

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.5 + phase: 0
(569 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
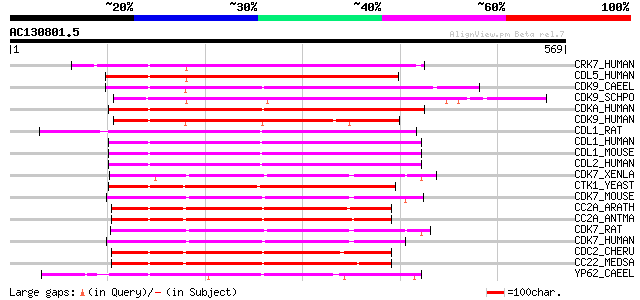
CRK7_HUMAN (Q9NYV4) Cell division cycle 2-related protein kinase... 284 4e-76
CDL5_HUMAN (Q14004) Cell division cycle 2-like protein kinase 5 ... 274 5e-73
CDK9_CAEEL (P46551) Putative cell division protein kinase 9 homo... 256 1e-67
CDK9_SCHPO (Q96WV9) Serine/threonine-protein kinase cdk9 (EC 2.7... 252 2e-66
CDKA_HUMAN (Q15131) Cell division protein kinase 10 (EC 2.7.1.37... 250 8e-66
CDK9_HUMAN (P50750) Cell division protein kinase 9 (EC 2.7.1.37)... 248 2e-65
CDL1_RAT (P46892) PITSLRE serine/threonine-protein kinase CDC2L1... 231 5e-60
CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2... 228 4e-59
CDL1_MOUSE (P24788) PITSLRE serine/threonine-protein kinase CDC2... 226 2e-58
CDL2_HUMAN (Q9UQ88) PITSLRE serine/threonine-protein kinase CDC2... 224 4e-58
CDK7_XENLA (P20911) Cell division protein kinase 7 (EC 2.7.1.37)... 222 2e-57
CTK1_YEAST (Q03957) CTD kinase alpha subunit (EC 2.7.1.-) (CTD k... 222 2e-57
CDK7_MOUSE (Q03147) Cell division protein kinase 7 (EC 2.7.1.37)... 220 7e-57
CC2A_ARATH (P24100) Cell division control protein 2 homolog A (E... 220 9e-57
CC2A_ANTMA (Q38772) Cell division control protein 2 homolog A (E... 219 2e-56
CDK7_RAT (P51952) Cell division protein kinase 7 (EC 2.7.1.37) (... 218 3e-56
CDK7_HUMAN (P50613) Cell division protein kinase 7 (EC 2.7.1.37)... 218 4e-56
CDC2_CHERU (P93101) Cell division control protein 2 homolog (EC ... 217 6e-56
CC22_MEDSA (Q05006) Cell division control protein 2 homolog 2 (E... 217 6e-56
YP62_CAEEL (Q09437) Putative serine/threonine-protein kinase B04... 216 1e-55
>CRK7_HUMAN (Q9NYV4) Cell division cycle 2-related protein kinase 7
(EC 2.7.1.37) (CDC2-related protein kinase 7) (CrkRS)
Length = 1490
Score = 284 bits (727), Expect = 4e-76
Identities = 162/373 (43%), Positives = 218/373 (58%), Gaps = 18/373 (4%)
Query: 64 PNPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYK 123
P P+ PP+ ++ P + + + W R D F+ I IG+GTY VYK
Sbjct: 686 PEPKAITPPQQPYKKRPKICCPRY--GERRQTESDWGKRCVDKFDIIGIIGEGTYGQVYK 743
Query: 124 AIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDHPNVIKLQGLVTSRMSC--- 179
A D TG++VALKKVR DN E E A REI ILR+L H +V+ ++ +VT +
Sbjct: 744 ARDKDTGELVALKKVRLDN-EKEGFPITAIREIKILRQLIHRSVVNMKEIVTDKQDALDF 802
Query: 180 -----SLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKG 234
+ YLVF+YM+HDL GL S ++ F+E IK +M QL+ GLE+CH + LHRDIK
Sbjct: 803 KKDKGAFYLVFEYMDHDLMGLLESGLVHFSEDHIKSFMKQLMEGLEYCHKKNFLHRDIKC 862
Query: 235 SNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSA 294
SN+L++N G +K+ADFGLA ++ P T++V+TLWYRPPELLLG Y ID+WS
Sbjct: 863 SNILLNNSGQIKLADFGLARLYNSEESRPYTNKVITLWYRPPELLLGEERYTPAIDVWSC 922
Query: 295 GCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKK-SKLPNATLFKPREPYKRCI 353
GCILGEL KPI E+ QL I +LCGSP W KLP KP++ Y+R +
Sbjct: 923 GCILGELFTKKPIFQANLELAQLELISRLCGSPCPAVWPDVIKLPYFNTMKPKKQYRRRL 982
Query: 354 RETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF-TTEPYACDPSSLPKYPPSKEM 412
RE F P +AL L+D +L +DP +R TA L+S+F E P LP + E+
Sbjct: 983 REEFSFIPSAALDLLDHMLTLDPSKRCTAEQTLQSDFLKDVELSKMAPPDLPHWQDCHEL 1042
Query: 413 DAKRRDDEVRRQR 425
+K+R RRQR
Sbjct: 1043 WSKKR----RRQR 1051
>CDL5_HUMAN (Q14004) Cell division cycle 2-like protein kinase 5 (EC
2.7.1.37) (Cholinesterase-related cell division
controller) (CDC2-related protein kinase 5)
Length = 418
Score = 274 bits (700), Expect = 5e-73
Identities = 148/311 (47%), Positives = 192/311 (61%), Gaps = 12/311 (3%)
Query: 99 WIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIII 157
W D F+ I IG+GTY VYKA D TG++VALKKVR DN E E A REI I
Sbjct: 83 WGKLCVDKFDIIGIIGEGTYGQVYKARDKDTGEMVALKKVRLDN-EKEGFPITAIREIKI 141
Query: 158 LRRLDHPNVIKLQGLVTSRMSC--------SLYLVFDYMEHDLAGLAASPVIRFTESQIK 209
LR+L H ++I ++ +VT + + YLVF+YM+HDL GL S ++ F E+ IK
Sbjct: 142 LRQLTHQSIINMKEIVTDKEDALDFKKDKGAFYLVFEYMDHDLMGLLESGLVHFYENHIK 201
Query: 210 CYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVV 269
+M QL+ GL++CH + LHRDIK SN+L++N G +K+ADFGLA + P T++V+
Sbjct: 202 SFMRQLMEGLDYCHKKNFLHRDIKCSNILLNNRGQIKLADFGLARLYSSEESRPYTNKVI 261
Query: 270 TLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSD 329
TLWYRPPELLLG Y ID+WS GCILGEL KPI E+ QL I ++CGSP
Sbjct: 262 TLWYRPPELLLGEERYTPAIDVWSCGCILGELFTKKPIFQANQELAQLELISRICGSPCP 321
Query: 330 EYWKK-SKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRS 388
W KLP KP++ Y+R +RE F P +AL L D +LA+DP +R TA AL+
Sbjct: 322 AVWPDVIKLPYFNTMKPKKQYRRKLREEFVFIPAAALDLFDYMLALDPSKRCTAEQALQC 381
Query: 389 EFF-TTEPYAC 398
EF EP C
Sbjct: 382 EFLRDVEPSKC 392
>CDK9_CAEEL (P46551) Putative cell division protein kinase 9 homolog
(EC 2.7.1.37)
Length = 730
Score = 256 bits (654), Expect = 1e-67
Identities = 154/396 (38%), Positives = 221/396 (54%), Gaps = 18/396 (4%)
Query: 99 WIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIII 157
W + +D+IG+GTY VYKA++++TG+ VALK+VR +N E E A REI I
Sbjct: 303 WYKTNLTHYTMLDQIGEGTYGQVYKAVNNLTGEQVALKRVRLEN-EKEGFPITAIREIKI 361
Query: 158 LRRLDHPNVIKLQGLVTSRMS--------CSLYLVFDYMEHDLAGLAASP-VIRFTESQI 208
LR+L H N+++L +V +S + YLVF+Y++HDL GL S ++ F + QI
Sbjct: 362 LRQLHHKNIVRLMDIVIDDISMDELKRTRANFYLVFEYVDHDLIGLLESKELVDFNKDQI 421
Query: 209 KCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRV 268
QLL GL + HN LHRDIK SN+L++N+G LKIAD GLA ++ T+RV
Sbjct: 422 CSLFKQLLEGLAYIHNTGFLHRDIKCSNILVNNKGELKIADLGLARLWEKE-SRLYTNRV 480
Query: 269 VTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPS 328
+TLWYRPPELLLG YG ID+WS GC+LGEL KP+ G E QL I K+CGSP+
Sbjct: 481 ITLWYRPPELLLGDERYGPAIDVWSTGCMLGELFTRKPLFNGNNEFGQLELISKVCGSPN 540
Query: 329 DEYWKK-SKLPNATLFKPREPYKRCIRETFKG-FPPSALPLIDKLLAIDPVERETASDAL 386
+ W + ++L F+ + Y+R IRE F+ P A+ L+DK+L ++P +R +A +AL
Sbjct: 541 VDNWPELTELVGWNTFRMKRTYQRRIREEFEHIMPREAVDLLDKMLTLNPEKRISAKEAL 600
Query: 387 RSEFF-TTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSM 445
+ + E P LP++ EM +K++ R R A + G H R S
Sbjct: 601 NHPWIRSLEHTTVQPLKLPQHQDCHEMWSKKQKKSARLGRQAEGSSGSG---HSIRATSH 657
Query: 446 KAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPP 481
P + +SN H ++ + PP
Sbjct: 658 PRAPTQPSTTTTKSNGSSNHHHHHHHSHHHASSLPP 693
>CDK9_SCHPO (Q96WV9) Serine/threonine-protein kinase cdk9 (EC
2.7.1.37) (Cyclin-dependent kinase cdk9)
Length = 591
Score = 252 bits (643), Expect = 2e-66
Identities = 163/479 (34%), Positives = 244/479 (50%), Gaps = 42/479 (8%)
Query: 107 FEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDHPN 165
+ ++K+G+GT+ VYK+ GKV ALK++ + E E A REI IL+ + H N
Sbjct: 36 YHLMEKLGEGTFGEVYKSQRRKDGKVYALKRILM-HTEKEGFPITAIREIKILKSIKHEN 94
Query: 166 VIKLQGLVTSRMSC------SLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGL 219
+I L + R S+Y+V YM+HDL+GL +P ++FTE QIKCYM QL +G
Sbjct: 95 IIPLSDMTVVRADKKHRRRGSIYMVTPYMDHDLSGLLENPSVKFTEPQIKCYMKQLFAGT 154
Query: 220 EHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFF-DPNYMN-----------PMTSR 267
++ H++ +LHRD+K +NLLIDN GILKIADFGLA + +Y N T
Sbjct: 155 KYLHDQLILHRDLKAANLLIDNHGILKIADFGLARVITEESYANKNPGLPPPNRREYTGC 214
Query: 268 VVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSP 327
VVT WYR PELLLG Y ID+WS GCI+ E+ G+PI+ G ++++QL KI++LCGSP
Sbjct: 215 VVTRWYRSPELLLGERRYTTAIDMWSVGCIMAEMYKGRPILQGSSDLDQLDKIFRLCGSP 274
Query: 328 SDEYWKK-SKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDAL 386
+ KLP + + R + F F L +L ++P ER +AS AL
Sbjct: 275 TQATMPNWEKLPGCEGVRSFPSHPRTLETAFFTFGKEMTSLCGAILTLNPDERLSASMAL 334
Query: 387 RSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSMK 446
E+FTT PY +PS L Y S E D +R+ ++ A + +G ++ R R
Sbjct: 335 EHEYFTTPPYPANPSELQSYSASHEYDKRRKREQRDANSHAFEQTANGKRQFRFMTRGPS 394
Query: 447 ----AMPAPEANAELQ----------SNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPL 492
+ P N++ Q N+DR R + + + + F P D P
Sbjct: 395 DPWYGIRRPNYNSQPQYQRGSYNREGGNMDRSRNVNY--QPKRQQNFKPLTSD----LPQ 448
Query: 493 GSSHHIDPDTIPADISFTSTTYTYSKEPFQAWSGPIGNAADIGVSKRKKYTAGE-AWDL 550
+S + + + + + P Q S + N +D G R+ + + AW++
Sbjct: 449 KNSEFSETNAMNQTSNHSHADGQRYYRPEQDRSQRLRNPSDYGRQGRQSSQSQQPAWNV 507
>CDKA_HUMAN (Q15131) Cell division protein kinase 10 (EC 2.7.1.37)
(Serine/threonine-protein kinase PISSLRE)
Length = 360
Score = 250 bits (638), Expect = 8e-66
Identities = 134/326 (41%), Positives = 196/326 (60%), Gaps = 4/326 (1%)
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRR 160
R FEK+++IG+GTY VY+A D+ T ++VALKKVR D E + I + REI +L R
Sbjct: 34 RSVKEFEKLNRIGEGTYGIVYRARDTQTDEIVALKKVRMDK-EKDGIPISSLREITLLLR 92
Query: 161 LDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLE 220
L HPN+++L+ +V S++LV Y E DLA L + F+E+Q+KC + Q+L GL+
Sbjct: 93 LRHPNIVELKEVVVGNHLESIFLVMGYCEQDLASLLENMPTPFSEAQVKCIVLQVLRGLQ 152
Query: 221 HCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLL 280
+ H ++HRD+K SNLL+ ++G +K ADFGLA + + PMT +VVTLWYR PELLL
Sbjct: 153 YLHRNFIIHRDLKVSNLLMTDKGCVKTADFGLARAYGVP-VKPMTPKVVTLWYRAPELLL 211
Query: 281 GATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKK-SKLPN 339
G T ID+W+ GCIL ELL +P++PG +E+ Q+ I +L G+PS+ W SKLP
Sbjct: 212 GTTTQTTSIDMWAVGCILAELLAHRPLLPGTSEIHQIDLIVQLLGTPSENIWPGFSKLPL 271
Query: 340 ATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACD 399
+ R+ ++ F + L L+ L DP +R TA D L S +F +P C+
Sbjct: 272 VGQYSLRKQPYNNLKHKFPWLSEAGLRLLHFLFMYDPKKRATAGDCLESSYFKEKPLPCE 331
Query: 400 PSSLPKYPPSKEMDAKRRDDEVRRQR 425
P +P +P + A E + +R
Sbjct: 332 PELMPTFPHHRNKRAAPATSEGQSKR 357
>CDK9_HUMAN (P50750) Cell division protein kinase 9 (EC 2.7.1.37)
(Serine/threonine-protein kinase PITALRE) (C-2K)
Length = 372
Score = 248 bits (634), Expect = 2e-65
Identities = 145/308 (47%), Positives = 193/308 (62%), Gaps = 18/308 (5%)
Query: 107 FEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDHPN 165
+EK+ KIGQGT+ V+KA TG+ VALKKV +N E E A REI IL+ L H N
Sbjct: 19 YEKLAKIGQGTFGEVFKARHRKTGQKVALKKVLMEN-EKEGFPITALREIKILQLLKHEN 77
Query: 166 VIKLQGLVTSRMS----C--SLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGL 219
V+ L + ++ S C S+YLVFD+ EHDLAGL ++ +++FT S+IK M LL+GL
Sbjct: 78 VVNLIEICRTKASPYNRCKGSIYLVFDFCEHDLAGLLSNVLVKFTLSEIKRVMQMLLNGL 137
Query: 220 EHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFD---PNYMNPMTSRVVTLWYRPP 276
+ H ++LHRD+K +N+LI +G+LK+ADFGLA F + N T+RVVTLWYRPP
Sbjct: 138 YYIHRNKILHRDMKAANVLITRDGVLKLADFGLARAFSLAKNSQPNRYTNRVVTLWYRPP 197
Query: 277 ELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSK 336
ELLLG DYG IDLW AGCI+ E+ PIM TE QL I +LCGS + E W
Sbjct: 198 ELLLGERDYGPPIDLWGAGCIMAEMWTRSPIMQANTEQHQLALISQLCGSITPEVW--PN 255
Query: 337 LPNATLFKPRE---PYKRCIRETFKGF--PPSALPLIDKLLAIDPVERETASDALRSEFF 391
+ N L++ E KR +++ K + P AL LIDKLL +DP +R + DAL +FF
Sbjct: 256 VDNYELYEKLELVKGQKRKVKDRLKAYVRDPYALDLIDKLLVLDPAQRIDSDDALNHDFF 315
Query: 392 TTEPYACD 399
++P D
Sbjct: 316 WSDPMPSD 323
>CDL1_RAT (P46892) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1)
Length = 436
Score = 231 bits (588), Expect = 5e-60
Identities = 137/391 (35%), Positives = 207/391 (52%), Gaps = 12/391 (3%)
Query: 31 TTASNGEEKNVVEIENDQKKKSDDSVQRSRRSKPNPRLSNPPKH--LRGEQVAAGWPSWL 88
TT+ + V EI KK V+ R+ P P + P L ++ P +L
Sbjct: 8 TTSWLFQSHEVTEILGRVKKNRKKLVKGLHRAGPPPEKNYLPDSPALSPIELKQELPKYL 67
Query: 89 TAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESI 148
A+ G R + F+ +++I +GTY VY+A D T ++VALK+++ + +
Sbjct: 68 PALQG-------CRSVEEFQCLNRIEEGTYGVVYRAKDKKTDEIVALKRLKMEKEKEGFP 120
Query: 149 KFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQI 208
REI + + HPN++ ++ +V +Y+V +Y+EHDL L + F ++
Sbjct: 121 LTSIREINTILKAQHPNIVTVREIVVGSNMDKIYIVMNYVEHDLKSLMETMKQPFLPGEV 180
Query: 209 KCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRV 268
K M QLLSG++H H+ +LHRD+K SNLL+ + GILK+ DFGLA + + + T V
Sbjct: 181 KTLMIQLLSGVKHLHDNWILHRDLKTSNLLLTHAGILKVGDFGLAREYG-SPLKAYTPVV 239
Query: 269 VTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPS 328
VTLWYR PELLLGA +Y D+WS GCI GELL KP+ PG+++++Q++KI+K G+PS
Sbjct: 240 VTLWYRAPELLLGAKEYSTACDMWSVGCIFGELLTQKPLFPGKSDIDQINKIFKDIGTPS 299
Query: 329 DEYWK-KSKLPNATLFKPRE-PYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDAL 386
++ W S+LP E PY + L++K L P R A D L
Sbjct: 300 EKIWPGYSELPAVKKMTFSELPYNNLRKRFGALLSDQGFDLMNKFLTYYPGRRINAEDGL 359
Query: 387 RSEFFTTEPYACDPSSLPKYPPSKEMDAKRR 417
+ E+F P DPS P +P E +R
Sbjct: 360 KHEYFRETPLPIDPSMFPTWPAKSEQQCVKR 390
>CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1) (CLK-1)
(CDK11) (p58 CLK-1)
Length = 795
Score = 228 bits (580), Expect = 4e-59
Identities = 122/324 (37%), Positives = 185/324 (56%), Gaps = 5/324 (1%)
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRR 160
R + F+ +++I +GTY VY+A D T ++VALK+++ + E E + REI + +
Sbjct: 433 RSVEEFQCLNRIEEGTYGVVYRAKDKKTDEIVALKRLKMEK-EKEGFPITSLREINTILK 491
Query: 161 LDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLE 220
HPN++ ++ +V +Y+V +Y+EHDL L + F ++K M QLL G++
Sbjct: 492 AQHPNIVTVREIVVGSNMDKIYIVMNYVEHDLKSLMETMKQPFLPGEVKTLMIQLLRGVK 551
Query: 221 HCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLL 280
H H+ +LHRD+K SNLL+ + GILK+ DFGLA + + + T VVTLWYR PELLL
Sbjct: 552 HLHDNWILHRDLKTSNLLLSHAGILKVGDFGLAREYG-SPLKAYTPVVVTLWYRAPELLL 610
Query: 281 GATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPN 339
GA +Y +D+WS GCI GELL KP+ PG++E++Q++K++K G+PS++ W S+LP
Sbjct: 611 GAKEYSTAVDMWSVGCIFGELLTQKPLFPGKSEIDQINKVFKDLGTPSEKIWPGYSELPA 670
Query: 340 ATLFKPRE-PYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYAC 398
E PY + L++K L P R +A D L+ E+F P
Sbjct: 671 VKKMTFSEHPYNNLRKRFGALLSDQGFDLMNKFLTYFPGRRISAEDGLKHEYFRETPLPI 730
Query: 399 DPSSLPKYPPSKEMDAKRRDDEVR 422
DPS P +P E +R R
Sbjct: 731 DPSMFPTWPAKSEQQRVKRGTSPR 754
>CDL1_MOUSE (P24788) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1)
Length = 434
Score = 226 bits (575), Expect = 2e-58
Identities = 123/324 (37%), Positives = 183/324 (55%), Gaps = 5/324 (1%)
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRR 160
R + F+ +++I +GTY VY+A D T ++VALK+++ + E E + REI + +
Sbjct: 72 RSVEEFQCLNRIEEGTYGVVYRAKDKKTDEIVALKRLKMEK-EKEGFPITSLREINTILK 130
Query: 161 LDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLE 220
HPN++ ++ +V +Y+V +Y+EHDL L + F ++K M QLLSG++
Sbjct: 131 AQHPNIVTVREIVVGSNMDKIYIVMNYVEHDLKSLMETMKQPFLPGEVKTLMIQLLSGVK 190
Query: 221 HCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLL 280
H H+ +LHRD+K SNLL+ + GILK+ DFGLA + + + T VVTLWYR PELLL
Sbjct: 191 HLHDNWILHRDLKTSNLLLTHAGILKVGDFGLAREYG-SPLKAYTPVVVTLWYRAPELLL 249
Query: 281 GATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPN 339
GA +Y D+WS GCI GELL KP+ PG+++++Q++KI+K GSPS++ W + LP
Sbjct: 250 GAKEYSTACDMWSVGCIFGELLTQKPLFPGKSDIDQINKIFKDLGSPSEKIWPGYNDLPA 309
Query: 340 ATLFKPRE-PYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYAC 398
E PY + L++K L P R A D L+ E+F P
Sbjct: 310 VKKMTFSEIPYNNLRKRFGALLSDQGFDLMNKFLTYYPGRRINAEDGLKHEYFRETPLPI 369
Query: 399 DPSSLPKYPPSKEMDAKRRDDEVR 422
DPS P +P E +R R
Sbjct: 370 DPSMFPTWPAKSEQQRVKRGTSPR 393
>CDL2_HUMAN (Q9UQ88) PITSLRE serine/threonine-protein kinase CDC2L2
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 2) (CDK11)
Length = 780
Score = 224 bits (572), Expect = 4e-58
Identities = 121/324 (37%), Positives = 183/324 (56%), Gaps = 5/324 (1%)
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRR 160
R + F+ +++I +GTY VY+A D T ++VALK+++ + E E + REI + +
Sbjct: 418 RSVEEFQCLNRIEEGTYGVVYRAKDKKTDEIVALKRLKMEK-EKEGFPITSLREINTILK 476
Query: 161 LDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLE 220
HPN++ ++ +V +Y+V +Y+EHDL L + F ++K M QLL G++
Sbjct: 477 AQHPNIVTVREIVVGSNMDKIYIVMNYVEHDLKSLMETMKQPFLPGEVKTLMIQLLRGVK 536
Query: 221 HCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLL 280
H H+ +LHRD+K SNLL+ + GILK+ DFGLA + + + T VVT WYR PELLL
Sbjct: 537 HLHDNWILHRDLKTSNLLLSHAGILKVGDFGLAREYG-SPLKAYTPVVVTQWYRAPELLL 595
Query: 281 GATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPN 339
GA +Y +D+WS GCI GELL KP+ PG +E++Q++K++K G+PS++ W S+LP
Sbjct: 596 GAKEYSTAVDMWSVGCIFGELLTQKPLFPGNSEIDQINKVFKELGTPSEKIWPGYSELPV 655
Query: 340 ATLFKPRE-PYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYAC 398
E PY + L++K L P R +A D L+ E+F P
Sbjct: 656 VKKMTFSEHPYNNLRKRFGALLSDQGFDLMNKFLTYFPGRRISAEDGLKHEYFRETPLPI 715
Query: 399 DPSSLPKYPPSKEMDAKRRDDEVR 422
DPS P +P E +R R
Sbjct: 716 DPSMFPTWPAKSEQQRVKRGTSPR 739
>CDK7_XENLA (P20911) Cell division protein kinase 7 (EC 2.7.1.37)
(40 kDa protein kinase) (P40 MO15)
(CDC2/CDK2,4-activating kinase)
Length = 352
Score = 222 bits (566), Expect = 2e-57
Identities = 134/346 (38%), Positives = 199/346 (56%), Gaps = 21/346 (6%)
Query: 103 KADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESI---KFMAREIIILR 159
+A +EK+D +G+G ++ VYKA D T ++VA+KK++ + + + REI +L+
Sbjct: 14 RAKQYEKLDFLGEGQFATVYKARDKNTDRIVAIKKIKLGHRAEANDGINRTALREIKLLQ 73
Query: 160 RLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGL 219
L HPN+I L + + SL VFD+ME DL + + T + IK YM L GL
Sbjct: 74 ELSHPNIIGLLDAFGHKSNISL--VFDFMETDLEVIIKDTSLVLTPAHIKSYMLMTLQGL 131
Query: 220 EHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLA-SFFDPNYMNPMTSRVVTLWYRPPEL 278
E+ H+ +LHRD+K +NLL+D G+LK+ADFGLA SF PN + T +VVT WYR PEL
Sbjct: 132 EYLHHLWILHRDLKPNNLLLDENGVLKLADFGLAKSFGSPNRI--YTHQVVTRWYRSPEL 189
Query: 279 LLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKL 337
L GA YGVG+D+W+ GCIL ELL+ P +PG ++++QL +I++ G+P++E W S L
Sbjct: 190 LFGARMYGVGVDMWAVGCILAELLLRVPFLPGDSDLDQLTRIFETLGTPTEEQWPGMSSL 249
Query: 338 PNATLFK--PREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEP 395
P+ FK P P + F L L+ L +P R TAS ALR +F+ P
Sbjct: 250 PDYVAFKSFPGTP----LHLIFIAAGDDLLELLQGLFTFNPCARCTASQALRKRYFSNRP 305
Query: 396 YACDPSSLPKYPPSKEMDAKRRDDE----VRRQRAASKAQVDGSKK 437
+ LP+ P+ ++A + ++R+R Q D +KK
Sbjct: 306 APTPGNLLPR--PNCSIEALKEQQNLNLGIKRKRTEGMDQKDIAKK 349
>CTK1_YEAST (Q03957) CTD kinase alpha subunit (EC 2.7.1.-) (CTD
kinase 58 kDa subunit) (CTDK-I alpha subunit)
Length = 528
Score = 222 bits (565), Expect = 2e-57
Identities = 123/300 (41%), Positives = 183/300 (61%), Gaps = 10/300 (3%)
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRR 160
R + +I ++G+GTY VYKA ++ T K+VALKK+R E E + REI +L+
Sbjct: 178 RSTSVYLRIMQVGEGTYGKVYKAKNTNTEKLVALKKLRLQG-EREGFPITSIREIKLLQS 236
Query: 161 LDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLE 220
DHPNV ++ ++ ++Y++F+Y ++DL+GL + ++ + SQ K QLL G+E
Sbjct: 237 FDHPNVSTIKEIMVESQK-TVYMIFEYADNDLSGLLLNKEVQISHSQCKHLFKQLLLGME 295
Query: 221 HCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLL 280
+ H+ ++LHRD+KGSN+LIDN+G LKI DFGLA N T+RV+TLWYRPPELLL
Sbjct: 296 YLHDNKILHRDVKGSNILIDNQGNLKITDFGLAR--KMNSRADYTNRVITLWYRPPELLL 353
Query: 281 GATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKK-SKLPN 339
G T+YG +D+W GC+L EL I G E+EQ+ I+K+ G+P+ W +P
Sbjct: 354 GTTNYGTEVDMWGCGCLLVELFNKTAIFQGSNELEQIESIFKIMGTPTINSWPTLYDMPW 413
Query: 340 ATLFKPRE--PYKRCIRETFKGFPPSA--LPLIDKLLAIDPVERETASDALRSEFFTTEP 395
+ P++ Y E FK PS+ L L LL D +R +A++AL+S++F EP
Sbjct: 414 FFMIMPQQTTKYVNNFSEKFKSVLPSSKCLQLAINLLCYDQTKRFSATEALQSDYFKEEP 473
>CDK7_MOUSE (Q03147) Cell division protein kinase 7 (EC 2.7.1.37)
(CDK-activating kinase) (CAK) (TFIIH basal transcription
factor complex kinase subunit) (39 kDa protein kinase)
(P39 Mo15) (Protein-tyrosine kinase MPK-7) (CR4 protein
kinase) (CRK4)
Length = 346
Score = 220 bits (561), Expect = 7e-57
Identities = 134/341 (39%), Positives = 192/341 (56%), Gaps = 24/341 (7%)
Query: 100 IPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEP--ESIKFMA-REII 156
+ +A +EK+D +G+G ++ VYKA D T ++VA+KK++ + + I A REI
Sbjct: 5 VKSRAKRYEKLDFLGEGQFATVYKARDKNTNQIVAIKKIKLGHRSEAKDGINRTALREIK 64
Query: 157 ILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLL 216
+L+ L HPN+I L + + SL VFD+ME DL + + T S IK YM L
Sbjct: 65 LLQELSHPNIIGLLDAFGHKSNISL--VFDFMETDLEVIIKDNSLVLTPSHIKAYMLMTL 122
Query: 217 SGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLA-SFFDPNYMNPMTSRVVTLWYRP 275
GLE+ H +LHRD+K +NLL+D G+LK+ADFGLA SF PN T +VVT WYR
Sbjct: 123 QGLEYLHQHWILHRDLKPNNLLLDENGVLKLADFGLAKSFGSPN--RAYTHQVVTRWYRA 180
Query: 276 PELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYW-KK 334
PELL GA YGVG+D+W+ GCIL ELL+ P +PG ++++QL +I++ G+P++E W
Sbjct: 181 PELLFGARMYGVGVDMWAVGCILAELLLRVPFLPGDSDLDQLTRIFETLGTPTEEQWPDM 240
Query: 335 SKLPNATLFK--PREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFT 392
LP+ FK P P ++ F L LI L +P R TAS AL++++F+
Sbjct: 241 CSLPDYVTFKSFPGVP----LQHIFIAAGDDLLELIQGLFLFNPCTRTTASQALKTKYFS 296
Query: 393 TEPYACDPSSLP---------KYPPSKEMDAKRRDDEVRRQ 424
P LP K P + + KR+ E Q
Sbjct: 297 NRPGPTPGCQLPRPNCPVEALKEPANPTVATKRKRAEALEQ 337
>CC2A_ARATH (P24100) Cell division control protein 2 homolog A (EC
2.7.1.37)
Length = 294
Score = 220 bits (560), Expect = 9e-57
Identities = 124/292 (42%), Positives = 181/292 (61%), Gaps = 11/292 (3%)
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDH 163
D +EK++KIG+GTY VYKA D +T + +ALKK+R + E E + A REI +L+ + H
Sbjct: 2 DQYEKVEKIGEGTYGVVYKARDKVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQH 60
Query: 164 PNVIKLQGLVTSRMSCSLYLVFDYMEHDLAG-LAASPVIRFTESQIKCYMNQLLSGLEHC 222
N++KLQ +V S LYLVF+Y++ DL + ++P IK Y+ Q+L G+ +C
Sbjct: 61 SNIVKLQDVVHSEKR--LYLVFEYLDLDLKKHMDSTPDFSKDLHMIKTYLYQILRGIAYC 118
Query: 223 HNRRVLHRDIKGSNLLIDNE-GILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLG 281
H+ RVLHRD+K NLLID LK+ADFGLA F + T VVTLWYR PE+LLG
Sbjct: 119 HSHRVLHRDLKPQNLLIDRRTNSLKLADFGLARAFGIP-VRTFTHEVVTLWYRAPEILLG 177
Query: 282 ATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPNA 340
+ Y +D+WS GCI E++ KP+ PG +E++QL KI+++ G+P ++ W+ + LP+
Sbjct: 178 SHHYSTPVDIWSVGCIFAEMISQKPLFPGDSEIDQLFKIFRIMGTPYEDTWRGVTSLPDY 237
Query: 341 TLFKPREPYKRCIRETF-KGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
P+ +K ETF P + L+ K+L +DP +R A AL E+F
Sbjct: 238 KSAFPK--WKPTDLETFVPNLDPDGVDLLSKMLLMDPTKRINARAALEHEYF 287
>CC2A_ANTMA (Q38772) Cell division control protein 2 homolog A (EC
2.7.1.37)
Length = 294
Score = 219 bits (557), Expect = 2e-56
Identities = 120/291 (41%), Positives = 182/291 (62%), Gaps = 9/291 (3%)
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDH 163
+ +EK++KIG+GTY VYKA D +T + +ALKK+R + E E + A REI +L+ + H
Sbjct: 2 EQYEKVEKIGEGTYGVVYKARDRVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQH 60
Query: 164 PNVIKLQGLVTSRMSCSLYLVFDYMEHDLAG-LAASPVIRFTESQIKCYMNQLLSGLEHC 222
N+++LQ +V S LYLVF+Y++ DL + + P +K ++ Q+L G+ +C
Sbjct: 61 GNIVRLQDVVHSEKR--LYLVFEYLDLDLKKHMDSCPEFSQDPRLVKMFLYQILRGIAYC 118
Query: 223 HNRRVLHRDIKGSNLLIDNE-GILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLG 281
H+ RVLHRD+K NLLID LK+ADFGLA F + T VVTLWYR PE+LLG
Sbjct: 119 HSHRVLHRDLKPQNLLIDRRTNALKLADFGLARAFGIP-VRTFTHEVVTLWYRAPEILLG 177
Query: 282 ATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPNA 340
+ Y +D+WS GCI E++ +P+ PG +E+++L KI+++ G+P++E W + LP+
Sbjct: 178 SRHYSTPVDVWSVGCIFAEMVNQRPLFPGDSEIDELFKIFRVMGTPNEETWPGVTSLPDF 237
Query: 341 TLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
P+ P K + S L L+DK+L +DP +R TA +AL+ E+F
Sbjct: 238 KSAFPKWPAKE-LAAVVPNLDASGLDLLDKMLRLDPSKRITARNALQHEYF 287
>CDK7_RAT (P51952) Cell division protein kinase 7 (EC 2.7.1.37)
(CDK-activating kinase) (CAK) (TFIIH basal transcription
factor complex kinase subunit) (39 protein kinase) (P39
Mo15) (Fragment)
Length = 329
Score = 218 bits (556), Expect = 3e-56
Identities = 135/339 (39%), Positives = 196/339 (56%), Gaps = 21/339 (6%)
Query: 104 ADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEP--ESIKFMA-REIIILRR 160
A+ EK+D +G+G ++ VYKA D T ++VA+KK++ + + I A REI +L+
Sbjct: 1 ANRNEKLDFLGEGQFATVYKARDKNTNQIVAIKKIKLGHRSEAKDGINRTALREIKLLQE 60
Query: 161 LDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLE 220
L HPN+I L + + SL VFD+ME DL + + T S IK YM L GLE
Sbjct: 61 LSHPNIIGLLDAFGHKSNISL--VFDFMETDLEVIIKDNSLVLTPSHIKAYMLMTLQGLE 118
Query: 221 HCHNRRVLHRDIKGSNLLIDNEGILKIADFGLA-SFFDPNYMNPMTSRVVTLWYRPPELL 279
+ H +LHRD+K +NLL+D G+LK+ADFGLA SF PN+ T +VVT WYR PELL
Sbjct: 119 YLHQHWILHRDLKPNNLLLDENGVLKLADFGLAKSFGSPNWA--YTHQVVTRWYRAPELL 176
Query: 280 LGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYW-KKSKLP 338
GA YGVG+D+W+ GCIL ELL+ P +PG ++++QL +I++ G+P++E W LP
Sbjct: 177 FGARMYGVGVDMWAVGCILAELLLRVPFLPGDSDLDQLTRIFETLGTPTEEQWPDMCSLP 236
Query: 339 NATLFK--PREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPY 396
+ FK P P ++ F L LI L +P R TAS ALR+++F+ P
Sbjct: 237 DYVTFKSFPGIP----LQHIFIAAGDDLLELIQGLFLFNPCTRITASQALRTKYFSNRPG 292
Query: 397 ACDPSSLPKYPPSKEMDAKRRDDE----VRRQRAASKAQ 431
LP+ P+ ++A + +R+RA + Q
Sbjct: 293 PTPGCQLPR--PNCPVEALKEQXNPAMATKRKRAEALEQ 329
>CDK7_HUMAN (P50613) Cell division protein kinase 7 (EC 2.7.1.37)
(CDK-activating kinase) (CAK) (TFIIH basal transcription
factor complex kinase subunit) (39 kDa protein kinase)
(P39 Mo15) (STK1) (CAK1)
Length = 346
Score = 218 bits (554), Expect = 4e-56
Identities = 127/313 (40%), Positives = 182/313 (57%), Gaps = 15/313 (4%)
Query: 100 IPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEP--ESIKFMA-REII 156
+ +A +EK+D +G+G ++ VYKA D T ++VA+KK++ + + I A REI
Sbjct: 5 VKSRAKRYEKLDFLGEGQFATVYKARDKNTNQIVAIKKIKLGHRSEAKDGINRTALREIK 64
Query: 157 ILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLL 216
+L+ L HPN+I L + + SL VFD+ME DL + + T S IK YM L
Sbjct: 65 LLQELSHPNIIGLLDAFGHKSNISL--VFDFMETDLEVIIKDNSLVLTPSHIKAYMLMTL 122
Query: 217 SGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLA-SFFDPNYMNPMTSRVVTLWYRP 275
GLE+ H +LHRD+K +NLL+D G+LK+ADFGLA SF PN T +VVT WYR
Sbjct: 123 QGLEYLHQHWILHRDLKPNNLLLDENGVLKLADFGLAKSFGSPN--RAYTHQVVTRWYRA 180
Query: 276 PELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYW-KK 334
PELL GA YGVG+D+W+ GCIL ELL+ P +PG ++++QL +I++ G+P++E W
Sbjct: 181 PELLFGARMYGVGVDMWAVGCILAELLLRVPFLPGDSDLDQLTRIFETLGTPTEEQWPDM 240
Query: 335 SKLPNATLFK--PREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFT 392
LP+ FK P P + F L LI L +P R TA+ AL+ ++F+
Sbjct: 241 CSLPDYVTFKSFPGIP----LHHIFSAAGDDLLDLIQGLFLFNPCARITATQALKMKYFS 296
Query: 393 TEPYACDPSSLPK 405
P LP+
Sbjct: 297 NRPGPTPGCQLPR 309
>CDC2_CHERU (P93101) Cell division control protein 2 homolog (EC
2.7.1.37) (p34cdc2)
Length = 294
Score = 217 bits (553), Expect = 6e-56
Identities = 121/293 (41%), Positives = 182/293 (61%), Gaps = 13/293 (4%)
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDH 163
D +EK++KIG+GTY VYKA D +T + +ALKK+R + E E + A REI +L+ + H
Sbjct: 2 DQYEKVEKIGEGTYGVVYKARDKVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQH 60
Query: 164 PNVIKLQGLVTSRMSCSLYLVFDYMEHDLAG-LAASPVIRFTESQIKCYMNQLLSGLEHC 222
N+++LQ +V S LYLVF+Y++ DL + + P IK ++ Q+L G+ +C
Sbjct: 61 GNIVRLQDVVHSEKR--LYLVFEYLDLDLKKHMDSCPDFAKDPRMIKRFLYQILRGIAYC 118
Query: 223 HNRRVLHRDIKGSNLLIDNE-GILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLG 281
H+ RVLHRD+K NLLID + LK+ADFGLA F + T VVTLWYR PE+LLG
Sbjct: 119 HSHRVLHRDLKPQNLLIDRQTNALKLADFGLARAFGIP-VRTFTHEVVTLWYRAPEILLG 177
Query: 282 ATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPNA 340
+ Y +D+WS GCI E++ KP+ PG +E+++L KI++ G+P++E W + LP+
Sbjct: 178 SRHYSTPVDVWSVGCIFAEMVNQKPLFPGDSEIDELFKIFRTLGTPNEETWPGVTSLPD- 236
Query: 341 TLFKPREP--YKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
FK P + + P+ + L++K+L +DP +R TA +AL E+F
Sbjct: 237 --FKSSFPKWISKDLSAVVPNLDPAGIDLLNKMLCLDPSKRITARNALEHEYF 287
>CC22_MEDSA (Q05006) Cell division control protein 2 homolog 2 (EC
2.7.1.37)
Length = 294
Score = 217 bits (553), Expect = 6e-56
Identities = 119/291 (40%), Positives = 179/291 (60%), Gaps = 9/291 (3%)
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDH 163
+ +EK++KIG+GTY VYKA D T + +ALKK+R + E E + A REI +L+ + H
Sbjct: 2 EQYEKVEKIGEGTYGVVYKARDRATNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQH 60
Query: 164 PNVIKLQGLVTSRMSCSLYLVFDYMEHDLAG-LAASPVIRFTESQIKCYMNQLLSGLEHC 222
N+++LQ +V S LYLVF+Y++ DL + +SP + QIK ++ Q+L G+ +C
Sbjct: 61 RNIVRLQDVVHSEKR--LYLVFEYLDLDLKKFMDSSPEFAKDQRQIKMFLYQILCGIAYC 118
Query: 223 HNRRVLHRDIKGSNLLID-NEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLG 281
H+ RVLHRD+K NLLID + +K+ADFGLA F + T VVTLWYR PE+LLG
Sbjct: 119 HSHRVLHRDLKPQNLLIDRSSNAVKLADFGLARAFGIP-VRTFTHEVVTLWYRAPEILLG 177
Query: 282 ATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPNA 340
+ Y +D+WS GCI E++ +P+ PG +E+++L KI+++ G+P++E W + LP+
Sbjct: 178 SRHYSTPVDVWSVGCIFAEMINQRPLFPGDSEIDELFKIFRITGTPNEETWPGVTSLPDF 237
Query: 341 TLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
P+ P K + P+ L L+ +DP R TA AL E+F
Sbjct: 238 KSAFPKWPAKDLATQV-PNLEPAGLDLLSSTCRLDPTRRITARGALEHEYF 287
>YP62_CAEEL (Q09437) Putative serine/threonine-protein kinase
B0495.2 (EC 2.7.1.37)
Length = 719
Score = 216 bits (550), Expect = 1e-55
Identities = 138/410 (33%), Positives = 219/410 (52%), Gaps = 40/410 (9%)
Query: 33 ASNGEEKNVVEIENDQKKKSDDSVQRSRRSKPNPRLSNPPKHLRGEQVAAGWPSWLTAVC 92
A + E+K + E+ + ++ ++ QR + R S K L Q+ +P +
Sbjct: 294 ADSDEDKYLKTPEDREWEEMTETEQRLHKEAMKKRASMKQKTLIA-QLPVFYPGLMGC-- 350
Query: 93 GEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA 152
R D +E ++++ +GT+ VY+ D T ++VALK+++ + E E A
Sbjct: 351 ---------RNIDEYECVNRVDEGTFGVVYRGKDKRTDEIVALKRLKMEK-EKEGFPITA 400
Query: 153 -REI-IILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIR---FTESQ 207
REI ++L+ +HPN++ ++ ++ +Y+ +++EHD+ L + R F+ +
Sbjct: 401 LREINMLLKAGNHPNIVNVKEILLGSNMDKIYMAMEFVEHDMKSLLDTMSRRNKRFSIGE 460
Query: 208 IKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLA-SFFDPNYMNPMTS 266
K + QLLSG+EH H +LHRD+K SNLL+ ++GILKIADFGLA + DP + TS
Sbjct: 461 QKTLLQQLLSGIEHMHKLWILHRDLKTSNLLMSHKGILKIADFGLAREYGDP--LKKFTS 518
Query: 267 RVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGS 326
VVTLWYR PELLLG Y +D+WS GCI+ E ++ KP+ PGR E+EQ+ KI+ G+
Sbjct: 519 IVVTLWYRSPELLLGTRLYSTPVDMWSVGCIMAEFILLKPLFPGRGELEQIKKIFMEMGT 578
Query: 327 PSDEYWKKSKLPNAT-------LFKPREPYKRCIRETFKG--FPPSALPLIDKLLAIDPV 377
P++ W P T L + PY + + G + L++ LL +DP
Sbjct: 579 PTESIW-----PGVTELDGWKALTFEKYPYNQLRKRFLAGRLLNDTGFKLLNGLLTLDPK 633
Query: 378 ERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMD-----AKRRDDEVR 422
R +A+ AL E+FT EPY P P +P E + AK++ E R
Sbjct: 634 NRFSATQALDHEWFTEEPYPVPPEEFPTFPAKSEQNKAPPPAKQKQQENR 683
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,875,837
Number of Sequences: 164201
Number of extensions: 3158353
Number of successful extensions: 13224
Number of sequences better than 10.0: 1696
Number of HSP's better than 10.0 without gapping: 1515
Number of HSP's successfully gapped in prelim test: 181
Number of HSP's that attempted gapping in prelim test: 8593
Number of HSP's gapped (non-prelim): 2089
length of query: 569
length of database: 59,974,054
effective HSP length: 116
effective length of query: 453
effective length of database: 40,926,738
effective search space: 18539812314
effective search space used: 18539812314
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC130801.5


