 bits (95), Expect = 0.003
Identities = 22/66 (33%), Positives = 33/66 (49%), Gaps = 13/66 (19%)
Query: 245 RRPKKKDALAEIICFNCGEKGHKSNACPSEIKR-------CFRCGKKGHKSNAEIICFNI 297
+RP+ + + + C NCG++GH + CP + CFRCG++GH S C N
Sbjct: 8 KRPRTESSTS---CRNCGKEGHYARECPEADSKGDERSTTCFRCGEEGHMSRE---CPNE 61
Query: 298 QHEGRA 303
G A
Sbjct: 62 ARSGAA 67
Score = 37.7 bits (86), Expect = 0.038
Identities = 21/76 (27%), Positives = 31/76 (40%), Gaps = 8/76 (10%)
Query: 221 KGQQSRPKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACP-------- 272
+G SR P S +G R + + C+ CG+ GH S CP
Sbjct: 105 EGHLSRDCPSSQGGSRGGYGQKRGRSGAQGGYSGDRTCYKCGDAGHISRDCPNGQGGYSG 164
Query: 273 SEIKRCFRCGKKGHKS 288
+ + C++CG GH S
Sbjct: 165 AGDRTCYKCGDAGHIS 180
Score = 37.4 bits (85), Expect = 0.050
Identities = 14/39 (35%), Positives = 22/39 (55%), Gaps = 8/39 (20%)
Query: 258 CFNCGEKGHKSNACP--------SEIKRCFRCGKKGHKS 288
C+ CG+ GH S CP + ++C++CG+ GH S
Sbjct: 170 CYKCGDAGHISRDCPNGQGGYSGAGDRKCYKCGESGHMS 208
Score = 33.9 bits (76), Expect = 0.56
Identities = 14/42 (33%), Positives = 20/42 (47%), Gaps = 11/42 (26%)
Query: 258 CFNCGEKGHKSNACPSE-----------IKRCFRCGKKGHKS 288
C+ CG+ GH S CP + C++CG+ GH S
Sbjct: 224 CYKCGKPGHISRECPEAGGSYGGSRGGGDRTCYKCGEAGHIS 265
>GAG_SIVAI (Q02843) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 513
Score = 45.8 bits (107), Expect = 1e-04
Identities = 21/42 (50%), Positives = 26/42 (61%), Gaps = 3/42 (7%)
Query: 247 PKKKDALAEIICFNCGEKGHKSNAC--PSEIKRCFRCGKKGH 286
P+KK + CFNCG+ GH C P +IK CF+CGK GH
Sbjct: 381 PQKKGPRGPLKCFNCGKFGHMQRECKAPRQIK-CFKCGKIGH 421
>GIS2_YEAST (P53849) Zinc-finger protein GIS2
Length = 153
Score = 44.3 bits (103), Expect = 4e-04
Identities = 22/71 (30%), Positives = 32/71 (44%), Gaps = 6/71 (8%)
Query: 250 KDALAEIICFNCGEKGHKSNACPS----EIKRCFRCGKKGHKSNAEII--CFNIQHEGRA 303
+D +E +C+NC + GH C E K+C+ CG+ GH + + CFN G
Sbjct: 17 EDCDSERLCYNCNKPGHVQTDCTMPRTVEFKQCYNCGETGHVRSECTVQRCFNCNQTGHI 76
Query: 304 SHRGRSPANRS 314
S P S
Sbjct: 77 SRECPEPKKTS 87
Score = 41.2 bits (95), Expect = 0.003
Identities = 19/65 (29%), Positives = 30/65 (45%), Gaps = 7/65 (10%)
Query: 247 PKKKDALAEIICFNCGEKGHKSNACPSEI----KRCFRCGKKGHKS---NAEIICFNIQH 299
PKK +++ C+ CG H + C E +C+ CG+ GH S + +C+N
Sbjct: 83 PKKTSRFSKVSCYKCGGPNHMAKDCMKEDGISGLKCYTCGQAGHMSRDCQNDRLCYNCNE 142
Query: 300 EGRAS 304
G S
Sbjct: 143 TGHIS 147
Score = 34.7 bits (78), Expect = 0.33
Identities = 20/70 (28%), Positives = 28/70 (39%), Gaps = 18/70 (25%)
Query: 258 CFNCGEKGHKSNACPSEIK-------RCFRCGKKGHKSN--------AEIICFNIQHEGR 302
CFNC + GH S CP K C++CG H + + + C+ G+
Sbjct: 67 CFNCNQTGHISRECPEPKKTSRFSKVSCYKCGGPNHMAKDCMKEDGISGLKCYTC---GQ 123
Query: 303 ASHRGRSPAN 312
A H R N
Sbjct: 124 AGHMSRDCQN 133
Score = 34.7 bits (78), Expect = 0.33
Identities = 13/34 (38%), Positives = 21/34 (61%), Gaps = 1/34 (2%)
Query: 253 LAEIICFNCGEKGHKSNACPSEIKRCFRCGKKGH 286
+++ C+ CG+ GH + C SE + C+ C K GH
Sbjct: 1 MSQKACYVCGKIGHLAEDCDSE-RLCYNCNKPGH 33
>GAG_SIVAT (P05892) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 519
Score = 43.9 bits (102), Expect = 5e-04
Identities = 20/49 (40%), Positives = 28/49 (56%), Gaps = 3/49 (6%)
Query: 239 QRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEIK-RCFRCGKKGH 286
Q MV+ PK++ + C+NCG+ GH CP K +C +CGK GH
Sbjct: 382 QNMVQQGGPKRQRP--PLRCYNCGKFGHMQRQCPEPRKTKCLKCGKLGH 428
>GAG_SIVSP (P19504) Gag polyprotein [Contains: Core protein p17;
Core protein p24]
Length = 507
Score = 42.7 bits (99), Expect = 0.001
Identities = 18/61 (29%), Positives = 35/61 (56%), Gaps = 15/61 (24%)
Query: 227 PKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEIKR-CFRCGKKG 285
P P++A KG++++++ C+NCG++GH + C + ++ C++CGK G
Sbjct: 377 PLPFAAVQQKGQRKIIK--------------CWNCGKEGHSARQCRAPRRQGCWKCGKAG 422
Query: 286 H 286
H
Sbjct: 423 H 423
>GAG_HV1MN (P05888) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 506
Score = 42.7 bits (99), Expect = 0.001
Identities = 17/33 (51%), Positives = 25/33 (75%), Gaps = 1/33 (3%)
Query: 256 IICFNCGEKGHKSNACPSEIKR-CFRCGKKGHK 287
I CFNCG++GH + C + KR C++CGK+GH+
Sbjct: 392 IKCFNCGKEGHIAKNCRAPRKRGCWKCGKEGHQ 424
>GAG_HV1EL (P04592) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 499
Score = 42.7 bits (99), Expect = 0.001
Identities = 17/33 (51%), Positives = 25/33 (75%), Gaps = 1/33 (3%)
Query: 256 IICFNCGEKGHKSNACPSEIKR-CFRCGKKGHK 287
I CFNCG++GH + C + K+ C+RCGK+GH+
Sbjct: 390 IKCFNCGKEGHIAKNCRAPRKKGCWRCGKEGHQ 422
>GAG_FIVPE (P16087) Gag polyprotein [Contains: Core protein p15;
Major core protein p24; Nucleic acid binding protein
p10]
Length = 450
Score = 42.7 bits (99), Expect = 0.001
Identities = 22/62 (35%), Positives = 34/62 (54%), Gaps = 8/62 (12%)
Query: 257 ICFNCGEKGHKSNACPSEIKRCFRCGKKGHKSNAEIICFNIQHEGRASHRGRSPANRSSA 316
+CFNC + GH + C E+K+C +CGK GH + C+ +G + G A R++A
Sbjct: 376 VCFNCKKPGHLARQC-REVKKCNKCGKPGHLAAK---CW----QGNRKNSGNWKAGRAAA 427
Query: 317 EV 318
V
Sbjct: 428 PV 429
>YL92_SCHPO (Q9HFF2) Hypothetical protein C683.02c in chromosome I
Length = 218
Score = 42.4 bits (98), Expect = 0.002
Identities = 30/93 (32%), Positives = 45/93 (48%), Gaps = 9/93 (9%)
Query: 200 SCRIYEDDTKAHYKIVNERKTKGQQSRPKPYSAPADKGKQRMVEDRRPKKKDALAEIICF 259
S + Y++ K + Q++R K A +G +V+D P+ KD ++ ICF
Sbjct: 49 SSKRYDERQKKKRSEYRRLRRINQRNRDKFCFACRQQG--HIVQDC-PEAKDNVS--ICF 103
Query: 260 NCGEKGHKSNAC----PSEIKRCFRCGKKGHKS 288
CG K H NAC P + +CF C + GH S
Sbjct: 104 RCGSKEHSLNACSKKGPLKFAKCFICHENGHLS 136
Score = 40.0 bits (92), Expect = 0.008
Identities = 21/70 (30%), Positives = 30/70 (42%), Gaps = 13/70 (18%)
Query: 234 ADKGKQRMVEDRRPKKKDALAEI----------ICFNCGEKGHKSNACP---SEIKRCFR 280
A G + ++R+ KK+ + CF C ++GH CP + CFR
Sbjct: 45 ASFGSSKRYDERQKKKRSEYRRLRRINQRNRDKFCFACRQQGHIVQDCPEAKDNVSICFR 104
Query: 281 CGKKGHKSNA 290
CG K H NA
Sbjct: 105 CGSKEHSLNA 114
>GAG_HV1ND (P18800) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 496
Score = 42.4 bits (98), Expect = 0.002
Identities = 19/52 (36%), Positives = 30/52 (57%), Gaps = 1/52 (1%)
Query: 237 GKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEIKR-CFRCGKKGHK 287
G V +R K I CFNCG++GH + C + K+ C++CG++GH+
Sbjct: 368 GSATAVMMQRGNFKGPRKSIKCFNCGKEGHTAKNCRAPRKKGCWKCGREGHQ 419
>CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 42.4 bits (98), Expect = 0.002
Identities = 23/81 (28%), Positives = 34/81 (41%), Gaps = 11/81 (13%)
Query: 236 KGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEIKRCFRCGKKGH--------K 287
+G+ DR + + IC+ CGE GH + C + C+ CG+ GH K
Sbjct: 32 RGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPK 91
Query: 288 SNAEIICFNIQHEGRASHRGR 308
E C+N G+ H R
Sbjct: 92 REREQCCYNC---GKPGHLAR 109
Score = 38.9 bits (89), Expect = 0.017
Identities = 18/63 (28%), Positives = 34/63 (53%), Gaps = 6/63 (9%)
Query: 245 RRPKKKDALAEIICFNCGEKGHKSNACP-SEIKRCFRCGKKGH--KSNAEIICFNIQHEG 301
+ PK++ E C+NCG+ GH + C ++ ++C+ CG+ GH K ++ C+ G
Sbjct: 88 KEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETG 144
Query: 302 RAS 304
+
Sbjct: 145 HVA 147
Score = 36.2 bits (82), Expect = 0.11
Identities = 15/34 (44%), Positives = 22/34 (64%), Gaps = 3/34 (8%)
Query: 255 EIICFNCGEKGHKSNAC--PSEIKRCFRCGKKGH 286
++ C+ CGE GH + C SE+ C+RCG+ GH
Sbjct: 134 KVKCYRCGETGHVAINCSKTSEV-NCYRCGESGH 166
>CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 42.4 bits (98), Expect = 0.002
Identities = 23/81 (28%), Positives = 34/81 (41%), Gaps = 11/81 (13%)
Query: 236 KGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEIKRCFRCGKKGH--------K 287
+G+ DR + + IC+ CGE GH + C + C+ CG+ GH K
Sbjct: 32 RGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPK 91
Query: 288 SNAEIICFNIQHEGRASHRGR 308
E C+N G+ H R
Sbjct: 92 REREQCCYNC---GKPGHLAR 109
Score = 38.9 bits (89), Expect = 0.017
Identities = 18/63 (28%), Positives = 34/63 (53%), Gaps = 6/63 (9%)
Query: 245 RRPKKKDALAEIICFNCGEKGHKSNACP-SEIKRCFRCGKKGH--KSNAEIICFNIQHEG 301
+ PK++ E C+NCG+ GH + C ++ ++C+ CG+ GH K ++ C+ G
Sbjct: 88 KEPKRE---REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETG 144
Query: 302 RAS 304
+
Sbjct: 145 HVA 147
Score = 36.2 bits (82), Expect = 0.11
Identities = 15/34 (44%), Positives = 22/34 (64%), Gaps = 3/34 (8%)
Query: 255 EIICFNCGEKGHKSNAC--PSEIKRCFRCGKKGH 286
++ C+ CGE GH + C SE+ C+RCG+ GH
Sbjct: 134 KVKCYRCGETGHVAINCSKTSEV-NCYRCGESGH 166
>ZCH3_HUMAN (Q9NUD5) Zinc finger CCHC domain containing protein 3
Length = 404
Score = 42.0 bits (97), Expect = 0.002
Identities = 16/31 (51%), Positives = 21/31 (67%), Gaps = 2/31 (6%)
Query: 258 CFNCGEKGHKSNACPSEIKRCFRCGKKGHKS 288
CF CG + H S +C + RCFRCG++GH S
Sbjct: 336 CFKCGSRTHMSGSCTQD--RCFRCGEEGHLS 364
Score = 36.6 bits (83), Expect = 0.086
Identities = 16/29 (55%), Positives = 18/29 (61%), Gaps = 1/29 (3%)
Query: 258 CFNCGEKGHKSNACPSEIKRCFRCGKKGH 286
CF CGE+GH S C I C CGK+GH
Sbjct: 354 CFRCGEEGHLSPYCRKGIV-CNLCGKRGH 381
>GAG_SIVCZ (P17282) Gag polyprotein [Contains: Core protein p18;
Core protein p25; Core protein p16]
Length = 508
Score = 42.0 bits (97), Expect = 0.002
Identities = 19/42 (45%), Positives = 28/42 (66%), Gaps = 6/42 (14%)
Query: 247 PKKKDALAEIICFNCGEKGHKSNACPS-EIKRCFRCGKKGHK 287
PK+K I CFNCG++GH + C + K C+RCG++GH+
Sbjct: 396 PKRK-----IKCFNCGKEGHLARNCKAPRRKGCWRCGQEGHQ 432
>GAG_SIVAG (P27978) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 521
Score = 42.0 bits (97), Expect = 0.002
Identities = 16/30 (53%), Positives = 19/30 (63%), Gaps = 1/30 (3%)
Query: 258 CFNCGEKGHKSNACPSEIK-RCFRCGKKGH 286
C+NCG+ GH CP K RC +CGK GH
Sbjct: 404 CYNCGKFGHMQRQCPEPRKMRCLKCGKPGH 433
>GAG_HV1U4 (P24736) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 492
Score = 42.0 bits (97), Expect = 0.002
Identities = 16/33 (48%), Positives = 25/33 (75%), Gaps = 1/33 (3%)
Query: 256 IICFNCGEKGHKSNACPSEIKR-CFRCGKKGHK 287
I CFNCG++GH + C + K+ C++CGK+GH+
Sbjct: 383 IKCFNCGKEGHLAKNCRAPRKKGCWKCGKEGHQ 415
>GAG_HV1RH (P05890) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 500
Score = 42.0 bits (97), Expect = 0.002
Identities = 26/81 (32%), Positives = 39/81 (48%), Gaps = 16/81 (19%)
Query: 250 KDALAEIICFNCGEKGHKSNACPSEIKR-CFRCGKKGHKSNAEIICFNIQHEGR------ 302
+D + CFNCG+ GH + C + K+ C++CGK+GH+ + +EGR
Sbjct: 383 RDQRKIVKCFNCGKVGHIAKNCRAPRKKGCWKCGKEGHQMK------DCTNEGRQANFLG 436
Query: 303 ---ASHRGRSPANRSSAEVPT 320
SH+GR S PT
Sbjct: 437 KIWPSHKGRPGNFLQSRPEPT 457
>GAG_HV1PV (P03350) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 511
Score = 42.0 bits (97), Expect = 0.002
Identities = 15/31 (48%), Positives = 24/31 (77%), Gaps = 1/31 (3%)
Query: 258 CFNCGEKGHKSNACPSEIKR-CFRCGKKGHK 287
CFNCG++GH + C + K+ C++CGK+GH+
Sbjct: 391 CFNCGKEGHTARNCRAPRKKGCWKCGKEGHQ 421
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.133 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,636,941
Number of Sequences: 164201
Number of extensions: 1593677
Number of successful extensions: 4719
Number of sequences better than 10.0: 99
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 83
Number of HSP's that attempted gapping in prelim test: 4413
Number of HSP's gapped (non-prelim): 271
length of query: 320
length of database: 59,974,054
effective HSP length: 110
effective length of query: 210
effective length of database: 41,911,944
effective search space: 8801508240
effective search space used: 8801508240
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC130806.3


