

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130802.9 + phase: 0 /pseudo
(937 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
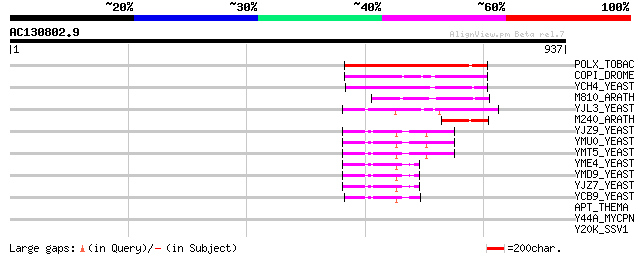
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 242 3e-63
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 160 1e-38
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 117 2e-25
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 110 2e-23
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 72 7e-12
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 55 9e-07
YJZ9_YEAST (P47100) Transposon Ty1 protein B 51 2e-05
YMU0_YEAST (Q04670) Transposon Ty1 protein B 50 3e-05
YMT5_YEAST (Q04214) Transposon Ty1 protein B 50 4e-05
YME4_YEAST (Q04711) Transposon Ty1 protein B 49 5e-05
YMD9_YEAST (Q03434) Transposon Ty1 protein B 49 5e-05
YJZ7_YEAST (P47098) Transposon Ty1 protein B 47 3e-04
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 47 3e-04
APT_THEMA (Q9X1A4) Adenine phosphoribosyltransferase (EC 2.4.2.7... 33 2.7
Y44A_MYCPN (P75151) Hypothetical lipoprotein MG440 homolog 1 pre... 33 4.6
Y20K_SSV1 (P20216) Hypothetical 20.4 kDa protein (ORF E-178) 32 6.0
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 242 bits (618), Expect = 3e-63
Identities = 119/242 (49%), Positives = 165/242 (68%), Gaps = 3/242 (1%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
+SLYGLKQ+PR+WY +FD F+ ++++ D CVY + +E + LLLYVDD+L+
Sbjct: 958 KSLYGLKQAPRQWYMKFDSFMKSQTYLKTYSDPCVYFKRFSENNFIILLLYVDDMLIVGK 1017
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
K I KLK L+ F+MKDLGPA+++LG+ I R R +L+LSQ Y+++ ERF M N
Sbjct: 1018 DKGLIAKLKGDLSKSFDMKDLGPAQQILGMKIVRERTSRKLWLSQEKYIERVLERFNMKN 1077
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
+K VSTPL H KLS + CP + +EK M PY+S VGS+MY MVC+RPD+A+AV +VS
Sbjct: 1078 AKPVSTPLAGHLKLSKKMCPTTVEEKGNMAKVPYSSAVGSLMYAMVCTRPDIAHAVGVVS 1137
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
RF+ NPG HW+A+KW+LRYL G+ L + + L+GY DAD AG++D RKS
Sbjct: 1138 RFLENPGKEHWEAVKWILRYLRGTTGDCLCF---GGSDPILKGYTDADMAGDIDNRKSST 1194
Query: 806 GF 807
G+
Sbjct: 1195 GY 1196
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 160 bits (406), Expect = 1e-38
Identities = 95/243 (39%), Positives = 141/243 (57%), Gaps = 10/243 (4%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKK-NEKVILYLLLYVDDILMAS 624
+++YGLKQ+ R W+ F++ L + FV S D C+Y+L K N +Y+LLYVDD+++A+
Sbjct: 1036 KAIYGLKQAARCWFEVFEQALKECEFVNSSVDRCIYILDKGNINENIYVLLYVDDVVIAT 1095
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
+ K L +F M DL K +GI I+ DK ++LSQ Y+KK +F M
Sbjct: 1096 GDMTRMNNFKRYLMEKFRMTDLNEIKHFIGIRIEMQEDK--IYLSQSAYVKKILSKFNME 1153
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
N VSTPL +K++ + ED TP S +G +MY M+C+RPDL AV+I+
Sbjct: 1154 NCNAVSTPLP--SKINYELLNSDEDCN-----TPCRSLIGCLMYIMLCTRPDLTTAVNIL 1206
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+ + WQ LK VLRYL G++ L + + E+ + GYVD+D+AG+ RKS
Sbjct: 1207 SRYSSKNNSELWQNLKRVLRYLKGTIDMKLIFKKNLAFENKIIGYVDSDWAGSEIDRKST 1266
Query: 805 VGF 807
G+
Sbjct: 1267 TGY 1269
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 117 bits (292), Expect = 2e-25
Identities = 78/240 (32%), Positives = 123/240 (50%), Gaps = 10/240 (4%)
Query: 568 LYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASSSK 627
+YGLKQ+P W + L K GF R + +Y ++ I Y+ +YVDD+L+A+ S
Sbjct: 40 MYGLKQAPLLWNEHINNTLKKIGFCRHEGEHGLYFRSTSDGPI-YIGVYVDDLLVAAPSP 98
Query: 628 DEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSNSK 687
++K+ L + MKDLG + LG++I ++ + G++ LS Y+ K ++ K
Sbjct: 99 KIYDRVKQELTKLYSMKDLGKVDKFLGLNIHQSTN-GDITLSLQDYIAKAASESEINTFK 157
Query: 688 TVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVSRF 747
TPL + L P +D TPY S VG +++ RPD++Y VS++SRF
Sbjct: 158 LTQTPLCNSKPLFETTSPHLKDI------TPYQSIVGQLLFCANTGRPDISYPVSLLSRF 211
Query: 748 MANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLVGF 807
+ P +H ++ + VLRYL + LKY +Q AL Y DA + D S G+
Sbjct: 212 LREPRAIHLESARRVLRYLYTTRSMCLKYRSGSQ--VALTVYCDASHGAIHDLPHSTGGY 269
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 110 bits (274), Expect = 2e-23
Identities = 74/202 (36%), Positives = 112/202 (54%), Gaps = 16/202 (7%)
Query: 611 LYLLLYVDDILMASSSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQ 670
+YLLLYVDDIL+ SS + L +L+ F MKDLGP LGI IK + LFLSQ
Sbjct: 1 MYLLLYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIKTH--PSGLFLSQ 58
Query: 671 LGYLKKGGERFRMSNSKTVST--PLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMY 728
Y ++ M + K +ST PL ++ +S + P D + S VG++ Y
Sbjct: 59 TKYAEQILNNAGMLDCKPMSTPLPLKLNSSVSTAKYPDPSD---------FRSIVGALQY 109
Query: 729 GMVCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEG 788
+ +RPD++YAV+IV + M P + + LK VLRY+ G++ GL + ++ ++
Sbjct: 110 -LTLTRPDISYAVNIVCQRMHEPTLADFDLLKRVLRYVKGTIFHGLYIHKNSKLN--VQA 166
Query: 789 YVDADYAGNVDTRKSLVGFCVY 810
+ D+D+AG TR+S GFC +
Sbjct: 167 FCDSDWAGCTSTRRSTTGFCTF 188
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 72.0 bits (175), Expect = 7e-12
Identities = 64/277 (23%), Positives = 119/277 (42%), Gaps = 20/277 (7%)
Query: 562 VSFEESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDIL 621
V ++LYGLKQSP+EW ++L G + Y +Y + E L + +YVDD +
Sbjct: 1410 VKLNKALYGLKQSPKEWNDHLRQYLNGIGLKDNSYTPGLY---QTEDKNLMIAVYVDDCV 1466
Query: 622 MASSSKDEIMKLKERLNGEFEMKDLGPA------KRVLGIDIKRNRDKGELFLSQLGYLK 675
+A+S++ + + +L FE+K G +LG+D+ N+ G + L+ ++
Sbjct: 1467 IAASNEQRLDEFINKLKSNFELKITGTLIDDVLDTDILGMDLVYNKRLGTIDLTLKSFIN 1526
Query: 676 KGGERFRMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGV-------GSIMY 728
+ +++ K + + H +S + +D Q+ E + GV G + Y
Sbjct: 1527 RMDKKYNEELKKIRKSSIPH---MSTYKIDPKKDVLQMSE-EEFRQGVLKLQQLLGELNY 1582
Query: 729 GMVCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEG 788
R D+ +AV V+R + P + + +++YL G+ Y R + +
Sbjct: 1583 VRHKCRYDIEFAVKKVARLVNYPHERVFYMIYKIIQYLVRYKDIGIHYDRDCNKDKKVIA 1642
Query: 789 YVDADYAGNVDTRKSLVGFCVYSL*HDD*LEGKSTIR 825
DA D + + Y + + KST R
Sbjct: 1643 ITDASVGSEYDAQSRIGVILWYGMNIFNVYSNKSTNR 1679
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 55.1 bits (131), Expect = 9e-07
Identities = 29/79 (36%), Positives = 48/79 (60%), Gaps = 2/79 (2%)
Query: 730 MVCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGY 789
+ +RPDL +AV+ +S+F + QA+ VL Y+ G++ GL Y +A + L+ +
Sbjct: 3 LTITRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFY--SATSDLQLKAF 60
Query: 790 VDADYAGNVDTRKSLVGFC 808
D+D+A DTR+S+ GFC
Sbjct: 61 ADSDWASCPDTRRSVTGFC 79
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 50.8 bits (120), Expect = 2e-05
Identities = 54/208 (25%), Positives = 94/208 (44%), Gaps = 32/208 (15%)
Query: 562 VSFEESLYGLKQSPREWYRRFDEFLL-KTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDI 620
+ ++SLYGLKQS WY +L+ + G SCV+ KN +V + L+VDD+
Sbjct: 1375 IRLKKSLYGLKQSGANWYETIKSYLIQQCGMEEVRGWSCVF---KNSQVT--ICLFVDDM 1429
Query: 621 LMASSSKDEIMKLKERLNGEFEMK--DLGPAKR-----VLGIDIKRNRDKGELFLSQLGY 673
++ S + + ++ E+L +++ K +LG + +LG++IK R K Y
Sbjct: 1430 VLFSKNLNSNKRIIEKLKMQYDTKIINLGESDEEIQYDILGLEIKYQRGK---------Y 1480
Query: 674 LKKGGERFRMSNSKTVSTPLG-HHTKLSI---------QQCPQSEDEKQLMEGTPYASGV 723
+K G E ++ PL KLS QQ + E++ M+ +
Sbjct: 1481 MKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKLI 1540
Query: 724 GSIMYGMVCSRPDLAYAVSIVSRFMANP 751
G Y R DL Y ++ +++ + P
Sbjct: 1541 GLASYVGYKFRFDLLYYINTLAQHILFP 1568
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 50.1 bits (118), Expect = 3e-05
Identities = 55/208 (26%), Positives = 91/208 (43%), Gaps = 32/208 (15%)
Query: 562 VSFEESLYGLKQSPREWYRRFDEFLLK-TGFVRSGYDSCVYMLKKNEKVILYLLLYVDDI 620
+ ++SLYGLKQS WY +L+K G SCV+ KN +V + L+VDD+
Sbjct: 948 IRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVF---KNSQVT--ICLFVDDM 1002
Query: 621 LMASSSKDEIMKLKERLNGEFEMK--DLGPAKR-----VLGIDIKRNRDKGELFLSQLGY 673
++ S + K+ L +++ K +LG + +LG++IK R K Y
Sbjct: 1003 ILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGK---------Y 1053
Query: 674 LKKGGERFRMSNSKTVSTPLG-HHTKLSI---------QQCPQSEDEKQLMEGTPYASGV 723
+K G E ++ PL KLS QQ + E++ M+ +
Sbjct: 1054 MKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKLI 1113
Query: 724 GSIMYGMVCSRPDLAYAVSIVSRFMANP 751
G Y R DL Y ++ +++ + P
Sbjct: 1114 GLASYVGYKFRFDLLYYINTLAQHILFP 1141
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 49.7 bits (117), Expect = 4e-05
Identities = 53/208 (25%), Positives = 94/208 (44%), Gaps = 32/208 (15%)
Query: 562 VSFEESLYGLKQSPREWYRRFDEFLLK-TGFVRSGYDSCVYMLKKNEKVILYLLLYVDDI 620
+ ++SLYGLKQS WY +L+K G SCV+ +N +V + L+VDD+
Sbjct: 948 IRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVF---ENSQVT--ICLFVDDM 1002
Query: 621 LMASSSKDEIMKLKERLNGEFEMK--DLGPAKR-----VLGIDIKRNRDKGELFLSQLGY 673
++ S + + ++ ++L +++ K +LG + +LG++IK R K Y
Sbjct: 1003 VLFSKNLNSNKRIIDKLKMQYDTKIINLGESDEEIQYDILGLEIKYQRGK---------Y 1053
Query: 674 LKKGGERFRMSNSKTVSTPLG-HHTKLSI---------QQCPQSEDEKQLMEGTPYASGV 723
+K G E ++ PL KLS QQ + E++ M+ +
Sbjct: 1054 MKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKLI 1113
Query: 724 GSIMYGMVCSRPDLAYAVSIVSRFMANP 751
G Y R DL Y ++ +++ + P
Sbjct: 1114 GLASYVGYKFRFDLLYYINTLAQHILFP 1141
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 49.3 bits (116), Expect = 5e-05
Identities = 43/139 (30%), Positives = 66/139 (46%), Gaps = 27/139 (19%)
Query: 562 VSFEESLYGLKQSPREWYRRFDEFLLK-TGFVRSGYDSCVYMLKKNEKVILYLLLYVDDI 620
+ ++SLYGLKQS WY +L+K G SCV+ KN +V + L+VDD+
Sbjct: 948 IRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVF---KNSQVT--ICLFVDDM 1002
Query: 621 LMASSSKDEIMKLKERLNGEFEMK--DLGPAKR-----VLGIDIKRNRDKGELFLSQLGY 673
++ S + K+ L +++ K +LG + +LG++IK R K Y
Sbjct: 1003 ILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGK---------Y 1053
Query: 674 LKKGGERFRMSNSKTVSTP 692
+K G M NS T P
Sbjct: 1054 MKLG-----MENSLTEKIP 1067
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 49.3 bits (116), Expect = 5e-05
Identities = 43/139 (30%), Positives = 66/139 (46%), Gaps = 27/139 (19%)
Query: 562 VSFEESLYGLKQSPREWYRRFDEFLLK-TGFVRSGYDSCVYMLKKNEKVILYLLLYVDDI 620
+ ++SLYGLKQS WY +L+K G SCV+ KN +V + L+VDD+
Sbjct: 948 IRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVF---KNSQVT--ICLFVDDM 1002
Query: 621 LMASSSKDEIMKLKERLNGEFEMK--DLGPAKR-----VLGIDIKRNRDKGELFLSQLGY 673
++ S + K+ L +++ K +LG + +LG++IK R K Y
Sbjct: 1003 ILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGK---------Y 1053
Query: 674 LKKGGERFRMSNSKTVSTP 692
+K G M NS T P
Sbjct: 1054 MKLG-----MENSLTEKIP 1067
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 46.6 bits (109), Expect = 3e-04
Identities = 42/139 (30%), Positives = 65/139 (46%), Gaps = 27/139 (19%)
Query: 562 VSFEESLYGLKQSPREWYRRFDEFLLK-TGFVRSGYDSCVYMLKKNEKVILYLLLYVDDI 620
+ ++S YGLKQS WY +L+K G SCV+ KN +V + L+VDD+
Sbjct: 1375 IRLKKSHYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVF---KNSQVT--ICLFVDDM 1429
Query: 621 LMASSSKDEIMKLKERLNGEFEMK--DLGPAKR-----VLGIDIKRNRDKGELFLSQLGY 673
++ S + K+ L +++ K +LG + +LG++IK R K Y
Sbjct: 1430 ILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGK---------Y 1480
Query: 674 LKKGGERFRMSNSKTVSTP 692
+K G M NS T P
Sbjct: 1481 MKLG-----MENSLTEKIP 1494
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 46.6 bits (109), Expect = 3e-04
Identities = 40/137 (29%), Positives = 67/137 (48%), Gaps = 24/137 (17%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRS--GYDSCVYMLKKNEKVILYLLLYVDDILMA 623
+SLYGLKQS WY +L+ ++ G+ SCV+ KN +V + L+VDD+++
Sbjct: 1394 KSLYGLKQSGANWYETIKSYLINCCDMQEVRGW-SCVF---KNSQVT--ICLFVDDMILF 1447
Query: 624 SSSKDEIMKLKERLNGEFEMK--DLGPAKR-----VLGIDIKRNRDKGELFLSQLGYLKK 676
S + K+ L +++ K +LG + +LG++IK R K Y+K
Sbjct: 1448 SKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRSK---------YMKL 1498
Query: 677 GGERFRMSNSKTVSTPL 693
G E+ ++ PL
Sbjct: 1499 GMEKSLTEKLPKLNVPL 1515
>APT_THEMA (Q9X1A4) Adenine phosphoribosyltransferase (EC 2.4.2.7)
(APRT)
Length = 173
Score = 33.5 bits (75), Expect = 2.7
Identities = 19/63 (30%), Positives = 35/63 (55%), Gaps = 12/63 (19%)
Query: 603 LKKNEKVILYLLLYVDDILMASSSKDEIMKLKERLNGE-------FEMKDLGPAKRVLGI 655
++K +KV++ VDD+L + + +++L ++L GE E+ L P KR+ G
Sbjct: 111 IEKGQKVLI-----VDDVLATGGTAEALIRLVKKLGGEVVSLAFLVELSYLEPRKRLEGY 165
Query: 656 DIK 658
D+K
Sbjct: 166 DVK 168
>Y44A_MYCPN (P75151) Hypothetical lipoprotein MG440 homolog 1
precursor (E09_orf277)
Length = 277
Score = 32.7 bits (73), Expect = 4.6
Identities = 20/107 (18%), Positives = 46/107 (42%), Gaps = 3/107 (2%)
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQ---LGYLKKGGERF 681
SS+D +++K G ++ ++ K +G+ DK E L Y F
Sbjct: 155 SSRDFTIQIKLNAKGNYKKSEVEKYKSQIGVQDNELGDKEEGTLEADLIFTYTPPASNLF 214
Query: 682 RMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMY 728
SN ++ + +T L ++ + E ++++ + + + + S+ Y
Sbjct: 215 SKSNFDKLTKSINFNTNLKLEMVGKDEVMRKILNSSAFTNNLASVNY 261
>Y20K_SSV1 (P20216) Hypothetical 20.4 kDa protein (ORF E-178)
Length = 178
Score = 32.3 bits (72), Expect = 6.0
Identities = 21/73 (28%), Positives = 33/73 (44%)
Query: 561 GVSFEESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDI 620
GV + + Y LK+ E +FDE + + R+ Y+ K + L+LY
Sbjct: 62 GVIEKTTPYSLKKEYEEQVEKFDEIINYEDYERARYELYKLSEKADNLFTSSLVLYASVF 121
Query: 621 LMASSSKDEIMKL 633
L S K EI+K+
Sbjct: 122 LSLSELKPEILKI 134
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.356 0.161 0.595
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 96,646,692
Number of Sequences: 164201
Number of extensions: 3884182
Number of successful extensions: 14381
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 14354
Number of HSP's gapped (non-prelim): 17
length of query: 937
length of database: 59,974,054
effective HSP length: 120
effective length of query: 817
effective length of database: 40,269,934
effective search space: 32900536078
effective search space used: 32900536078
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 71 (32.0 bits)
Medicago: description of AC130802.9


