

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.11 - phase: 0
(275 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
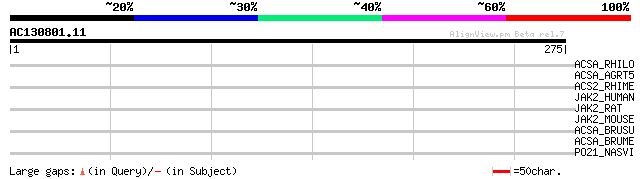
Score E
Sequences producing significant alignments: (bits) Value
ACSA_RHILO (Q98ET8) Acetyl-coenzyme A synthetase (EC 6.2.1.1) (A... 37 0.052
ACSA_AGRT5 (Q8UBV5) Acetyl-coenzyme A synthetase (EC 6.2.1.1) (A... 35 0.26
ACS2_RHIME (Q92KX2) Acetyl-coenzyme A synthetase 2 (EC 6.2.1.1) ... 33 0.58
JAK2_HUMAN (O60674) Tyrosine-protein kinase JAK2 (EC 2.7.1.112) ... 32 1.7
JAK2_RAT (Q62689) Tyrosine-protein kinase JAK2 (EC 2.7.1.112) (J... 31 3.7
JAK2_MOUSE (Q62120) Tyrosine-protein kinase JAK2 (EC 2.7.1.112) ... 31 3.7
ACSA_BRUSU (Q8FYQ3) Acetyl-coenzyme A synthetase (EC 6.2.1.1) (A... 31 3.7
ACSA_BRUME (Q8YJ48) Acetyl-coenzyme A synthetase (EC 6.2.1.1) (A... 31 3.7
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 30 6.4
>ACSA_RHILO (Q98ET8) Acetyl-coenzyme A synthetase (EC 6.2.1.1)
(Acetate--CoA ligase) (Acyl-activating enzyme)
Length = 651
Score = 37.0 bits (84), Expect = 0.052
Identities = 19/55 (34%), Positives = 30/55 (54%), Gaps = 2/55 (3%)
Query: 30 PLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWLWAWEEI--VLEECR 82
PL+E + D+A + V N+L +R + G GW+ R LW +E+ V EC+
Sbjct: 196 PLKENTDKAIDIAAKNFVMVKNVLVVRRTGGKVGWAPGRDLWYHDEVATVKAECK 250
>ACSA_AGRT5 (Q8UBV5) Acetyl-coenzyme A synthetase (EC 6.2.1.1)
(Acetate--CoA ligase) (Acyl-activating enzyme)
Length = 651
Score = 34.7 bits (78), Expect = 0.26
Identities = 16/47 (34%), Positives = 26/47 (55%)
Query: 30 PLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWLWAWEEI 76
PL+E + D+A +Y V +L +R + G GW+ R +W +EI
Sbjct: 195 PLKENTDKAIDIAAKQYVIVNKVLVVRRTGGKTGWAPGRDIWYHQEI 241
>ACS2_RHIME (Q92KX2) Acetyl-coenzyme A synthetase 2 (EC 6.2.1.1)
(Acetate--CoA ligase 2) (Acyl-activating enzyme 2)
Length = 649
Score = 33.5 bits (75), Expect = 0.58
Identities = 19/56 (33%), Positives = 29/56 (50%), Gaps = 2/56 (3%)
Query: 27 GGAP--LREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWLWAWEEIVLEE 80
GG P L+E D+A ++ TV+ +L +R + G GW+ R LW +E E
Sbjct: 190 GGKPVALKENTDTAIDIAARQHVTVSKVLVVRRTGGKVGWAPGRDLWYHQETAAAE 245
>JAK2_HUMAN (O60674) Tyrosine-protein kinase JAK2 (EC 2.7.1.112)
(Janus kinase 2) (JAK-2)
Length = 1132
Score = 32.0 bits (71), Expect = 1.7
Identities = 15/45 (33%), Positives = 24/45 (53%)
Query: 197 EQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLL 241
+ L L G CV D NI+ FV+F L + K ++ + ++W L
Sbjct: 607 KHLVLNYGVCVCGDENILVQEFVKFGSLDTYLKKNKNCINILWKL 651
>JAK2_RAT (Q62689) Tyrosine-protein kinase JAK2 (EC 2.7.1.112)
(Janus kinase 2) (JAK-2)
Length = 1132
Score = 30.8 bits (68), Expect = 3.7
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 1/61 (1%)
Query: 182 SDHLFLSCPFFAEL-WEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWL 240
S+ F + ++L + L L G CV + NI+ FV+F L + K ++ + ++W
Sbjct: 591 SESFFEAASMMSQLSHKHLVLNYGVCVCGEENILVQEFVKFGSLDTYLKKNKNSINILWK 650
Query: 241 L 241
L
Sbjct: 651 L 651
>JAK2_MOUSE (Q62120) Tyrosine-protein kinase JAK2 (EC 2.7.1.112)
(Janus kinase 2) (JAK-2)
Length = 1129
Score = 30.8 bits (68), Expect = 3.7
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 1/61 (1%)
Query: 182 SDHLFLSCPFFAEL-WEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWL 240
S+ F + ++L + L L G CV + NI+ FV+F L + K ++ + ++W
Sbjct: 591 SESFFEAASMMSQLSHKHLVLNYGVCVCGEENILVQEFVKFGSLDTYLKKNKNSINILWK 650
Query: 241 L 241
L
Sbjct: 651 L 651
>ACSA_BRUSU (Q8FYQ3) Acetyl-coenzyme A synthetase (EC 6.2.1.1)
(Acetate--CoA ligase) (Acyl-activating enzyme)
Length = 651
Score = 30.8 bits (68), Expect = 3.7
Identities = 18/56 (32%), Positives = 26/56 (46%), Gaps = 2/56 (3%)
Query: 27 GGAP--LREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWLWAWEEIVLEE 80
GG P L+E D+A +Y V +L +R + G W R LW +E+ E
Sbjct: 190 GGKPVALKENTDTAIDIAAKQYVMVNKVLVVRRTGGKVSWGRGRDLWYHQEVASVE 245
>ACSA_BRUME (Q8YJ48) Acetyl-coenzyme A synthetase (EC 6.2.1.1)
(Acetate--CoA ligase) (Acyl-activating enzyme)
Length = 651
Score = 30.8 bits (68), Expect = 3.7
Identities = 18/56 (32%), Positives = 26/56 (46%), Gaps = 2/56 (3%)
Query: 27 GGAP--LREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWLWAWEEIVLEE 80
GG P L+E D+A +Y V +L +R + G W R LW +E+ E
Sbjct: 190 GGKPVALKENTDTAIDIAAKQYVMVNKVLVVRRTGGKVSWGRGRDLWYHQEVASVE 245
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 30.0 bits (66), Expect = 6.4
Identities = 22/80 (27%), Positives = 31/80 (38%), Gaps = 6/80 (7%)
Query: 117 LTATGNSITDSA---VDLIWHHQVPLKVSIFAW-RLMRDRLSTRSNLA--SRGILQLDTA 170
LT G + D+ W + + + W + R++ L SRG A
Sbjct: 785 LTTDGKDLRDTRKAEASYSWIRDIHVAIPASVWIKYHHTRINALPTLMRMSRGRRTNGNA 844
Query: 171 LCVTGCGSIETSDHLFLSCP 190
LC GCG ET H+ CP
Sbjct: 845 LCRAGCGLPETLYHVVQQCP 864
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.483
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,522,219
Number of Sequences: 164201
Number of extensions: 1397290
Number of successful extensions: 3312
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 3306
Number of HSP's gapped (non-prelim): 9
length of query: 275
length of database: 59,974,054
effective HSP length: 109
effective length of query: 166
effective length of database: 42,076,145
effective search space: 6984640070
effective search space used: 6984640070
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC130801.11


