

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.12 + phase: 0
(124 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
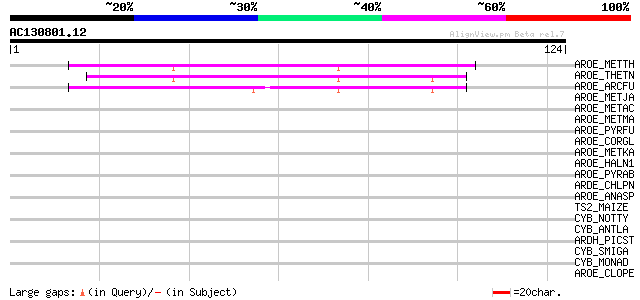
Sequences producing significant alignments: (bits) Value
AROE_METTH (O26344) Shikimate dehydrogenase (EC 1.1.1.25) 46 1e-05
AROE_THETN (Q8RAG2) Shikimate dehydrogenase (EC 1.1.1.25) 42 2e-04
AROE_ARCFU (O27957) Shikimate dehydrogenase (EC 1.1.1.25) 42 3e-04
AROE_METJA (Q58484) Shikimate dehydrogenase (EC 1.1.1.25) 40 0.001
AROE_METAC (Q8THC3) Shikimate dehydrogenase (EC 1.1.1.25) 39 0.002
AROE_METMA (Q8PXE6) Shikimate dehydrogenase (EC 1.1.1.25) 37 0.006
AROE_PYRFU (Q8U0A6) Shikimate dehydrogenase (EC 1.1.1.25) 37 0.010
AROE_CORGL (Q9X5C9) Shikimate dehydrogenase (EC 1.1.1.25) 37 0.010
AROE_METKA (Q8TZ24) Shikimate dehydrogenase (EC 1.1.1.25) 36 0.013
AROE_HALN1 (Q9HS68) Shikimate dehydrogenase (EC 1.1.1.25) 35 0.030
AROE_PYRAB (Q9V1H7) Shikimate dehydrogenase (EC 1.1.1.25) 34 0.051
ARDE_CHLPN (Q9Z6M4) Shikimate biosynthesis protein aroDE [Includ... 33 0.11
AROE_ANASP (Q8YVC1) Shikimate dehydrogenase (EC 1.1.1.25) 32 0.33
TS2_MAIZE (P50160) Sex determination protein tasselseed 2 31 0.43
CYB_NOTTY (O03478) Cytochrome b 31 0.56
CYB_ANTLA (O20434) Cytochrome b 31 0.56
ARDH_PICST (P50167) D-arabinitol 2-dehydrogenase [ribulose formi... 31 0.56
CYB_SMIGA (Q9XP83) Cytochrome b 30 0.73
CYB_MONAD (Q34957) Cytochrome b 30 0.73
AROE_CLOPE (Q8XMI8) Shikimate dehydrogenase (EC 1.1.1.25) 30 0.73
>AROE_METTH (O26344) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 283
Score = 46.2 bits (108), Expect = 1e-05
Identities = 30/97 (30%), Positives = 53/97 (53%), Gaps = 6/97 (6%)
Query: 14 LFVRAGGAGKALAFGAKTRGAR-IVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGA 72
L + AGGA +A AF +GA I I + +K+R LA + ++ + ++ ++G+
Sbjct: 126 LILGAGGAARACAFQLAEKGASDITILNRTPEKARLLAEDMADKLGFEASYGGYELIQGS 185
Query: 73 -----ILANATPIGMHPDTDRIPVAEYQLVFDAVYLH 104
IL + TP+GMHP TD P+ +L+ + + +H
Sbjct: 186 VKSADILIDTTPVGMHPHTDDRPLVGAELMHEGLVVH 222
>AROE_THETN (Q8RAG2) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 280
Score = 42.0 bits (97), Expect = 2e-04
Identities = 30/95 (31%), Positives = 47/95 (48%), Gaps = 10/95 (10%)
Query: 18 AGGAGKALAFGAKTRGAR-IVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGA---- 72
AGGA KA+ F G IVI + +K+++LA + E + + + + V+
Sbjct: 132 AGGASKAICFALAREGVESIVIANRTLNKAKALAEYIREEFKMKCDYCSIEEVEKFNEID 191
Query: 73 ILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
IL N T +GMHP+ PV+E V+D +Y
Sbjct: 192 ILINTTSVGMHPEVGNSPVSEEVVAKANFVYDLIY 226
>AROE_ARCFU (O27957) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 269
Score = 41.6 bits (96), Expect = 3e-04
Identities = 33/98 (33%), Positives = 48/98 (48%), Gaps = 10/98 (10%)
Query: 14 LFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAV--FGEVQSYKNLVNFQPVKG 71
L V AGGAGKA A G+ +++ + +K R + +GE + L + +KG
Sbjct: 117 LVVGAGGAGKAAALALLDMGSTVIVANRTEEKGREAVEMLRRYGEC-IFWPLSRVEELKG 175
Query: 72 A--ILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
++ NATP+GM IPV +LVFD VY
Sbjct: 176 KVDVVVNATPLGMRGFKAEIPVPPSMLDGVELVFDTVY 213
>AROE_METJA (Q58484) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 282
Score = 39.7 bits (91), Expect = 0.001
Identities = 33/99 (33%), Positives = 50/99 (50%), Gaps = 17/99 (17%)
Query: 18 AGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAV-------FGEVQSYKNL-VNFQPV 69
AGGA +A+AF + I+I + +K+ +LA + FGE + L V+ V
Sbjct: 131 AGGAARAVAFEL-AKDNNIIIANRTVEKAEALAKEIAEKLNKKFGEEVKFSGLDVDLDGV 189
Query: 70 KGAILANATPIGMHPDTDRIPVA------EYQLVFDAVY 102
I+ NATPIGM+P+ D P+ E +V D +Y
Sbjct: 190 D--IIINATPIGMYPNIDVEPIVKAEKLREDMVVMDLIY 226
>AROE_METAC (Q8THC3) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 280
Score = 39.3 bits (90), Expect = 0.002
Identities = 32/98 (32%), Positives = 44/98 (44%), Gaps = 15/98 (15%)
Query: 18 AGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGA----- 72
AGGA +A+AF GA I + + +++ LA V S +N + G
Sbjct: 129 AGGAARAVAFQLAADGAEITVVNRTEERAVELAKDV--AAASLPGKINGTGLSGLKELLR 186
Query: 73 ---ILANATPIGMHPDTDRIPVAEYQL-----VFDAVY 102
IL N T +GMHP+TD +L VFD VY
Sbjct: 187 DADILINTTTLGMHPNTDTTIATAEELHSGLTVFDIVY 224
>AROE_METMA (Q8PXE6) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 280
Score = 37.4 bits (85), Expect = 0.006
Identities = 30/100 (30%), Positives = 44/100 (44%), Gaps = 19/100 (19%)
Query: 18 AGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGE----------VQSYKNLVNFQ 67
AGGA +A+AF GA I + + +++ LA + + K L+
Sbjct: 129 AGGAARAVAFQLAADGADITVINRTEERAVGLAREISAADLPGKIRGTGLSGLKELLR-- 186
Query: 68 PVKGAILANATPIGMHPDTDRIPVAEYQL-----VFDAVY 102
+L N T +GMHP+TD V +L VFD VY
Sbjct: 187 --DADVLINTTTLGMHPNTDATIVTAEELHSGLTVFDIVY 224
>AROE_PYRFU (Q8U0A6) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 271
Score = 36.6 bits (83), Expect = 0.010
Identities = 28/92 (30%), Positives = 47/92 (50%), Gaps = 6/92 (6%)
Query: 14 LFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAI 73
L + AGGAGKA+A+ ++ A +V+ + +K++ L FG V + + I
Sbjct: 125 LILGAGGAGKAIAY-ELSKVANVVVLNRTIEKAKRL--EKFGIVGDSLEALPYYVEWADI 181
Query: 74 LANATPIGMHPDTDRIP---VAEYQLVFDAVY 102
L NAT +GM+ + +P + +V D VY
Sbjct: 182 LINATSVGMNEEKSLVPKNLLRPGLVVMDIVY 213
>AROE_CORGL (Q9X5C9) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 283
Score = 36.6 bits (83), Expect = 0.010
Identities = 33/105 (31%), Positives = 54/105 (51%), Gaps = 11/105 (10%)
Query: 10 LDGRLFVRAGGAGKALAFGAKTRGA-RIVIFDIDFDKSRSLACAVFGEVQSYKNL-VNFQ 67
LD + V AGG G A+A+ T G ++ + D+D ++++LA + V + V+ +
Sbjct: 127 LDSVVQVGAGGVGNAVAYALVTHGVQKLQVADLDTSRAQALADVINNAVGREAVVGVDAR 186
Query: 68 PVKGAILA-----NATPIGM--HPDT--DRIPVAEYQLVFDAVYL 103
++ I A NATP+GM HP T D + + V D VY+
Sbjct: 187 GIEDVIAAADGVVNATPMGMPAHPGTAFDVSCLTKDHWVGDVVYM 231
>AROE_METKA (Q8TZ24) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 290
Score = 36.2 bits (82), Expect = 0.013
Identities = 31/99 (31%), Positives = 50/99 (50%), Gaps = 14/99 (14%)
Query: 18 AGGAGKALAFGAKTRGA-RIVIFDIDFDKSRSLACAVFGEVQSYKNLVNF------QPVK 70
AGGA +A++F T GA IVI + D++ LA + +V + + ++
Sbjct: 137 AGGAARAVSFKLATEGADEIVIANRTVDRAERLAEELKEKVGVKARAIGLDGDEIERELR 196
Query: 71 GA-ILANATPIGMHPDTDRIPVA------EYQLVFDAVY 102
A +L +ATP+GM+P+ D P+ E +V D VY
Sbjct: 197 DADLLVDATPVGMYPNEDEPPLVTADQMHEDLIVNDLVY 235
>AROE_HALN1 (Q9HS68) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 266
Score = 35.0 bits (79), Expect = 0.030
Identities = 32/92 (34%), Positives = 41/92 (43%), Gaps = 7/92 (7%)
Query: 16 VRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILA 75
V AGGAG+A AF GA + I + + LA V G +L +L
Sbjct: 123 VGAGGAGRAAAFALADAGATVRIANRTRAAADELAADVGGTAVGLGDLPR-SLADATVLV 181
Query: 76 NATPIGMHPDTDRIPVAEYQL-----VFDAVY 102
+AT +GM D D PV+ L V DAVY
Sbjct: 182 HATTVGM-DDPDTSPVSADALHDDLAVLDAVY 212
>AROE_PYRAB (Q9V1H7) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 264
Score = 34.3 bits (77), Expect = 0.051
Identities = 32/100 (32%), Positives = 50/100 (50%), Gaps = 12/100 (12%)
Query: 10 LDGR--LFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLA--CAVFGEVQSYKNLVN 65
++GR L + AGGAGKA+A+ ++ A IV+ + K++SL G + N V
Sbjct: 113 IEGRNVLILGAGGAGKAIAY-ELSKIANIVVLNRTPSKAKSLEKFGVKGGSLDELPNYVG 171
Query: 66 FQPVKGAILANATPIGMHPDTDRIP---VAEYQLVFDAVY 102
+ V L NAT +GM + +P + +V D VY
Sbjct: 172 WADV----LINATSVGMGTNESLVPRRLLRRELIVMDIVY 207
>ARDE_CHLPN (Q9Z6M4) Shikimate biosynthesis protein aroDE [Includes:
3-dehydroquinate dehydratase (EC 4.2.1.10)
(3-dehydroquinase) (Type I DHQase); Shikimate
dehydrogenase (EC 1.1.1.25)]
Length = 477
Score = 33.1 bits (74), Expect = 0.11
Identities = 21/64 (32%), Positives = 32/64 (49%), Gaps = 2/64 (3%)
Query: 16 VRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILA 75
V AGGA KA+A +GA + IF+ + +LA G+ +L NF+ + I+
Sbjct: 338 VGAGGAAKAIAATLAMQGANLHIFNRTLSSAAALATCCKGKAYPLGSLENFKTID--III 395
Query: 76 NATP 79
N P
Sbjct: 396 NCLP 399
>AROE_ANASP (Q8YVC1) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 289
Score = 31.6 bits (70), Expect = 0.33
Identities = 17/56 (30%), Positives = 29/56 (51%), Gaps = 9/56 (16%)
Query: 56 EVQSYKNLVNFQPVKGAILANATPIGMHPDTDRIPVAEYQLV--------FDAVYL 103
+V ++ L P + +L N TPIGM+P D P++ +LV +D +Y+
Sbjct: 178 QVHTWDYLAKLIP-QANLLVNTTPIGMYPQVDESPLSAEELVNLQTGTIAYDLIYI 232
>TS2_MAIZE (P50160) Sex determination protein tasselseed 2
Length = 336
Score = 31.2 bits (69), Expect = 0.43
Identities = 20/51 (39%), Positives = 29/51 (56%), Gaps = 3/51 (5%)
Query: 10 LDGRLFVRAGGA---GKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEV 57
LDG++ + GGA G+A+ GAR+VI DID +LA A+ +V
Sbjct: 53 LDGKVAIVTGGARGIGEAIVRLFAKHGARVVIADIDDAAGEALASALGPQV 103
>CYB_NOTTY (O03478) Cytochrome b
Length = 381
Score = 30.8 bits (68), Expect = 0.56
Identities = 16/40 (40%), Positives = 23/40 (57%)
Query: 79 PIGMHPDTDRIPVAEYQLVFDAVYLHFYIKLGGSKAVLFP 118
P G++PD D+IP Y + DA+ L F + L S A+ P
Sbjct: 208 PSGINPDADKIPFHPYYTIKDALGLLFLLLLLLSLALFSP 247
>CYB_ANTLA (O20434) Cytochrome b
Length = 381
Score = 30.8 bits (68), Expect = 0.56
Identities = 15/40 (37%), Positives = 24/40 (59%)
Query: 79 PIGMHPDTDRIPVAEYQLVFDAVYLHFYIKLGGSKAVLFP 118
P G++PD+D+IP Y + DA+ L F + + S A+ P
Sbjct: 208 PSGINPDSDKIPFHPYYTIKDALGLMFLLLILLSLALFSP 247
>ARDH_PICST (P50167) D-arabinitol 2-dehydrogenase [ribulose
forming] (EC 1.1.1.250) (ARDH)
Length = 278
Score = 30.8 bits (68), Expect = 0.56
Identities = 16/49 (32%), Positives = 29/49 (58%), Gaps = 3/49 (6%)
Query: 10 LDGRLFVRAGGAGKALAFGAKT---RGARIVIFDIDFDKSRSLACAVFG 55
LDGRL + GG+G A ++ +GA + + D++ ++++S A V G
Sbjct: 14 LDGRLAIITGGSGGLAAVISRALLAQGADVALIDMNLERTKSAAKEVLG 62
>CYB_SMIGA (Q9XP83) Cytochrome b
Length = 381
Score = 30.4 bits (67), Expect = 0.73
Identities = 15/40 (37%), Positives = 24/40 (59%)
Query: 79 PIGMHPDTDRIPVAEYQLVFDAVYLHFYIKLGGSKAVLFP 118
P G++PD+D+IP Y + DA+ L F + + S A+ P
Sbjct: 208 PSGINPDSDKIPFHPYYTIKDALGLMFLLLVLLSLALFSP 247
>CYB_MONAD (Q34957) Cytochrome b
Length = 382
Score = 30.4 bits (67), Expect = 0.73
Identities = 15/43 (34%), Positives = 25/43 (57%)
Query: 76 NATPIGMHPDTDRIPVAEYQLVFDAVYLHFYIKLGGSKAVLFP 118
++ P G++PD+D+IP Y + DA+ L I + S A+ P
Sbjct: 205 SSNPTGINPDSDKIPFHPYYTIKDALGLILMILILMSLAMFSP 247
>AROE_CLOPE (Q8XMI8) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 271
Score = 30.4 bits (67), Expect = 0.73
Identities = 25/91 (27%), Positives = 43/91 (46%), Gaps = 8/91 (8%)
Query: 18 AGGAGKA-LAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILAN 76
AGGA ++ L + ++ +IV+ D +K+ SY L + L N
Sbjct: 126 AGGAARSILKYLEDSKAKKIVLVSRDKEKAFKKFKDFNINFMSYGELEEIN--EEFALIN 183
Query: 77 ATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
TP GM+P+T+ + V+E +++ D VY
Sbjct: 184 TTPCGMYPNTNSVAVSEKVIKKFKVAVDIVY 214
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.143 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,721,454
Number of Sequences: 164201
Number of extensions: 541122
Number of successful extensions: 2186
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 73
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 2104
Number of HSP's gapped (non-prelim): 98
length of query: 124
length of database: 59,974,054
effective HSP length: 100
effective length of query: 24
effective length of database: 43,553,954
effective search space: 1045294896
effective search space used: 1045294896
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC130801.12


