

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.9 + phase: 0
(487 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
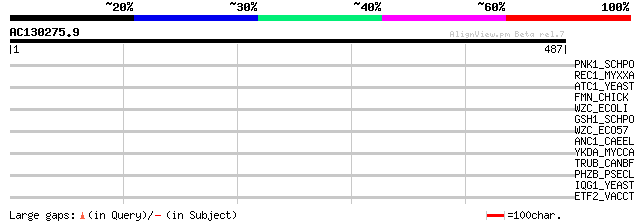
Score E
Sequences producing significant alignments: (bits) Value
PNK1_SCHPO (O13911) Bifunctional polynucleotide phosphatase/kina... 35 0.33
REC1_MYXXA (P48291) RecA protein 1 (Recombinase A 1) 33 1.3
ATC1_YEAST (P13586) Calcium-transporting ATPase 1 (EC 3.6.3.8) (... 33 1.7
FMN_CHICK (Q05858) Formin (Limb deformity protein) 32 2.8
WZC_ECOLI (P76387) Tyrosine-protein kinase wzc (EC 2.7.1.112) 32 3.7
GSH1_SCHPO (Q09768) Glutamate--cysteine ligase (EC 6.3.2.2) (Gam... 32 4.8
WZC_ECO57 (Q8X7L9) Tyrosine-protein kinase wzc (EC 2.7.1.112) 31 6.3
ANC1_CAEEL (Q9N4M4) Nuclear anchorage protein 1 (Anchorage 1 pro... 31 6.3
YKDA_MYCCA (P45615) Hypothetical lipoprotein in kdtB 5'region pr... 31 8.2
TRUB_CANBF (P59877) tRNA pseudouridine synthase B (EC 4.2.1.70) ... 31 8.2
PHZB_PSECL (Q51519) Anthranilate synthase, phenazine specific (E... 31 8.2
IQG1_YEAST (Q12280) Ras GTPase-activating-like protein IQG1 (IQG... 31 8.2
ETF2_VACCT (Q9JF93) Early transcription factor 82 kDa subunit (V... 31 8.2
>PNK1_SCHPO (O13911) Bifunctional polynucleotide phosphatase/kinase
(Polynucleotide kinase-3'-phosphatase) (DNA
5'-kinase/3'-phosphatase) [Includes: Polynucleotide
3'-phosphatase (EC 3.1.3.32) (2'(3')-polynucleotidase);
Polynucleotide 5'-hydroxy-kinase (E
Length = 421
Score = 35.4 bits (80), Expect = 0.33
Identities = 40/175 (22%), Positives = 65/175 (36%), Gaps = 23/175 (13%)
Query: 49 VPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGSKLEEACK 108
VPPN + PK Y+ N S PYH+ QE+ VL + P+ + E
Sbjct: 226 VPPNFESFHPKNYLVRNSSSHPYHFKKSEHQEIVVL-------VGFPSSGKSTLAESQIV 278
Query: 109 FLEKEEIN------ARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDNEFKHVPSLS 162
E +N C I+ E K +++ II +SDN + S
Sbjct: 279 TQGYERVNQDILKTKSKCIKAAIEALKKE--KSVVIGMYSIISTTYAISDNTNPTIESRK 336
Query: 163 RWMLPTIRREM------IMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKF 211
W+ I +E I L++ + +F N P+++F + +F
Sbjct: 337 MWI--DIAQEFEIPIRCIHLQSSEELARHNNVFRYIHHNQKQLPEIAFNSFKSRF 389
>REC1_MYXXA (P48291) RecA protein 1 (Recombinase A 1)
Length = 342
Score = 33.5 bits (75), Expect = 1.3
Identities = 15/71 (21%), Positives = 36/71 (50%)
Query: 248 KLIGEEPRGCQSQMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIG 307
++ G E G + +H+I +++ G ++ +D+S+ +K G+ V+EL + G
Sbjct: 65 EVFGNESSGKTTLTLHAIAQVQAAGGVAAFIDAEHALDVSYARKLGVRVEELLVSQPDTG 124
Query: 308 DHRGTVFRNIV 318
+ + ++V
Sbjct: 125 EQALEITEHLV 135
>ATC1_YEAST (P13586) Calcium-transporting ATPase 1 (EC 3.6.3.8)
(Golgi Ca(2+)-ATPase)
Length = 950
Score = 33.1 bits (74), Expect = 1.7
Identities = 20/58 (34%), Positives = 29/58 (49%), Gaps = 7/58 (12%)
Query: 405 FRASLVHNYLTSWAVGLSTLGALLALYFTFIQAICGFADAATKLEKAKFSSVIRSLII 462
F N + ++AVGLS LG + A+Y F Q+I K EK S ++ L+I
Sbjct: 871 FEIGFFTNKMFNYAVGLSLLGQMCAIYIPFFQSI-------FKTEKLGISDILLLLLI 921
>FMN_CHICK (Q05858) Formin (Limb deformity protein)
Length = 1213
Score = 32.3 bits (72), Expect = 2.8
Identities = 25/91 (27%), Positives = 44/91 (47%), Gaps = 9/91 (9%)
Query: 50 PPNMLNVEPKA---YIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANG---SKL 103
PP + E K Y + H +H +E+E LK++F R+F G + ++L
Sbjct: 485 PPKANHEEVKVGLKYTEAEYQAAILHLKREHKEEIETLKSQFELRVFHIRGEHAVSTAQL 544
Query: 104 EEACKFLEKE---EINARNCYMGEIKLSSDE 131
EE L+ E ++N RN +I +S+++
Sbjct: 545 EETIAHLKNELDNKLNRRNEEARDIGVSTED 575
>WZC_ECOLI (P76387) Tyrosine-protein kinase wzc (EC 2.7.1.112)
Length = 720
Score = 32.0 bits (71), Expect = 3.7
Identities = 15/53 (28%), Positives = 29/53 (54%)
Query: 417 WAVGLSTLGALLALYFTFIQAICGFADAATKLEKAKFSSVIRSLIIIPFRDPP 469
W +G++T+ AL A+ +TF ADA ++E+ +S+++ + PP
Sbjct: 33 WVIGITTVFALCAVVYTFFATPIYSADALVQIEQNSGNSLVQDIGSALANKPP 85
>GSH1_SCHPO (Q09768) Glutamate--cysteine ligase (EC 6.3.2.2)
(Gamma-glutamylcysteine synthetase) (Gamma-ECS) (GCS)
Length = 669
Score = 31.6 bits (70), Expect = 4.8
Identities = 27/105 (25%), Positives = 48/105 (45%), Gaps = 14/105 (13%)
Query: 297 KELTIPPLYIGDHRGTVFRNIVAFEKCH-KRCNPDMTT------YMFFLNRLINSANDVS 349
++LTI L+ G+HR F ++ + + CNPD T Y+ +++ N +
Sbjct: 561 RQLTIDELFNGEHRENGFPGLITIVRSYLYSCNPDAKTICLIERYIRLISQRANGQCLTA 620
Query: 350 ALHYKGVI-------HHSLGSDEHVAELINNIAKEIVPDMNESYL 387
A + I S+ +DE +LI IAK + D +++ L
Sbjct: 621 ASWIRNFITTHPSYKQDSVVNDEINYDLIRRIAKIVDGDYDDTLL 665
>WZC_ECO57 (Q8X7L9) Tyrosine-protein kinase wzc (EC 2.7.1.112)
Length = 720
Score = 31.2 bits (69), Expect = 6.3
Identities = 14/53 (26%), Positives = 29/53 (54%)
Query: 417 WAVGLSTLGALLALYFTFIQAICGFADAATKLEKAKFSSVIRSLIIIPFRDPP 469
W +G++ + AL A+ +TF ADA ++E++ +S+++ + PP
Sbjct: 33 WVIGITAVFALCAVVYTFFATPIYSADALVQIEQSSGNSLVQDIGSALANKPP 85
>ANC1_CAEEL (Q9N4M4) Nuclear anchorage protein 1 (Anchorage 1 protein)
(Nesprin homolog)
Length = 8545
Score = 31.2 bits (69), Expect = 6.3
Identities = 18/67 (26%), Positives = 30/67 (43%)
Query: 337 FLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNEANE 396
F++++ N N+V LH + V E + + NI K + D+N S + E N
Sbjct: 2910 FVDKIRNKINEVQKLHDEAVQDEKDELKEKLVAKVQNIGKTSIDDVNVSDFEEIEREING 2969
Query: 397 YLGCWRA 403
L + A
Sbjct: 2970 SLEAFEA 2976
>YKDA_MYCCA (P45615) Hypothetical lipoprotein in kdtB 5'region
precursor (ORFA)
Length = 655
Score = 30.8 bits (68), Expect = 8.2
Identities = 53/224 (23%), Positives = 93/224 (40%), Gaps = 35/224 (15%)
Query: 7 KLIKRKFHGTAVKSSKFWRGLVEKELQSIKPEEKNESVCIYRVPPNMLNVEPKAYIPSNI 66
K+IK K S L K L IK E + +++ + LN K I + I
Sbjct: 61 KIIKAIEKSDFNKISNINLELTIKFLTRIKNELETKTI-------SQLNKNDKLDILTKI 113
Query: 67 SIGPYHYGSQHLQEMEVLKNKFFHRL-------------FDPNGANGSKLEEA-CKFLEK 112
+ H GS +L E+ + ++ ++L + N N ++++ + LE
Sbjct: 114 KV---HLGSLNLIELVNIVDELVNKLNQKEEIKNTHKDKIEKNKDNIEDIDDSKLEILES 170
Query: 113 EEINARNCYMGEIK----LSSDEFLKMMLVDGSFIIQLLRDLSDNEFKHVPSLSRWMLPT 168
+ I ++ Y +K +S++E K L D +F I+ L L D + + W+L
Sbjct: 171 KYIPNQHNYPDYVKNFKTVSAEEIYKE-LYDRTFSIKFLVKLKDGGLLSNGTGTGWLLDY 229
Query: 169 IRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFF 212
+ N+ MF+ T L L D +NSL + N F ++
Sbjct: 230 HKYSNT---NKYKMFIATNLHVLADFSNSLTDEQ---NKEFNYY 267
>TRUB_CANBF (P59877) tRNA pseudouridine synthase B (EC 4.2.1.70)
(tRNA pseudouridine 55 synthase) (Psi55 synthase)
(Pseudouridylate synthase) (Uracil hydrolyase)
Length = 255
Score = 30.8 bits (68), Expect = 8.2
Identities = 16/59 (27%), Positives = 31/59 (52%), Gaps = 2/59 (3%)
Query: 371 INNIAKEIVPDMNESYLYNVVNEANEYLGCWR--ARFRASLVHNYLTSWAVGLSTLGAL 427
+N+I + + +Y+ ++VN+ EYLGC R +V Y+ S V + T+ ++
Sbjct: 168 LNSIIELEIHCSKGTYVRSIVNDIGEYLGCGAHIISLRRLMVGQYIASMMVNIKTIESI 226
>PHZB_PSECL (Q51519) Anthranilate synthase, phenazine specific (EC
4.1.3.27) [Includes: Glutamine amidotransferase]
Length = 637
Score = 30.8 bits (68), Expect = 8.2
Identities = 28/116 (24%), Positives = 44/116 (37%), Gaps = 10/116 (8%)
Query: 39 EKNESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHR------- 91
++ E+ +Y V L + + +GPY HL E HR
Sbjct: 234 DRKEADELYMVVDEELKMMARICEDGGHVLGPYLKEMAHLAHTEYFIEGKTHRDVREILR 293
Query: 92 --LFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQ 145
LF P GS LE AC+ +++ E R Y G L + +D + +I+
Sbjct: 294 ETLFAPT-VTGSPLESACRVIQRYEPQGRAYYSGMAALIGSDGKGGRSLDSAILIR 348
>IQG1_YEAST (Q12280) Ras GTPase-activating-like protein IQG1
(IQGAP-related protein 1) (Cytokinesis protein 1)
Length = 1495
Score = 30.8 bits (68), Expect = 8.2
Identities = 40/183 (21%), Positives = 80/183 (42%), Gaps = 21/183 (11%)
Query: 216 LQSESRKTPECQTSYKFKIEHVLDLLRYNIRPKLIGEEPRGCQSQMIHSITELKEGGVKI 275
L++++ + T K+E + L+ ++ KL+G+ +++ +++
Sbjct: 53 LKTKTSASNSSATILSKKVESSVSKLKPSLPNKLVGK----------YTVDLSNYSKIEL 102
Query: 276 KACESRELMDISFGKKWGIMVKELTIPP---LYIGDHRGTVFRNIVAFEKCHKRCNPDMT 332
+ E L +S K W V E +P L +GD RN V K +R NPD+T
Sbjct: 103 RYYEF--LCRVSEVKIWIEAVIEEALPSEIELCVGDS----LRNGVFLAKLTQRINPDLT 156
Query: 333 TYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNES--YLYNV 390
T +F + + + + G++ H D EL + K+ +P + E+ L ++
Sbjct: 157 TVIFPAGDKLQFKHTQNINAFFGLVEHVGVPDSFRFELQDLYNKKNIPQVFETLHILISM 216
Query: 391 VNE 393
+N+
Sbjct: 217 INK 219
>ETF2_VACCT (Q9JF93) Early transcription factor 82 kDa subunit (VETF
large subunit)
Length = 710
Score = 30.8 bits (68), Expect = 8.2
Identities = 19/74 (25%), Positives = 36/74 (47%), Gaps = 2/74 (2%)
Query: 331 MTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNV 390
MT +L L++ +D+ +K + +S + + K ++ N+S+LYNV
Sbjct: 63 MTQLKGYLGNLLDIKDDIIIYSHKNNLEYSYVDNTIFNPFVYTQKKTLLK--NDSFLYNV 120
Query: 391 VNEANEYLGCWRAR 404
+ A ++L W AR
Sbjct: 121 YSGACDFLVIWVAR 134
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 57,494,516
Number of Sequences: 164201
Number of extensions: 2423160
Number of successful extensions: 5811
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 5803
Number of HSP's gapped (non-prelim): 15
length of query: 487
length of database: 59,974,054
effective HSP length: 114
effective length of query: 373
effective length of database: 41,255,140
effective search space: 15388167220
effective search space used: 15388167220
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC130275.9


