

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130804.6 + phase: 0 /pseudo
(99 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
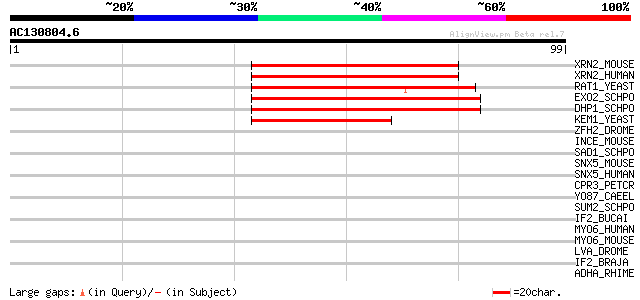
XRN2_MOUSE (Q9DBR1) 5'-3' exoribonuclease 2 (EC 3.1.11.-) (Dhm1 ... 48 5e-06
XRN2_HUMAN (Q9H0D6) 5'-3' exoribonuclease 2 (EC 3.1.11.-) 48 5e-06
RAT1_YEAST (Q02792) Ribonucleic acid trafficking protein 1 (5'-3... 48 5e-06
EXO2_SCHPO (P40383) Exonuclease II (EC 3.1.11.-) (Exo II) (P140) 46 1e-05
DHP1_SCHPO (P40848) Protein dhp1 44 5e-05
KEM1_YEAST (P22147) Strand exchange protein 1 (KAR(-) enhancing ... 44 9e-05
ZFH2_DROME (P28167) Zinc finger protein 2 (Zinc finger homeodoma... 32 0.21
INCE_MOUSE (Q9WU62) Inner centromere protein 31 0.61
SAD1_SCHPO (Q09825) Spindle pole body associated protein sad1 30 1.0
SNX5_MOUSE (Q9D8U8) Sorting nexin 5 30 1.4
SNX5_HUMAN (Q9Y5X3) Sorting nexin 5 30 1.4
CPR3_PETCR (Q99091) Light-inducible protein CPRF-3 29 1.8
YO87_CAEEL (P34623) Hypothetical protein ZK1236.7 in chromosome III 29 2.3
SUM2_SCHPO (Q9HGL3) Sum2 protein 29 2.3
IF2_BUCAI (P57458) Translation initiation factor IF-2 29 2.3
MYO6_HUMAN (Q9UM54) Myosin VI 28 3.0
MYO6_MOUSE (Q64331) Myosin VI 28 4.0
LVA_DROME (Q8MSS1) Lava lamp protein 28 4.0
IF2_BRAJA (Q89WA9) Translation initiation factor IF-2 28 4.0
ADHA_RHIME (O31186) Alcohol dehydrogenase (EC 1.1.1.1) 28 4.0
>XRN2_MOUSE (Q9DBR1) 5'-3' exoribonuclease 2 (EC 3.1.11.-) (Dhm1
protein)
Length = 951
Score = 47.8 bits (112), Expect = 5e-06
Identities = 21/37 (56%), Positives = 29/37 (77%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKE 80
IDGV PRAKMN+QR+RRFR +K+ E++R+R+E
Sbjct: 102 IDGVAPRAKMNQQRSRRFRASKEGMEAAVEKQRVREE 138
>XRN2_HUMAN (Q9H0D6) 5'-3' exoribonuclease 2 (EC 3.1.11.-)
Length = 950
Score = 47.8 bits (112), Expect = 5e-06
Identities = 21/37 (56%), Positives = 29/37 (77%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKE 80
IDGV PRAKMN+QR+RRFR +K+ E++R+R+E
Sbjct: 102 IDGVAPRAKMNQQRSRRFRASKEGMEAAVEKQRVREE 138
>RAT1_YEAST (Q02792) Ribonucleic acid trafficking protein 1 (5'-3'
exoribonuclease) (EC 3.1.11.-) (P116)
Length = 1006
Score = 47.8 bits (112), Expect = 5e-06
Identities = 23/41 (56%), Positives = 32/41 (77%), Gaps = 1/41 (2%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEM-REAEEERLRKEFEM 83
+DGV PRAKMN+QR RRFR+A+D ++ EA EE +R+ E+
Sbjct: 101 VDGVAPRAKMNQQRARRFRSARDAQIENEAREEIMRQREEV 141
>EXO2_SCHPO (P40383) Exonuclease II (EC 3.1.11.-) (Exo II) (P140)
Length = 1328
Score = 46.2 bits (108), Expect = 1e-05
Identities = 22/41 (53%), Positives = 28/41 (67%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEME 84
+DG PRAKMN+QR+RRFRTAKD + ER ++F E
Sbjct: 85 VDGCAPRAKMNQQRSRRFRTAKDAHDARLKAERNGEDFPEE 125
>DHP1_SCHPO (P40848) Protein dhp1
Length = 991
Score = 44.3 bits (103), Expect = 5e-05
Identities = 19/41 (46%), Positives = 31/41 (75%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEME 84
IDGV PRAKMN+QR+RRFR++++ ++E E + +E + +
Sbjct: 103 IDGVAPRAKMNQQRSRRFRSSREAALKEEELQAFIEEAKQQ 143
>KEM1_YEAST (P22147) Strand exchange protein 1 (KAR(-) enhancing
mutation protein) (5'-3' exoribonuclease) (DNA strand
transfer protein beta) (STP-beta) (P175)
Length = 1528
Score = 43.5 bits (101), Expect = 9e-05
Identities = 20/25 (80%), Positives = 21/25 (84%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNE 68
IDGV PRAKMN+QR RRFRTA D E
Sbjct: 85 IDGVAPRAKMNQQRARRFRTAMDAE 109
>ZFH2_DROME (P28167) Zinc finger protein 2 (Zinc finger homeodomain
protein 2)
Length = 3005
Score = 32.3 bits (72), Expect = 0.21
Identities = 20/73 (27%), Positives = 35/73 (47%), Gaps = 8/73 (10%)
Query: 34 LIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEM-------REAEEERLRKEFEMEAN 86
L+A P +PD+ G P ++ K + + + RE ++E+LR + E N
Sbjct: 2573 LVASPQIIPDVPGYKPSLRIPKSTSDEAPYIGETSLEQATELQREKQDEQLRIDRPSEEN 2632
Query: 87 KFFLNKNVKCQNL 99
+NKN K +N+
Sbjct: 2633 DLSMNKN-KVENI 2644
>INCE_MOUSE (Q9WU62) Inner centromere protein
Length = 880
Score = 30.8 bits (68), Expect = 0.61
Identities = 14/35 (40%), Positives = 23/35 (65%)
Query: 50 RAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEME 84
R + +++ RR + K+ E RE E+ERLR + EM+
Sbjct: 672 REQERREQERREQERKEQERREQEQERLRAKREMQ 706
>SAD1_SCHPO (Q09825) Spindle pole body associated protein sad1
Length = 514
Score = 30.0 bits (66), Expect = 1.0
Identities = 21/77 (27%), Positives = 33/77 (42%), Gaps = 8/77 (10%)
Query: 6 SNKFACHLLANTLKQKIHETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAK 65
+N H L K+H+ + K I PPT I+ +TP+ + F A
Sbjct: 26 ANSAQIHQNTADLASKMHKLRYTK-----IRSPPTRVSIESITPKQRF---PAPNFEQAY 77
Query: 66 DNEMREAEEERLRKEFE 82
+ +R +EE +EFE
Sbjct: 78 HSNIRYEQEESDNEEFE 94
>SNX5_MOUSE (Q9D8U8) Sorting nexin 5
Length = 404
Score = 29.6 bits (65), Expect = 1.4
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 10/65 (15%)
Query: 22 IHETLH--AKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRK 79
+H+TL + +I P PT PD DG PR KM K + M + E ++++
Sbjct: 75 LHDTLTETTDYAGLIIPPAPTKPDFDG--PREKMQK------LGEGEGSMTKEEFAKMKQ 126
Query: 80 EFEME 84
E E E
Sbjct: 127 ELEAE 131
>SNX5_HUMAN (Q9Y5X3) Sorting nexin 5
Length = 404
Score = 29.6 bits (65), Expect = 1.4
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 10/65 (15%)
Query: 22 IHETL--HAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRK 79
+H+TL + +I P PT PD DG PR KM K + M + E ++++
Sbjct: 75 LHDTLIETTDYAGLIIPPAPTKPDFDG--PREKMQK------LGEGEGSMTKEEFAKMKQ 126
Query: 80 EFEME 84
E E E
Sbjct: 127 ELEAE 131
>CPR3_PETCR (Q99091) Light-inducible protein CPRF-3
Length = 296
Score = 29.3 bits (64), Expect = 1.8
Identities = 15/56 (26%), Positives = 27/56 (47%)
Query: 25 TLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKE 80
+L + N +A P + DG+ P ++N +R + + K + A RLRK+
Sbjct: 163 SLEVRSNPLDVAAPGAIVVHDGMLPDQRVNDERELKRQRRKQSNRESARRSRLRKQ 218
>YO87_CAEEL (P34623) Hypothetical protein ZK1236.7 in chromosome III
Length = 324
Score = 28.9 bits (63), Expect = 2.3
Identities = 15/38 (39%), Positives = 22/38 (57%)
Query: 50 RAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
R + +++ R AK+ +AEEERLRKE E + K
Sbjct: 161 RREEEREKKRDEERAKEEADEKAEEERLRKEREEKERK 198
>SUM2_SCHPO (Q9HGL3) Sum2 protein
Length = 426
Score = 28.9 bits (63), Expect = 2.3
Identities = 17/62 (27%), Positives = 31/62 (49%), Gaps = 4/62 (6%)
Query: 38 PPTLPDIDGVTPRAKMNKQ-RTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLNKNVKC 96
P T PD PR + + Q ++F++ KD+ ++ +E +EF FF N+ C
Sbjct: 293 PTTQPDASAAKPRTEFDFQTANQKFQSMKDDLLKGKNDEE-AEEFYKPKQSFF--DNISC 349
Query: 97 QN 98
++
Sbjct: 350 ES 351
>IF2_BUCAI (P57458) Translation initiation factor IF-2
Length = 864
Score = 28.9 bits (63), Expect = 2.3
Identities = 13/44 (29%), Positives = 26/44 (58%), Gaps = 2/44 (4%)
Query: 50 RAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEM--EANKFFLN 91
+ ++NK +R T K+N+++ + ++ K+F E+N F LN
Sbjct: 133 KIELNKSDSRSLNTKKENKLKISNKDEQNKKFNQHRESNSFDLN 176
>MYO6_HUMAN (Q9UM54) Myosin VI
Length = 1285
Score = 28.5 bits (62), Expect = 3.0
Identities = 15/36 (41%), Positives = 21/36 (57%), Gaps = 3/36 (8%)
Query: 52 KMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
+M K+R RR +D + R EEE R + EMEA +
Sbjct: 932 EMEKERKRR---EEDEKRRRKEEEERRMKLEMEAKR 964
>MYO6_MOUSE (Q64331) Myosin VI
Length = 1265
Score = 28.1 bits (61), Expect = 4.0
Identities = 15/36 (41%), Positives = 20/36 (54%), Gaps = 3/36 (8%)
Query: 52 KMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
+M K+R RR +D E R EEE R + EME +
Sbjct: 935 EMEKERKRR---EEDEERRRKEEEERRMKLEMEPKR 967
>LVA_DROME (Q8MSS1) Lava lamp protein
Length = 2779
Score = 28.1 bits (61), Expect = 4.0
Identities = 14/46 (30%), Positives = 23/46 (49%)
Query: 35 IAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKE 80
I+PPPT+ D+ + + Q A++ + E EE RL+ E
Sbjct: 1780 ISPPPTIEDLQRNVSDLEKHAQDLETKLLARNQNLAEQEERRLQLE 1825
>IF2_BRAJA (Q89WA9) Translation initiation factor IF-2
Length = 902
Score = 28.1 bits (61), Expect = 4.0
Identities = 17/47 (36%), Positives = 22/47 (46%), Gaps = 2/47 (4%)
Query: 36 APPPTLPDI--DGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKE 80
APPP P GV R +R+ R D ++RE EE R +E
Sbjct: 78 APPPAPPRNAGSGVVLRTLTEDERSARASALADAKLREVEERRQAEE 124
>ADHA_RHIME (O31186) Alcohol dehydrogenase (EC 1.1.1.1)
Length = 340
Score = 28.1 bits (61), Expect = 4.0
Identities = 15/38 (39%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Query: 23 HETLHAKFNRWLIAP-PPTLPDIDGVTPRAKMNKQRTR 59
H LHA W + P PP +P +GV AK+ + TR
Sbjct: 41 HTDLHAAKGDWPVRPNPPFIPGHEGVGYVAKLGAEVTR 78
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,630,266
Number of Sequences: 164201
Number of extensions: 414528
Number of successful extensions: 1793
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1748
Number of HSP's gapped (non-prelim): 63
length of query: 99
length of database: 59,974,054
effective HSP length: 75
effective length of query: 24
effective length of database: 47,658,979
effective search space: 1143815496
effective search space used: 1143815496
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC130804.6


