

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.7 - phase: 0 /pseudo
(833 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
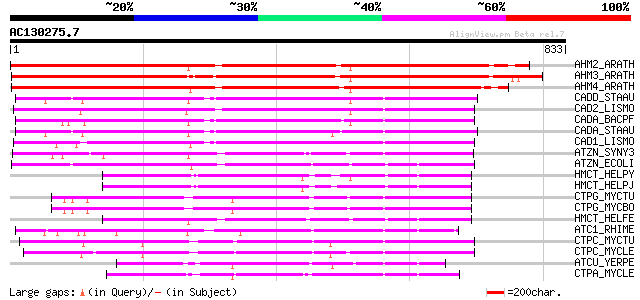
Score E
Sequences producing significant alignments: (bits) Value
AHM2_ARATH (O64474) Potential cadmium/zinc-transporting ATPase 2... 779 0.0
AHM3_ARATH (Q9SZW4) Potential cadmium/zinc-transporting ATPase 3... 766 0.0
AHM4_ARATH (Q9SZW5) Potential cadmium/zinc-transporting ATPase 4... 741 0.0
CADD_STAAU (P37386) Probable cadmium-transporting ATPase (EC 3.6... 334 5e-91
CAD2_LISMO (Q60048) Probable cadmium-transporting ATPase (EC 3.6... 334 7e-91
CADA_BACPF (P30336) Probable cadmium-transporting ATPase (EC 3.6... 333 1e-90
CADA_STAAU (P20021) Probable cadmium-transporting ATPase (EC 3.6... 331 4e-90
CAD1_LISMO (P58414) Probable cadmium-transporting ATPase (EC 3.6... 327 1e-88
ATZN_SYNY3 (Q59998) Zinc-transporting ATPase (EC 3.6.3.5) (Zn(II... 293 1e-78
ATZN_ECOLI (P37617) Lead, cadmium, zinc and mercury transporting... 275 3e-73
HMCT_HELPY (Q59465) Cadmium, zinc and cobalt transporting ATPase... 259 2e-68
HMCT_HELPJ (Q9ZL53) Cadmium, zinc and cobalt transporting ATPase... 251 6e-66
CTPG_MYCTU (P63689) Probable cation-transporting ATPase G (EC 3.... 251 7e-66
CTPG_MYCBO (P63690) Probable cation-transporting ATPase G (EC 3.... 251 7e-66
HMCT_HELFE (Q9RQB4) Cadmium, zinc and cobalt transporting ATPase... 242 3e-63
ATC1_RHIME (P58341) Copper-transporting ATPase 1 (EC 3.6.3.4) 242 3e-63
CTPC_MYCTU (P96875) Probable cation-transporting P-type ATPase C... 215 3e-55
CTPC_MYCLE (Q9CCL1) Probable cation-transporting P-type ATPase C... 215 4e-55
ATCU_YERPE (Q8ZCA7) Copper-transporting P-type ATPase (EC 3.6.3.4) 213 1e-54
CTPA_MYCLE (P46839) Cation-transporting P-type ATPase A (EC 3.6.... 201 5e-51
>AHM2_ARATH (O64474) Potential cadmium/zinc-transporting ATPase 2
(EC 3.6.3.3) (EC 3.6.3.5)
Length = 1172
Score = 779 bits (2012), Expect = 0.0
Identities = 414/796 (52%), Positives = 546/796 (68%), Gaps = 49/796 (6%)
Query: 2 VENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIA 61
V+ +++S F+VLG+CC +E ++E ILK L GVK SVIVP+RTV VVHD LLIS +IA
Sbjct: 13 VKKLQKSYFDVLGICCTSEVPIIENILKSLDGVKEYSVIVPSRTVIVVHDSLLISPFQIA 72
Query: 62 DALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGF 121
ALN ARLEA+ R GET+ + K P + GLLL LSFLK++Y PL WLA+ +VA G
Sbjct: 73 KALNEARLEANVRVNGETSFKNKWPSPFAVVSGLLLLLSFLKFVYSPLRWLAVAAVAAGI 132
Query: 122 PKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKAL 181
+L +A ASI+ ++INILV++ V T A+QDF + ++FLF+I+ WLETRA++KA
Sbjct: 133 YPILAKAFASIKRPRIDINILVIITVIATLAMQDFMEAAAVVFLFTISDWLETRASYKAT 192
Query: 182 VAMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKML 241
M SL ++APQ AI+AETGE V+V++VK++T++AVKAG+ IP+DGIVV+G CEVDEK L
Sbjct: 193 SVMQSLMSLAPQKAIIAETGEEVEVDEVKVDTVVAVKAGETIPIDGIVVDGNCEVDEKTL 252
Query: 242 TGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNV 296
TGE+FPV K+ DS VWAGTIN+N Y + D ++ K + + S + +
Sbjct: 253 TGEAFPVPKQRDSTVWAGTINLNGYICVKTTSLAGDCVVAKMAKLVEEAQSSKTKSQRLI 312
Query: 297 KTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALI 356
C + + +I ++ +C+A+VP + V +++ WFHLALVVL+SGCPC LI
Sbjct: 313 DKCSQYYTPAII----------LVSACVAIVPVIMKVHNLKHWFHLALVVLVSGCPCGLI 362
Query: 357 LSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDD 415
LSTPVA FCALTKAA SGLL+K DYL+TLS+IK VAFDKTGTITRGEF V DF S D
Sbjct: 363 LSTPVATFCALTKAATSGLLIKSADYLDTLSKIKIVAFDKTGTITRGEFIVIDFKSLSRD 422
Query: 416 INNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRD 475
IN +LLYW+SS+ESKSSHP+A +VDYA+ S++P PE VE++QNFPGEGI+G IDG D
Sbjct: 423 INLRSLLYWVSSVESKSSHPMAATIVDYAKSVSVEPRPEEVEDYQNFPGEGIYGKIDGND 482
Query: 476 IYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGAL 529
I+IGNK++ RA C E+ TK E L G F+L DACRSG
Sbjct: 483 IFIGNKKIASRAGCS------TVPEIEVDTKGGKTVGYVYVGERLAGFFNLSDACRSGVS 536
Query: 530 EAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI 589
+AM ELK LG+++ MLTGD+ AM+ Q QL N +D+VH +LLP +K+++I+ FKKEGP
Sbjct: 537 QAMAELKSLGIKTAMLTGDNQAAAMHAQEQLGNVLDVVHGDLLPEDKSRIIQEFKKEGPT 596
Query: 590 AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL 649
AM+GDG+NDAPALATADIGISMGISGSALA +T N ILMSNDIR++P+A++LAR+ RK+
Sbjct: 597 AMVGDGVNDAPALATADIGISMGISGSALATQTGNIILMSNDIRRIPQAVKLARRARRKV 656
Query: 650 VENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTK 709
VENV +S+ K ILALA AG+PL+W AVL DVGTCLLVI NSML+L+E K ++ +
Sbjct: 657 VENVCLSIILKAGILALAFAGHPLIWAAVLVDVGTCLLVIFNSMLLLREKKKIGNKKCYR 716
Query: 710 GSK---YGKFLEDKTKPLLNKQSNIDEEKGLLS---GEECGKDCCKNATHYVETSKERND 763
S G+ LE + +D E GLL+ +C CC ++N
Sbjct: 717 ASTSKLNGRKLEG------DDDYVVDLEAGLLTKSGNGQCKSSCC---------GDKKNQ 761
Query: 764 ESCGVSKLSLLNGNDH 779
E+ + K S +DH
Sbjct: 762 ENVVMMKPSSKTSSDH 777
>AHM3_ARATH (Q9SZW4) Potential cadmium/zinc-transporting ATPase 3
(EC 3.6.3.3) (EC 3.6.3.5)
Length = 951
Score = 766 bits (1979), Expect = 0.0
Identities = 420/829 (50%), Positives = 555/829 (66%), Gaps = 47/829 (5%)
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD 62
+ + +S F+VLG+CC +E L+E IL + GVK SVIVP+RTV VVHD L++S+ +I
Sbjct: 4 KKMTKSYFDVLGICCTSEVPLIENILNSMDGVKEFSVIVPSRTVIVVHDTLILSQFQIVK 63
Query: 63 ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFP 122
ALN A+LEA+ R GETN + K P + G+LL LSF KY+Y P WLA+ +V G
Sbjct: 64 ALNQAQLEANVRVTGETNFKNKWPSPFAVVSGILLLLSFFKYLYSPFRWLAVAAVVAGIY 123
Query: 123 KVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALV 182
+L +A+AS+ ++INILV++ V T +QD+T+ V++FLF+IA+WL++RA++KA
Sbjct: 124 PILAKAVASLARFRIDINILVVVTVGATIGMQDYTEAAVVVFLFTIAEWLQSRASYKASA 183
Query: 183 AMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLT 242
M SL ++APQ A++AETGE V+V+++K NT++AVKAG+ IP+DG+VV+G CEVDEK LT
Sbjct: 184 VMQSLMSLAPQKAVIAETGEEVEVDELKTNTVIAVKAGETIPIDGVVVDGNCEVDEKTLT 243
Query: 243 GESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RS 302
GE+FPV K DS VWAGTIN+N Y I +T C + K +N KT +
Sbjct: 244 GEAFPVPKLKDSTVWAGTINLNGY--ITVNTTAL-AEDCVVAKMAKLVEEAQNSKTETQR 300
Query: 303 F*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVA 362
F K +K + ++ C +P AL V +++ W HLALVVL+S CPC LILSTPVA
Sbjct: 301 FIDK--CSKYYTPAIILISICFVAIPFALKVHNLKHWVHLALVVLVSACPCGLILSTPVA 358
Query: 363 IFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDINNETL 421
FCALTKAA SGLL+KG DYLETL++IK VAFDKTGTITRGEF V DF S +DI+ ++L
Sbjct: 359 TFCALTKAATSGLLIKGADYLETLAKIKIVAFDKTGTITRGEFIVMDFQSLSEDISLQSL 418
Query: 422 LYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNK 481
LYW+SS ESKSSHP+A A+VDYAR S++P PE VE++QNFPGEGI+G IDG+++YIGNK
Sbjct: 419 LYWVSSTESKSSHPMAAAVVDYARSVSVEPKPEAVEDYQNFPGEGIYGKIDGKEVYIGNK 478
Query: 482 RVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGALEAMEEL 535
R+ RA C + ++ TK ETL GVF+L DACRSG +AM+EL
Sbjct: 479 RIASRAGC------LSVPDIDVDTKGGKTIGYVYVGETLAGVFNLSDACRSGVAQAMKEL 532
Query: 536 KLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK-EGPIAMIGD 594
K LG++ MLTGD+ AM+ Q QL NA+DIV AELLP +K+++I+ K+ EGP AM+GD
Sbjct: 533 KSLGIKIAMLTGDNHAAAMHAQEQLGNAMDIVRAELLPEDKSEIIKQLKREEGPTAMVGD 592
Query: 595 GINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVI 654
G+NDAPALATADIGISMG+SGSALA ET N ILMSNDIR++P+AI+LA++ RK+VENV+
Sbjct: 593 GLNDAPALATADIGISMGVSGSALATETGNIILMSNDIRRIPQAIKLAKRAKRKVVENVV 652
Query: 655 ISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKYG 714
IS+ K AILALA AG+PL+W AVL DVGTCLLVILNSML+L + HK + + S
Sbjct: 653 ISITMKGAILALAFAGHPLIWAAVLADVGTCLLVILNSMLLLSDKHKTGNKCYRESSSSS 712
Query: 715 KFLEDKTKPLLNKQSNIDEEKGLL---SGEECGKDCCKNATH-----------------Y 754
+ +K L + D E GLL S + C CC T
Sbjct: 713 VLIAEK----LEGDAAGDMEAGLLPKISDKHCKPGCCGTKTQEKAMKPAKASSDHSHSGC 768
Query: 755 VETSKERN----DESCGVSKLSLLNGNDHGMFMFVEVHVVKHCVMIQDS 799
ET ++ N +SC + L +G+D G +H V +Q S
Sbjct: 769 CETKQKDNVTVVKKSCCAEPVDLGHGHDSGCCGDKSQQPHQHEVQVQQS 817
>AHM4_ARATH (Q9SZW5) Potential cadmium/zinc-transporting ATPase 4
(EC 3.6.3.3) (EC 3.6.3.5)
Length = 760
Score = 741 bits (1913), Expect = 0.0
Identities = 398/764 (52%), Positives = 524/764 (68%), Gaps = 46/764 (6%)
Query: 4 NIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADA 63
N++ S F+V+G+CC++E ++V +L+ + GVK SVIVP+RTV VVHD LIS +I A
Sbjct: 11 NLQTSYFDVVGICCSSEVSIVGNVLRQVDGVKEFSVIVPSRTVIVVHDTFLISPLQIVKA 70
Query: 64 LNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFPK 123
LN ARLEAS RP GET+ + + P + G+LL LSF KY Y PL WLA+ +V G
Sbjct: 71 LNQARLEASVRPYGETSLKSQWPSPFAIVSGVLLVLSFFKYFYSPLEWLAIVAVVAGVFP 130
Query: 124 VLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVA 183
+L +A+AS+ L+IN L L+AV T +QDFT+ I+FLFS+A WLE+ A HKA +
Sbjct: 131 ILAKAVASVTRFRLDINALTLIAVIATLCMQDFTEAATIVFLFSVADWLESSAAHKASIV 190
Query: 184 MSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTG 243
MSSL ++AP+ A++A+TG VDV++V +NT+++VKAG++IP+DG+VV+G C+VDEK LTG
Sbjct: 191 MSSLMSLAPRKAVIADTGLEVDVDEVGINTVVSVKAGESIPIDGVVVDGSCDVDEKTLTG 250
Query: 244 ESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCKNYSVGKRHSSGKNVKT 298
ESFPV+K+ +S V A TIN+N Y K+ D ++ K + + + + +
Sbjct: 251 ESFPVSKQRESTVMAATINLNGYIKVKTTALARDCVVAKMTKLVEEAQKSQTKTQRFIDK 310
Query: 299 C*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILS 358
C R + V+ V +C AV+P L V D+ WFHLALVVL+SGCPC LILS
Sbjct: 311 CSRYYTPAVV----------VSAACFAVIPVLLKVQDLSHWFHLALVVLVSGCPCGLILS 360
Query: 359 TPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDIN 417
TPVA FCALTKAA SG L+K GD LETL++IK VAFDKTGTIT+ EF V+DF S IN
Sbjct: 361 TPVATFCALTKAATSGFLIKTGDCLETLAKIKIVAFDKTGTITKAEFMVSDFRSLSPSIN 420
Query: 418 NETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIY 477
LLYW+SSIE KSSHP+A AL+DYAR S++P P+ VENFQNFPGEG++G IDG+DIY
Sbjct: 421 LHKLLYWVSSIECKSSHPMAAALIDYARSVSVEPKPDIVENFQNFPGEGVYGRIDGQDIY 480
Query: 478 IGNKRVGVRAICKRDNCEVQFQRPEI-STKKNN-------CEETLVGVFSLVDACRSGAL 529
IGNKR+ RA C DN P+I +T K L G F+L+D CR G
Sbjct: 481 IGNKRIAQRAGCLTDNV------PDIEATMKRGKTIGYIYMGAKLTGSFNLLDGCRYGVA 534
Query: 530 EAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI 589
+A++ELK LG+++ MLTGD+ AM Q QL+NA+DIVH+ELLP +KA++I++FK +GP
Sbjct: 535 QALKELKSLGIQTAMLTGDNQDAAMSTQEQLENALDIVHSELLPQDKARIIDDFKIQGPT 594
Query: 590 AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL 649
M+GDG+NDAPALA ADIGISMGISGSALA ET + ILMSNDIRK+P+ +RLA+++ +K+
Sbjct: 595 MMVGDGLNDAPALAKADIGISMGISGSALATETGDIILMSNDIRKIPKGMRLAKRSHKKV 654
Query: 650 VENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEK---RE 706
+ENV++SV K AI+ L GYPLVW AVL D GTCLLVILNSM++L++ + R
Sbjct: 655 IENVVLSVSIKGAIMVLGFVGYPLVWAAVLADAGTCLLVILNSMMLLRDEREAVSTCYRA 714
Query: 707 STKGSKYGKFLEDKTKPLLNKQSNIDEEKGLL--SGEECGKDCC 748
ST S K ED+ + D E GLL S E K CC
Sbjct: 715 ST--SSPVKLEEDEVE---------DLEVGLLQKSEETSKKSCC 747
>CADD_STAAU (P37386) Probable cadmium-transporting ATPase (EC
3.6.3.3) (Cadmium efflux ATPase)
Length = 804
Score = 334 bits (857), Expect = 5e-91
Identities = 226/725 (31%), Positives = 378/725 (51%), Gaps = 46/725 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDV-------------LLIS 56
+ V G CA A E+ +K L GV+ V A + V + L +
Sbjct: 93 YRVEGFSCANCAGKFEKNVKQLAGVQDAKVNFGASKIDVYGNASVEELEKAGAFENLKVI 152
Query: 57 ESKIADALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPP-----LGW 111
K+A+ A E + P+ E K L A LL+A +L +
Sbjct: 153 PEKLANPSIQAVKEDTKAPKEEKIPFYKKHSTLLFAT-LLIAFGYLSHFVNGEDNLVTSM 211
Query: 112 LALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQW 171
L + S+ IG + ++ ++ L+ +AV G A + ++ + +++ LF+I++
Sbjct: 212 LFVSSIVIGGYSLFKVGFQNLIRFDFDMKTLMTVAVIGAAIIGEWAEASIVVILFAISEA 271
Query: 172 LETRATHKALVAMSSLTNMAPQMAIVAETGERV--DVNDVKMNTILAVKAGDAIPLDGIV 229
LE + +A ++ SL ++AP+ A+V G+ + V+D+ + I+ VK G+ I +DGI+
Sbjct: 272 LERFSMDRARQSIRSLMDIAPKEALVRRNGQEIMIHVDDIAVGDIMIVKPGEKIAMDGII 331
Query: 230 VEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNY 284
+ G V++ +TGES PV K D V+AGT+N K + DT +I
Sbjct: 332 INGVSAVNQAAITGESVPVAKTVDDEVFAGTLNEEGLLEVKITKYVEDTTISKIIHLVEE 391
Query: 285 SVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLAL 344
+ G+R + ++F K K + V+ + +AVVP + W + L
Sbjct: 392 AQGERAPA--------QAFVDKF--AKYYTPIIMVIAALVAVVPPLFFGGSWDTWVYQGL 441
Query: 345 VVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGE 404
VL+ GCPCAL+++TP++I A+ AA G+L+KGG YLE L IK +AFDKTGT+T+G
Sbjct: 442 AVLVVGCPCALVITTPISIVSAIGNAAKKGVLIKGGVYLEELGAIKAIAFDKTGTLTKGV 501
Query: 405 FSVTDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFP 463
VTDF D + + L I+++E +S HP+A A++ A +I V++F +
Sbjct: 502 PVVTDFKVLNDQVEEKELFSIITALEYRSQHPLASAIMKKAEQDNITYSDVRVKDFTSIT 561
Query: 464 GEGIFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNNC-----EETLVGVF 518
G GI G IDG YIG+ R+ + E + + + + ++T++GV
Sbjct: 562 GRGIQGNIDGTTYYIGSPRLFKELNVSDFSLEFENKVKVLQNQGKTAMIIGTDQTILGVI 621
Query: 519 SLVDACRSGALEAMEELKLLGVR-SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKA 577
++ D R + + +L LG++ ++MLTGD+ A + + + + + +ELLP +K
Sbjct: 622 AVADEVRETSKNVILKLHQLGIKQTIMLTGDNQGTAEAIGAHV--GVSDIQSELLPQDKL 679
Query: 578 KLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVP 636
I+ K E G +AMIGDG+NDAPALA + +GI+MG +G+ A ET++ LM +D+ K+P
Sbjct: 680 DYIKKMKAEHGNVAMIGDGVNDAPALAASTVGIAMGGAGTDTAIETADIALMGDDLSKLP 739
Query: 637 EAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
A+RL+RKT + N+ ++G K L L I G+ +W+A+L+D+G +LV LNS+ ++
Sbjct: 740 FAVRLSRKTLNIIKANITFAIGIKIIALLLVIPGWLTLWIAILSDMGATILVALNSLRLM 799
Query: 697 QENHK 701
+ K
Sbjct: 800 RVKDK 804
>CAD2_LISMO (Q60048) Probable cadmium-transporting ATPase (EC
3.6.3.3) (Cadmium efflux ATPase)
Length = 711
Score = 334 bits (856), Expect = 7e-91
Identities = 216/718 (30%), Positives = 381/718 (52%), Gaps = 38/718 (5%)
Query: 6 KRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLL--ISESKIADA 63
+++ + V G+ C A ER +K + GV V A +TV + + + ++ +
Sbjct: 3 EKTVYRVDGLSCTNCAAKFERNVKEIEGVTEAIVNFGASKITVTGEASIQQVEQAGAFEH 62
Query: 64 LNTARLEASFR-PQGETNNEKKC-PDILTMACGLLLALSFLKYI-----YPPLGWLALGS 116
L + SF P+ T+++ + + GL +A+ + I + L + +
Sbjct: 63 LKIIPEKESFTDPEHFTDHQSFIRKNWRLLLSGLFIAVGYASQIMNGEDFYLTNALFIFA 122
Query: 117 VAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRA 176
+ IG + ++ + L+ +A+ G A + ++ +G +++ LF++++ LE +
Sbjct: 123 IFIGGYSLFKEGFKNLLKFEFTMETLMTIAIIGAAFIGEWAEGSIVVILFAVSEALERYS 182
Query: 177 THKALVAMSSLTNMAPQMAIVAETG--ERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKC 234
KA ++ SL ++AP+ A+V +G V V+D+++ I+ +K G I +DG VV+G
Sbjct: 183 MDKARQSIRSLMDIAPKEALVRRSGTDRMVHVDDIQIGDIMIIKPGQKIAMDGHVVKGYS 242
Query: 235 EVDEKMLTGESFPVTKESDSLVWAGTINMN-----AYPKIIFDTYDFRIYKCKNYSVGKR 289
V++ +TGES PV K D V+AGT+N A K + DT +I + G+R
Sbjct: 243 AVNQAAITGESIPVEKNIDDSVFAGTLNEEGLLEVAVTKRVEDTTISKIIHLVEEAQGER 302
Query: 290 HSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLS 349
+ V T + + +I V+ + IA VP L + E W + L VL+
Sbjct: 303 APAQAFVDTFAKYYTPAII----------VIAALIATVPPLLFGGNWETWVYQGLSVLVV 352
Query: 350 GCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTD 409
GCPCAL++STPVAI A+ AA +G+L+KGG YLE + +K +AFDKTGT+T+G VTD
Sbjct: 353 GCPCALVVSTPVAIVTAIGNAAKNGVLVKGGVYLEEIGGLKAIAFDKTGTLTKGVPVVTD 412
Query: 410 F---SAGDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEG 466
+ + +I + ++++E S HP+A A++ Y + NV +F + G+G
Sbjct: 413 YIELTEATNIQHNKNYIIMAALEQLSQHPLASAIIKYGETREMDLTSINVNDFTSITGKG 472
Query: 467 IFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNNC-----EETLVGVFSLV 521
I GT+DG Y+G+ + + + + Q ++ K + L+ + ++
Sbjct: 473 IRGTVDGNTYYVGSPVLFKELLASQFTDSIHRQVSDLQLKGKTAMLFGTNQKLISIVAVA 532
Query: 522 DACRSGALEAMEELKLLGV-RSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLI 580
D RS + ++ L LG+ +++MLTGD+ A + Q+ + + EL+P +K I
Sbjct: 533 DEVRSSSQHVIKRLHELGIEKTIMLTGDNQATAQAIGQQV--GVSEIEGELMPQDKLDYI 590
Query: 581 ENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAI 639
+ K G +AM+GDGINDAPALA A +GI+MG +G+ A ET++ LM +D++K+P +
Sbjct: 591 KQLKINFGKVAMVGDGINDAPALAAATVGIAMGGAGTDTAIETADVALMGDDLQKLPFTV 650
Query: 640 RLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
+L+RKT + + +N+ S+ K L L I G+ +W+A++ D+G LLV LN + +++
Sbjct: 651 KLSRKTLQIIKQNITFSLVIKLIALLLVIPGWLTLWIAIMADMGATLLVTLNGLRLMK 708
>CADA_BACPF (P30336) Probable cadmium-transporting ATPase (EC
3.6.3.3) (Cadmium efflux ATPase)
Length = 723
Score = 333 bits (854), Expect = 1e-90
Identities = 223/721 (30%), Positives = 384/721 (52%), Gaps = 49/721 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALNTARL 69
+ V G CA A E+ +K L GV+ V A + V + I E + A A ++
Sbjct: 16 YRVQGFTCANCAGKFEKNVKQLSGVEDAKVNFGASKIAVYGNAT-IEELEKAGAFENLKV 74
Query: 70 --EASFRPQG----ETNNEKKCP----DILTMACGLLLALSFLK-YIYPPLG----WLAL 114
E S R E E K P + LL+ +L Y+ L L
Sbjct: 75 TPEKSARQASQEVKEDTKEDKVPFYKKHSTLLYASLLITFGYLSSYVNGEENIVTTLLFL 134
Query: 115 GSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLET 174
S+ IG + + ++ ++ L+ +AV G A + ++ + +++ LF+I++ LE
Sbjct: 135 ASMFIGGLSLFKVGLQNLLRFEFDMKTLMTVAVIGGAIIGEWAEVAIVVILFAISEALER 194
Query: 175 RATHKALVAMSSLTNMAPQMAIVAETGERV--DVNDVKMNTILAVKAGDAIPLDGIVVEG 232
+ +A ++ SL ++AP+ A+V G+ + V+D+ + I+ VK G I +DG+VV G
Sbjct: 195 FSMDRARQSIRSLMDIAPKEALVKRNGQEIMIHVDDIAVGDIMIVKPGQKIAMDGVVVSG 254
Query: 233 KCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVG 287
V++ +TGES PV K D+ V+AGT+N K++ DT +I + G
Sbjct: 255 YSAVNQTAITGESVPVEKTVDNEVFAGTLNEEGLLEVEITKLVEDTTISKIIHLVEEAQG 314
Query: 288 KRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVL 347
+R S ++F K K + ++ + +A+VP E W + L VL
Sbjct: 315 ERAPS--------QAFVDKF--AKYYTPIIMIIATLVAIVPPLFFDGSWETWIYQGLAVL 364
Query: 348 LSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSV 407
+ GCPCAL++STP++I A+ AA G+L+KGG YLE + +K +AFDKTGT+T+G +V
Sbjct: 365 VVGCPCALVISTPISIVSAIGNAAKKGVLVKGGVYLEEMGALKAIAFDKTGTLTKGVPAV 424
Query: 408 TDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEG 466
TD++ IN + LL I+++E +S HP+A A++ A +I VE+F + G+G
Sbjct: 425 TDYNVLNKQINEKELLSIITALEYRSQHPLASAIMKKAEEENITYSDVQVEDFSSITGKG 484
Query: 467 IFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNN--------CEETLVGVF 518
I G ++G YIG+ ++ + + +++ ++T +N E+ ++ V
Sbjct: 485 IKGIVNGTTYYIGSPKLFKELLTNDFDKDLE---QNVTTLQNQGKTAMIIGTEKEILAVI 541
Query: 519 SLVDACRSGALEAMEELKLLGVR-SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKA 577
++ D R + E +++L LG++ ++MLTGD+ A + Q+ + + AEL+P +K
Sbjct: 542 AVADEVRESSKEILQKLHQLGIKKTIMLTGDNKGTANAIGGQV--GVSDIEAELMPQDKL 599
Query: 578 KLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVP 636
I+ + E G +AM+GDG+NDAPALA + +GI+MG +G+ A ET++ LM +D+RK+P
Sbjct: 600 DFIKQLRSEYGNVAMVGDGVNDAPALAASTVGIAMGGAGTDTALETADVALMGDDLRKLP 659
Query: 637 EAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
++L+RKT + N+ ++ K L I G+ +W+A+L+D+G LLV LN + ++
Sbjct: 660 STVKLSRKTLNIIKANITFAIAIKFIASLLVIPGWLTLWIAILSDMGATLLVALNGLRLM 719
Query: 697 Q 697
+
Sbjct: 720 R 720
>CADA_STAAU (P20021) Probable cadmium-transporting ATPase (EC
3.6.3.3) (Cadmium efflux ATPase)
Length = 727
Score = 331 bits (849), Expect = 4e-90
Identities = 224/727 (30%), Positives = 382/727 (51%), Gaps = 50/727 (6%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDV-------------LLIS 56
+ V G CA A E+ +K + GV+ V A + V + L +S
Sbjct: 16 YRVQGFTCANCAGKFEKNVKKIPGVQDAKVNFGASKIDVYGNASVEELEKAGAFENLKVS 75
Query: 57 ESKIAD-ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPP-----LG 110
K+A+ + + + + +T KK +L LL+A +L +
Sbjct: 76 PEKLANQTIQRVKDDTKAHKEEKTPFYKKHSTLLFAT--LLIAFGYLSHFVNGEDNLVTS 133
Query: 111 WLALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQ 170
L +GS+ IG + ++ ++ L+ +AV G + + + +++ LF+I++
Sbjct: 134 MLFVGSIVIGGYSLFKVGFQNLIRFDFDMKTLMTVAVIGATIIGKWAEASIVVILFAISE 193
Query: 171 WLETRATHKALVAMSSLTNMAPQMAIVAETGERV--DVNDVKMNTILAVKAGDAIPLDGI 228
LE + ++ ++ SL ++AP+ A+V G+ + V+D+ + I+ VK G+ I +DGI
Sbjct: 194 ALERFSMDRSRQSIRSLMDIAPKEALVRRNGQEIIIHVDDIAVGDIMIVKPGEKIAMDGI 253
Query: 229 VVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKN 283
+V G V++ +TGES PV+K D V+AGT+N K + DT +I
Sbjct: 254 IVNGLSAVNQAAITGESVPVSKAVDDEVFAGTLNEEGLIEVKITKYVEDTTITKIIHLVE 313
Query: 284 YSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLA 343
+ G+R + ++F K K + V+ + +AVVP + W +
Sbjct: 314 EAQGERAPA--------QAFVDKF--AKYYTPIIMVIAALVAVVPPLFFGGSWDTWVYQG 363
Query: 344 LVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRG 403
L VL+ GCPCAL++STP++I A+ AA G+L+KGG YLE L IKTVAFDKTGT+T+G
Sbjct: 364 LAVLVVGCPCALVISTPISIVSAIGNAAKKGVLVKGGVYLEKLGAIKTVAFDKTGTLTKG 423
Query: 404 EFSVTDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNF 462
VTDF D + + L I+++E +S HP+A A++ A +I VE F +
Sbjct: 424 VPVVTDFEVLNDQVEEKELFSIITALEYRSQHPLASAIMKKAEQDNIPYSNVQVEEFTSI 483
Query: 463 PGEGIFGTIDGRDIYIGNKR------VGVRAICKRDNCEVQFQRPEISTKKNNCEETLVG 516
G GI G ++G YIG+ + V ++ +N ++ Q + E+T++G
Sbjct: 484 TGRGIKGIVNGTTYYIGSPKLFKELNVSDFSLGFENNVKI-LQNQGKTAMIIGTEKTILG 542
Query: 517 VFSLVDACRSGALEAMEELKLLGVR-SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHE 575
V ++ D R + +++L LG++ ++MLTGD+ A + + + + + +EL+P +
Sbjct: 543 VIAVADEVRETSKNVIQKLHQLGIKQTIMLTGDNQGTANAIGTHV--GVSDIQSELMPQD 600
Query: 576 KAKLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRK 634
K I+ + E +AMIGDG+NDAPALA + +GI+MG +G+ A ET++ LM +D+ K
Sbjct: 601 KLDYIKKMQSEYDNVAMIGDGVNDAPALAASTVGIAMGGAGTDTAIETADIALMGDDLSK 660
Query: 635 VPEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSML 694
+P A+RL+RKT + N+ ++G K L L I G+ +W+A+L+D+G +LV LNS+
Sbjct: 661 LPFAVRLSRKTLNIIKANITFAIGIKIIALLLVIPGWLTLWIAILSDMGATILVALNSLR 720
Query: 695 ILQENHK 701
+++ K
Sbjct: 721 LMRVKDK 727
>CAD1_LISMO (P58414) Probable cadmium-transporting ATPase (EC
3.6.3.3) (Cadmium efflux ATPase)
Length = 707
Score = 327 bits (837), Expect = 1e-88
Identities = 203/721 (28%), Positives = 390/721 (53%), Gaps = 51/721 (7%)
Query: 6 KRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALN 65
K++ + V GM C A E+ +K L GV V A ++V + IS+ + A A
Sbjct: 6 KQTTYRVDGMSCTNCAGKFEKNVKNLEGVTDAKVNFGAGKISVYGETS-ISQIEKAGAFE 64
Query: 66 TARL----EASFRPQGETNNEKKCPDILTMACGLLLAL------------SFLKYIYPPL 109
R+ + S +P + + KK ++ ++LA + + Y+
Sbjct: 65 NLRVTDEKDYSSKPAKKESFLKKNWHLVVSIIFIILAFISQNISGEDSTTTIILYVI--- 121
Query: 110 GWLALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIA 169
++ +G + A++ L + L+ +A+ G + + ++ +G +++ LF+ +
Sbjct: 122 ------AIVVGGFNLFKEGFANLIKLDFTMESLMTIAIIGASIIGEWAEGSIVVILFAFS 175
Query: 170 QWLETRATHKALVAMSSLTNMAPQMAIVA--ETGERVDVNDVKMNTILAVKAGDAIPLDG 227
+ LE + KA ++ SL ++AP+ A++ + + + V+D+++ I+ +K G I +DG
Sbjct: 176 EVLERYSMDKARQSIRSLMDIAPKEALIRRDDVEQMIAVSDIQIGDIMIIKPGQKIAMDG 235
Query: 228 IVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCK 282
+V++G +++ +TGES PV K+ D V+AGT+N ++ + DT +I
Sbjct: 236 VVIKGYSAINQSAITGESIPVEKKVDDEVFAGTLNEEGLLEVKVTKHVEDTTISKIIHLV 295
Query: 283 NYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHL 342
+ G+R + ++F K K + ++ + VVP D + W +
Sbjct: 296 EEAQGERAPA--------QAFVDKF--AKYYTPTIMLIALLVVVVPPLFFGGDWDTWVYQ 345
Query: 343 ALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITR 402
L +L+ GCPC+L++STPV+I A+ +A +G+L+KGG YLE + ++ +AFDKTGT+T+
Sbjct: 346 GLSLLVVGCPCSLVISTPVSIVSAIGNSAKNGVLVKGGIYLEEIGGLQAIAFDKTGTLTK 405
Query: 403 GEFSVTDFSA-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQN 461
G+ VTDF + ++ + L I+++E+ S HP+A A++ A + ++ ++NF +
Sbjct: 406 GKPVVTDFIPYSEHMDEQNSLSIITALETMSQHPLASAIISKAMIDNVDYKSIEIDNFSS 465
Query: 462 FPGEGIFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKN---NCEETLVGVF 518
G+G+ G ++G YIG+ ++ ++ K + +Q + K E ++ +
Sbjct: 466 ITGKGVKGEVNGITYYIGSSKLFESSLEKSQSISQTYQSLQKQGKTAMLFGTESNILAII 525
Query: 519 SLVDACRSGALEAMEELKLLGV-RSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKA 577
++ D R + E + +L LG+ ++MLTGD++ A ++ ++ + + AEL+P +K
Sbjct: 526 AVADEVRESSKEVIAQLHKLGIAHTIMLTGDNNDTAQFIGKEI--GVSDIKAELMPEDKL 583
Query: 578 KLIENFKKE-GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVP 636
I+ K+ G +AMIGDG+NDAPALA + +GI+MG +G+ A ET++ LM +D++K+P
Sbjct: 584 TYIKELKQTYGKVAMIGDGVNDAPALAASTVGIAMGGAGTDTALETADVALMGDDLKKLP 643
Query: 637 EAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLIL 696
+ L+RKT + + +N+ S+G K L L + G+ +W+A++ D+G LLV LN + ++
Sbjct: 644 FIVNLSRKTLKIIKQNITFSLGIKLLALLLVLPGWLTLWIAIVADMGATLLVTLNGLRLM 703
Query: 697 Q 697
+
Sbjct: 704 K 704
>ATZN_SYNY3 (Q59998) Zinc-transporting ATPase (EC 3.6.3.5)
(Zn(II)-translocating P-type ATPase)
Length = 721
Score = 293 bits (751), Expect = 1e-78
Identities = 217/736 (29%), Positives = 358/736 (48%), Gaps = 68/736 (9%)
Query: 5 IKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD-- 62
+K +V GM C + +E L+ L GV SV V +TV +D +SE I +
Sbjct: 7 LKTQQMQVGGMDCTSCKLKIEGSLERLKGVAEASVTVATGRLTVTYDPKQVSEITIQERI 66
Query: 63 --------------ALNTARLEASFRPQG--------ETNNEKKCPDILTMACGLLLALS 100
LN + S R +G E N +++ +LT +A+
Sbjct: 67 AALGYTLAEPKSSVTLNGHKHPHSHREEGHSHSHGAGEFNLKQELLPVLTAIALFTIAIL 126
Query: 101 FLKYIYPPLGWLALGSVAIGFPKVLFRAIASIRALTLNI--------NILVLLAVCGTAA 152
F + ++ G +A +V I P L ++ NI N L+ +A G A
Sbjct: 127 FEQPLHNTPGQIAEFAVII--PAYLLSGWTVLKTAGRNILRGQIFDENFLMTIATLGALA 184
Query: 153 LQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETG--ERVDVNDVK 210
+ + ++ F + + + + ++ ++ +L P A + G ++V V+
Sbjct: 185 IHQLPEAVAVMLFFRVGELFQEYSVGRSRRSIKALLEARPDTANLKRNGTVQQVSPETVQ 244
Query: 211 MNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY---- 266
++ ++ VK G+ +PLDG ++ G +VD LTGES P T + + AG IN +
Sbjct: 245 VDDLILVKPGEKVPLDGEILGGTSQVDTSALTGESVPGTVKPGDTILAGMINQSGVLTIR 304
Query: 267 -PKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIA 325
K+ ++ ++ + K+ S+ K + R + ++ + +A
Sbjct: 305 VTKLFSESSIAKVLDLVENASSKKASTEKFITQFARYYTPVIVFLSL----------AVA 354
Query: 326 VVPAALSVP--DMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYL 383
++P L +P D W + ALV+L+ CPC L++S P+ F + AA G+L+KG +L
Sbjct: 355 LLPP-LFIPGADRADWVYRALVLLVISCPCGLVISIPLGYFGGIGGAAKHGILIKGSTFL 413
Query: 384 ETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAGALVDY 443
++L+ +KTV FDKTGT+T+G F VT + + LL + ES S+HP+A ++ +
Sbjct: 414 DSLTAVKTVVFDKTGTLTKGTFKVTQVVTKNGFSESELLTLAAKAESHSTHPIALSIRE- 472
Query: 444 ARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEI 503
A SI V +++ G GI + + + GN R+ R D C+V +
Sbjct: 473 AYAQSI--ADSEVADYEEIAGHGIRAVVQNQVVIAGNDRLLHREKIDHDTCDVAGTVVHL 530
Query: 504 STKKNNCEETLVGVFSLVDACRSGALEAMEELKLLGV-RSVMLTGDSSQVAMYVQSQLKN 562
+ + G + D + A++A+ +LK +GV ++VMLTGDS VA V Q+
Sbjct: 531 AV-----DGRYGGYILIADEIKEDAVQAIRDLKRMGVEKTVMLTGDSEIVAQSVAQQI-- 583
Query: 563 AIDIVHAELLPHEKAKLIENF---KKEGPIAMIGDGINDAPALATADIGISMGISGSALA 619
+D AELLP EK IE + +A +GDGINDAP +A AD+GI+MG GS A
Sbjct: 584 GLDAFVAELLPEEKVDEIEQLLDPSGKAKLAFVGDGINDAPVIARADVGIAMGGLGSDAA 643
Query: 620 NETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVL 679
ET++ +LM++ KV EAI +ARKT + +V+N+++++G K +AL G +W AV
Sbjct: 644 IETADVVLMTDAPSKVAEAIHVARKTRQIVVQNIVLALGIKALFIALGTIGLATLWEAVF 703
Query: 680 TDVGTCLLVILNSMLI 695
DVG LL ILN+ I
Sbjct: 704 ADVGVALLAILNATRI 719
>ATZN_ECOLI (P37617) Lead, cadmium, zinc and mercury transporting
ATPase (EC 3.6.3.3) (EC 3.6.3.5)
Length = 732
Score = 275 bits (704), Expect = 3e-73
Identities = 215/712 (30%), Positives = 353/712 (49%), Gaps = 41/712 (5%)
Query: 3 ENIK--RSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVV--HDVLLISES 58
EN+ R +++V GM CA A VE ++ L GV V V+ + V +D+ ES
Sbjct: 43 ENVSGTRYSWKVSGMDCAACARKVENAVRQLAGVNQVQVLFATEKLVVDADNDIRAQVES 102
Query: 59 KIADALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSF-LKYIYPPLGWLA-LGS 116
+ A + R E + + ++ P I + +++A+S+ L+ P G LA + +
Sbjct: 103 ALQKAGYSLRDEQAAEEPQASRLKENLPLITLI---VMMAISWGLEQFNHPFGQLAFIAT 159
Query: 117 VAIGFPKVLFRAIASIRALT-LNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETR 175
+G + +A+ I++ + I L+ +A G + + +++ LF I + LE
Sbjct: 160 TLVGLYPIARQALRLIKSGSYFAIETLMSVAAIGALFIGATAEAAMVLLLFLIGERLEGW 219
Query: 176 ATHKALVAMSSLTNMAPQMAIVAETGER--VDVNDVKMNTILAVKAGDAIPLDGIVVEGK 233
A +A +S+L + P+ A GER V +N ++ ++ V AG +P DG ++
Sbjct: 220 AASRARQGVSALMALKPETATRLRKGEREEVAINSLRPGDVIEVAAGGRLPADGKLLSPF 279
Query: 234 CEVDEKMLTGESFPVTKESDSLVWAGTINMNAYPKIIF------DTYDFRIYKCKNYSVG 287
DE LTGES PV + + V AG +++ + D RI K +
Sbjct: 280 ASFDESALTGESIPVERATGDKVPAGATSVDRLVTLEVLSEPGASAID-RILKLIEEAEE 338
Query: 288 KRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVL 347
+R + + R + +++ + + +VP L + W + L +L
Sbjct: 339 RRAPIERFIDRFSRIYTPAIMAVAL----------LVTLVPPLLFAASWQEWIYKGLTLL 388
Query: 348 LSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSV 407
L GCPCAL++STP AI L AA G L+KGG LE L R+ VAFDKTGT+T G+ V
Sbjct: 389 LIGCPCALVISTPAAITSGLAAAARRGALIKGGAALEQLGRVTQVAFDKTGTLTVGKPRV 448
Query: 408 TDFSAGDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGI 467
T I+ LL +++E ++HP+A A+V A++ + +P E+ + G GI
Sbjct: 449 TAIHPATGISESELLTLAAAVEQGATHPLAQAIVREAQVAEL-AIP-TAESQRALVGSGI 506
Query: 468 FGTIDGRDIYI--GNKRVGVRAICKRDNCEVQFQRPEISTKKNNCEETLVGVFSLVDACR 525
++G + I K + E Q + + ++ ++GV +L D R
Sbjct: 507 EAQVNGERVLICAAGKHPADAFTGLINELESAGQTVVLVVRNDD----VLGVIALQDTLR 562
Query: 526 SGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK 585
+ A A+ EL LGV+ V+LTGD+ + A + +L A LLP +K K + +
Sbjct: 563 ADAATAISELNALGVKGVILTGDNPRAAAAIAGELGLEF---KAGLLPEDKVKAVTELNQ 619
Query: 586 EGPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKT 645
P+AM+GDGINDAPA+ A IGI+MG SG+ +A ET++A L N +R + + I LAR T
Sbjct: 620 HAPLAMVGDGINDAPAMKAAAIGIAMG-SGTDVALETADAALTHNHLRGLVQMIELARAT 678
Query: 646 TRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
+ +N+ I++G K L + G +WLAVL D G +LV N++ +L+
Sbjct: 679 HANIRQNITIALGLKGIFLVTTLLGMTGLWLAVLADTGATVLVTANALRLLR 730
>HMCT_HELPY (Q59465) Cadmium, zinc and cobalt transporting ATPase
(EC 3.6.3.3) (EC 3.6.3.5)
Length = 686
Score = 259 bits (663), Expect = 2e-68
Identities = 176/576 (30%), Positives = 302/576 (51%), Gaps = 52/576 (9%)
Query: 140 NILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAE 199
N L+L+A + + + I+ +S ++L+ A ++ ++ +L ++AP +A + +
Sbjct: 136 NALMLIATIAAFFVGAYEESVSIMVFYSAGEFLQKLAVSRSKKSLKALVDVAPNLAYLKK 195
Query: 200 TGERVDV--NDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVW 257
E V V D+++N I+ VK G+ +P+DG+VV+G+ +DE+ L+GES PV +S V
Sbjct: 196 GDELVSVAPEDLRVNDIVVVKVGEKVPVDGVVVKGESLLDERALSGESMPVNVSENSKVL 255
Query: 258 AGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FR 317
G++N+ A +I + ++YK + S+ K + T +S +K I TK R +
Sbjct: 256 GGSLNLKAVLEIQVE----KMYK--DSSIAKVVDLVQQA-TNEKSETEKFI-TKFSRYYT 307
Query: 318 -KVLYSC--IAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISG 374
VL+ IAV+P S+ + W + LV L+ CPCAL++S P+ F + A+ G
Sbjct: 308 PSVLFIALMIAVLPPLFSMGSFDEWIYRGLVALMVSCPCALVISVPLGYFGGVGAASRKG 367
Query: 375 LLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSH 434
+L+KG LE L++ K++AFDKTGT+T+G F VTD + + E +L++ S + S+H
Sbjct: 368 ILMKGVHVLEVLTQAKSIAFDKTGTLTKGVFKVTDIVPQNGHSKEEVLHYASCSQLLSTH 427
Query: 435 PVA--------GALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRVGVR 486
P+A L D H IK N++ G G+ I GN+++
Sbjct: 428 PIALSIQKACEEMLKDDKHQHDIK-------NYEEVSGMGVKAQCHTDLIIAGNEKM--- 477
Query: 487 AICKRDNCEVQFQRPEISTKKNNC------EETLVGVFSLVDACRSGALEAMEELKLLGV 540
QF +K+N +T VG + D + A+E + +LK+ G+
Sbjct: 478 --------LDQFHIAHSPSKENGTIVHVAFNQTYVGYIVISDEIKDDAIECLRDLKVQGI 529
Query: 541 RS-VMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK--EGPIAMIGDGIN 597
+ +L+GD + L HA LLP EK + + FK+ + P +GDGIN
Sbjct: 530 ENFCILSGDRKSATESIAQTLGCE---YHASLLPEEKTSVFKTFKERYKAPAIFVGDGIN 586
Query: 598 DAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISV 657
DAP LA+AD+GI MG GS L+ ++++ ++ ++ + + + + +A+KT + +N++ ++
Sbjct: 587 DAPTLASADVGIGMG-KGSELSKQSADIVITNDSLNSLVKVLAIAKKTKSIIWQNILFAL 645
Query: 658 GFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSM 693
G K + L + G +W AV DVG LL + NSM
Sbjct: 646 GIKAVFIVLGLMGVASLWEAVFGDVGVTLLALANSM 681
>HMCT_HELPJ (Q9ZL53) Cadmium, zinc and cobalt transporting ATPase
(EC 3.6.3.3) (EC 3.6.3.5)
Length = 686
Score = 251 bits (641), Expect = 6e-66
Identities = 172/572 (30%), Positives = 298/572 (52%), Gaps = 44/572 (7%)
Query: 140 NILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAE 199
N L+L+A + + + I+ +S ++L+ A ++ ++ +L ++AP +A + +
Sbjct: 136 NALMLIATIAAFCVGAYEESVSIMVFYSAGEFLQILAIARSKKSLKALVDVAPNLAYLKK 195
Query: 200 TGERVDV--NDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVW 257
V V D+++N I+ VK G+ +P+DG+V++G+ +DE+ L+GES PV S V
Sbjct: 196 GDTLVSVAPEDLRINDIVVVKVGEKVPVDGVVIKGESLLDERALSGESMPVNVSERSKVL 255
Query: 258 AGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FR 317
G++N+ A +I + ++YK + S+ K + T +S +K I TK R +
Sbjct: 256 GGSLNLKAVLEIQVE----KLYK--DSSIAKVVDLVQQA-TNEKSETEKFI-TKFSRYYT 307
Query: 318 -KVLYSC--IAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISG 374
VL+ IAV+P S+ + W + LV L+ CPCAL++S P+ F + A+ G
Sbjct: 308 PSVLFIALMIAVLPPLFSMGSFDEWIYRGLVALMVSCPCALVISVPLGYFGGVGAASRKG 367
Query: 375 LLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSH 434
+L+KG LE L++ K++AFDKTGT+T+G F V D + + E +L++ S + S+H
Sbjct: 368 ILMKGVHVLEVLTQTKSIAFDKTGTLTKGVFKVVDIVPQNGHSKEEVLHYASCSQLLSTH 427
Query: 435 PVA--------GALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRVGVR 486
P+A L D H IK N++ G G+ I GN+++
Sbjct: 428 PIALSIQKACEEMLKDDKHQHDIK-------NYEELSGMGVKAQCHTDLIIAGNEKM--- 477
Query: 487 AICKRDNCEVQFQ-RPEISTKKNNC-EETLVGVFSLVDACRSGALEAMEELKLLGVRS-V 543
D + PE T + +T +G + D + A+E + +LK G+ +
Sbjct: 478 ----LDQFHIAHSPSPENGTIVHVAFNQTYIGYIVISDEIKDDAIECLRDLKAQGIENFC 533
Query: 544 MLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK--EGPIAMIGDGINDAPA 601
+L+GD + L HA LLP EK + + FK+ + P +GDGINDAP
Sbjct: 534 ILSGDRKSATESIARTLGCE---YHASLLPEEKTSVFKTFKERYKAPAIFVGDGINDAPT 590
Query: 602 LATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKC 661
LA+AD+GI MG GS L+ ++++ ++ ++ + + + + +A+KT + +N++ ++G K
Sbjct: 591 LASADVGIGMG-KGSELSKQSADIVITNDSLSSLVKVLAIAKKTKSIIWQNILFALGIKA 649
Query: 662 AILALAIAGYPLVWLAVLTDVGTCLLVILNSM 693
+ L + G +W AV DVG LL + NSM
Sbjct: 650 VFIVLGLMGVASLWEAVFGDVGVTLLALANSM 681
>CTPG_MYCTU (P63689) Probable cation-transporting ATPase G (EC
3.6.3.-)
Length = 771
Score = 251 bits (640), Expect = 7e-66
Identities = 187/658 (28%), Positives = 320/658 (48%), Gaps = 51/658 (7%)
Query: 64 LNTARLEASFRPQGETN--------NEKKCPDILTMAC--------GLLLALSFLKYIYP 107
L+ R + + +P GET+ NE + P+ L G+LL S +
Sbjct: 116 LSGVRRDVAAQPSGETSDACCDGEDNEDREPEQLWQVAKLRRAAFSGVLLTASLVAAWAY 175
Query: 108 PLGWLALG----SVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVII 163
PL + LG ++A+G + ++ + + + L+ +A G AL + + +
Sbjct: 176 PLWPVVLGLKALALAVGASTFVPSSLKRLAEGRVGVGTLMTIAALGAVALGELGEAATLA 235
Query: 164 FLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETGERVDVNDVKMNT--ILAVKAGD 221
FLFSI++ LE AT + + +L ++ P A V G V +++ + VK G+
Sbjct: 236 FLFSISEGLEEYATARTRRGLRALLSLVPDQATVLREGTETIVASTELHVGDQMIVKPGE 295
Query: 222 AIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKC 281
+ DGI+ G+ +D +TGES PV V+AG+IN + +
Sbjct: 296 RLATDGIIRAGRTALDVSAITGESVPVEVGPGDEVFAGSIN------------GLGVLQV 343
Query: 282 KNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALS-----VPDM 336
+ +S + V +K S ++ + L I + A ++ + +
Sbjct: 344 GVTATAANNSLARIVHIVEAEQVRKGASQRLADCIARPLVPSIMIAAALIAGTGSVLGNP 403
Query: 337 EPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDK 396
W ALVVL++ PCAL ++ PV + ++ A+ G+L+KGG LETL I+ VA DK
Sbjct: 404 LVWIERALVVLVAAAPCALAIAVPVTVVASIGAASRLGVLIKGGAALETLGTIRAVALDK 463
Query: 397 TGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENV 456
TGT+T V D + + E +L +++E++S HP+A A++ + +
Sbjct: 464 TGTLTANRPVVIDVATTNGATREEVLAVAAALEARSEHPLAVAVLAATQATTA------A 517
Query: 457 ENFQNFPGEGIFGTIDGRDIYIGNKR-VGVRAICKRDNCEVQFQRPEISTKKNNCEETLV 515
+ Q PG G+ G +DGR + +G + + C Q + ++ ++ L+
Sbjct: 518 SDVQAVPGAGLIGRLDGRVVRLGRPGWLDAAELADHVACMQQAGATAVLVER---DQQLL 574
Query: 516 GVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHE 575
G ++ D R A E + L+ G + MLTGD+ A + +Q I+ VHAEL P +
Sbjct: 575 GAIAVRDELRPEAAEVVAGLRTGGYQVTMLTGDNHATAAALAAQA--GIEQVHAELRPED 632
Query: 576 KAKLIENFKKEGPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKV 635
KA L+ + P AM+GDG+NDAPALA AD+GI+MG G+ +A ET++ LM D+R +
Sbjct: 633 KAHLVAQLRARQPTAMVGDGVNDAPALAAADLGIAMGAMGTDVAIETADVALMGQDLRHL 692
Query: 636 PEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSM 693
P+A+ AR++ + +V+NV +S+ ++ LA+ G + VL T ++VI N +
Sbjct: 693 PQALDHARRSRQIMVQNVGLSLSIITVLMPLALFGILGLAAVVLVHEFTEVIVIANGV 750
>CTPG_MYCBO (P63690) Probable cation-transporting ATPase G (EC
3.6.3.-)
Length = 771
Score = 251 bits (640), Expect = 7e-66
Identities = 187/658 (28%), Positives = 320/658 (48%), Gaps = 51/658 (7%)
Query: 64 LNTARLEASFRPQGETN--------NEKKCPDILTMAC--------GLLLALSFLKYIYP 107
L+ R + + +P GET+ NE + P+ L G+LL S +
Sbjct: 116 LSGVRRDVAAQPSGETSDACCDGEDNEDREPEQLWQVAKLRRAAFSGVLLTASLVAAWAY 175
Query: 108 PLGWLALG----SVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVII 163
PL + LG ++A+G + ++ + + + L+ +A G AL + + +
Sbjct: 176 PLWPVVLGLKALALAVGASTFVPSSLKRLAEGRVGVGTLMTIAALGAVALGELGEAATLA 235
Query: 164 FLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETGERVDVNDVKMNT--ILAVKAGD 221
FLFSI++ LE AT + + +L ++ P A V G V +++ + VK G+
Sbjct: 236 FLFSISEGLEEYATARTRRGLRALLSLVPDQATVLREGTETIVASTELHVGDQMIVKPGE 295
Query: 222 AIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKC 281
+ DGI+ G+ +D +TGES PV V+AG+IN + +
Sbjct: 296 RLATDGIIRAGRTALDVSAITGESVPVEVGPGDEVFAGSIN------------GLGVLQV 343
Query: 282 KNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALS-----VPDM 336
+ +S + V +K S ++ + L I + A ++ + +
Sbjct: 344 GVTATAANNSLARIVHIVEAEQVRKGASQRLADCIARPLVPSIMIAAALIAGTGSVLGNP 403
Query: 337 EPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDK 396
W ALVVL++ PCAL ++ PV + ++ A+ G+L+KGG LETL I+ VA DK
Sbjct: 404 LVWIERALVVLVAAAPCALAIAVPVTVVASIGAASRLGVLIKGGAALETLGTIRAVALDK 463
Query: 397 TGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENV 456
TGT+T V D + + E +L +++E++S HP+A A++ + +
Sbjct: 464 TGTLTANRPVVIDVATTNGATREEVLAVAAALEARSEHPLAVAVLAATQATTA------A 517
Query: 457 ENFQNFPGEGIFGTIDGRDIYIGNKR-VGVRAICKRDNCEVQFQRPEISTKKNNCEETLV 515
+ Q PG G+ G +DGR + +G + + C Q + ++ ++ L+
Sbjct: 518 SDVQAVPGAGLIGRLDGRVVRLGRPGWLDAAELADHVACMQQAGATAVLVER---DQQLL 574
Query: 516 GVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHE 575
G ++ D R A E + L+ G + MLTGD+ A + +Q I+ VHAEL P +
Sbjct: 575 GAIAVRDELRPEAAEVVAGLRTGGYQVTMLTGDNHATAAALAAQA--GIEQVHAELRPED 632
Query: 576 KAKLIENFKKEGPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKV 635
KA L+ + P AM+GDG+NDAPALA AD+GI+MG G+ +A ET++ LM D+R +
Sbjct: 633 KAHLVAQLRARQPTAMVGDGVNDAPALAAADLGIAMGAMGTDVAIETADVALMGQDLRHL 692
Query: 636 PEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSM 693
P+A+ AR++ + +V+NV +S+ ++ LA+ G + VL T ++VI N +
Sbjct: 693 PQALDHARRSRQIMVQNVGLSLSIITVLMPLALFGILGLAAVVLVHEFTEVIVIANGV 750
>HMCT_HELFE (Q9RQB4) Cadmium, zinc and cobalt transporting ATPase
(EC 3.6.3.3) (EC 3.6.3.5)
Length = 681
Score = 242 bits (618), Expect = 3e-63
Identities = 157/564 (27%), Positives = 287/564 (50%), Gaps = 33/564 (5%)
Query: 140 NILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAI--V 197
N L+L A + + I+ +S ++L+ A ++ ++ +L ++AP A+ V
Sbjct: 136 NTLMLSATIAAFGVGAHEEAVSIMVFYSAGEFLQQLAIARSKQSLHALLDVAPNHAVRKV 195
Query: 198 AETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVW 257
+ + V+ + + + ++ VK G+ IP DG+V+ G+ +D+K LTGES PV + S V
Sbjct: 196 GQELQEVEPSALAIGDVVIVKVGEKIPTDGVVLRGESLLDQKALTGESLPVNVKEHSAVL 255
Query: 258 AGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKI 312
G+IN+ K+ D+ +I + +V ++ + K + T R + V + +
Sbjct: 256 GGSINLKGVLEIQVSKLYADSSVAKIVDLVSNAVSQKSQTEKFITTFARYYTPAVFAIAL 315
Query: 313 HR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAI 372
IA+VP L D + W + L L+ CPCAL++S P+ F + A+
Sbjct: 316 ----------LIALVPPLLGHGDFDTWIYRGLFALMVSCPCALVISVPLGYFGGVGAASR 365
Query: 373 SGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKS 432
+G+L+K LE LS++K +AFDKTGT+++GEF+V D + E +L + + S
Sbjct: 366 AGILIKSVQTLEALSQVKNIAFDKTGTLSKGEFNVIDVVPIAPFSKEDVLQHATCAQILS 425
Query: 433 SHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRVGVRAICKRD 492
+HP+A ++ + + V+++Q G G+ + I GN ++ + D
Sbjct: 426 THPIAISIQKAYK----RQCQHEVKDYQEIGGLGVQASCHSHLIIAGNDKMLHKYNIPHD 481
Query: 493 NCEVQFQRPEISTKKNNCEETLVGVFSLVDACRSGALEAMEELKLLGVRSV-MLTGDSSQ 551
C ++ ++ + +G + D + A E ++ LK G+ + +L+GD
Sbjct: 482 TCSLEGTIVHVAVDGKH-----IGYIVVADTLKDNAKECLDGLKHAGIEHMCILSGDHEY 536
Query: 552 VAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKE--GPIAMIGDGINDAPALATADIGI 609
V +L +A LLP +K ++F+ + +GDGINDAP LA AD+ +
Sbjct: 537 STKRVAKELDCP---YYANLLPEDKLNAFKDFQAQHAHKSMFVGDGINDAPTLARADVSM 593
Query: 610 SMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIA 669
SMG S S ++ E+++ ++ +N + V + ++A+KT R ++EN+I ++ K + L ++
Sbjct: 594 SMG-SASQISKESADIVITNNSLESVLKVFKIAKKTKRIIIENIIFALAIKAMFIVLGLS 652
Query: 670 GYPLVWLAVLTDVGTCLLVILNSM 693
G +W AVL DVG L+ + NSM
Sbjct: 653 GDASLWEAVLGDVGVTLIALANSM 676
>ATC1_RHIME (P58341) Copper-transporting ATPase 1 (EC 3.6.3.4)
Length = 826
Score = 242 bits (618), Expect = 3e-63
Identities = 211/727 (29%), Positives = 347/727 (47%), Gaps = 83/727 (11%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVH----DVLLISESKIADALN 65
F + GM CA+ + VE+ L+ + GV SV + TV DV I E+ + DA
Sbjct: 86 FGIEGMTCASCVSRVEKALRTVPGVADASVNLATEKGTVRFVSGVDVAAI-EAAVRDAGY 144
Query: 66 TARLE----ASFRPQGETNNEKKCPDILTMACGLLLALSFL------------KYIYPPL 109
R A+ P+ E + L + +L FL ++I +
Sbjct: 145 DVRKAKASGATAEPEDRRELETRTLKRLVILSAVLTLPLFLVEMGSHFMPGVHEWIMENI 204
Query: 110 GW-------LALGSVAIGFPKVLF--RAIASIRALTLNINILVLL---AVCGTAALQDFT 157
G AL + + P + F + + ++ T ++N LV+L A G + + F
Sbjct: 205 GMRHNLYIQFALATAVLFGPGLRFFRKGVPNLLRWTPDMNSLVVLGTTAAWGYSVVATFA 264
Query: 158 DG--------------GVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETGER 203
G VI+ L + ++LE RA + A+ L + P+ A VA E
Sbjct: 265 SGLLPSGTANVYYEAAAVIVTLILLGRYLEARAKGRTSQAIKRLLGLQPKTAFVAHGDEF 324
Query: 204 VDV--NDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTI 261
V++ +DV + ++ ++ G+ IP+DG V++G VDE M+TGE PV K + + V GTI
Sbjct: 325 VEIQISDVVVGDVIRIRPGEKIPVDGTVLDGNSYVDESMITGEPVPVQKAAGAEVVGGTI 384
Query: 262 NMNA-----YPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*F 316
N N K+ DT +I K + G + + + K+ F
Sbjct: 385 NKNGSFTFRATKVGGDTLLAQIIKMVETAQGSKLPI-------------QALVDKVTAWF 431
Query: 317 RKVLYSCIAVVPAALSVPDMEPWFHLALV----VLLSGCPCALILSTPVAIFCALTKAAI 372
+ + AA V P ALV VL+ CPCA+ L+TP +I +AA
Sbjct: 432 VPAVILVAVLTFAAWYVFGPSPALTFALVNAVAVLIIACPCAMGLATPTSIMVGTGRAAE 491
Query: 373 SGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKS 432
G+L + G+ L++L +A DKTGT+T+G +TD D + +L +++S+E+ S
Sbjct: 492 LGILFRKGEALQSLREADVIALDKTGTLTKGRPELTDIVPADGFEADEVLSFVASLEALS 551
Query: 433 SHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRVGVRAICKRD 492
HP+A A+V A+ I VP +F+ PG G+ G + G + +G R
Sbjct: 552 EHPIAEAIVSAAKSRGIALVP--ATDFEATPGFGVRGAVSGLPVQVGADRAFSGVGIDVS 609
Query: 493 NCEVQFQRPEISTKK---NNCEETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDS 549
V+ +R S K + L + ++ D + +A++ L LG++ M+TGD+
Sbjct: 610 PFVVEAERLGNSGKSPLYAAIDGRLAAIIAVSDPIKDTTPQAIKALHDLGLKVAMITGDN 669
Query: 550 SQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEG-PIAMIGDGINDAPALATADIG 608
+ A + QL ID V AE+LP K ++ ++ G +A IGDGINDAPAL AD+G
Sbjct: 670 RRTADAIARQL--GIDEVVAEVLPDGKVDAVKRLREGGRKVAFIGDGINDAPALTEADVG 727
Query: 609 ISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAI 668
I++G +G+ +A E+++ +LMS D+ VP+AI L++ T R + +N+ + + +++ +A
Sbjct: 728 IAVG-TGTDIAIESADVVLMSGDLIGVPKAIALSKATIRNIKQNLFWAFAYNVSLVPVA- 785
Query: 669 AG--YPL 673
AG YPL
Sbjct: 786 AGVLYPL 792
>CTPC_MYCTU (P96875) Probable cation-transporting P-type ATPase C
(EC 3.6.3.-) (Metal transporting ATPase Mta72)
Length = 718
Score = 215 bits (548), Expect = 3e-55
Identities = 198/713 (27%), Positives = 335/713 (46%), Gaps = 52/713 (7%)
Query: 16 CCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALN-TARLEASFR 74
C + A VE + +GV+ V +V V + + + A+ A + A
Sbjct: 22 CDSRRAVAVEEAVAKQNGVRVVHAYPRTGSVVVWYSPRRADRAAVLAAIKGAAHVAAELI 81
Query: 75 PQGETNN-EKKCPDILTMACG--LLLALSFLKYIY--PPL-----GWLALG-SVAIGFPK 123
P ++ E + D+L M G L L +Y++ PPL +A G ++ G+P
Sbjct: 82 PARAPHSAEIRNTDVLRMVIGGVALALLGVRRYVFARPPLLGTTGRTVATGVTIFTGYP- 140
Query: 124 VLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVA 183
L A+ S+R+ + LV A + L++ +++L +I ++L+ + A
Sbjct: 141 FLRGALRSLRSGKAGTDALVSAATVASLILRENVVALTVLWLLNIGEYLQDLTLRRTRRA 200
Query: 184 MSSLTNMAPQMAIV----------AETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGK 233
+S L A V A T +V ++ V++ + V AIP+DG VV+G+
Sbjct: 201 ISELLRGNQDTAWVRLTDPSAGSDAATEIQVPIDTVQIGDEVVVHEHVAIPVDGEVVDGE 260
Query: 234 CEVDEKMLTGESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSG 293
V++ +TGE+ PV+ + V AG++ + + + ++VG + + G
Sbjct: 261 AIVNQSAITGENLPVSVVVGTRVHAGSVVVRGRVVV------------RAHAVGNQTTIG 308
Query: 294 KNVKTC*RSF*QKVISTKIHR*F--RKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGC 351
+ + + + + F R V S I A L D+ A+ +LL C
Sbjct: 309 RIISRVEEAQLDRAPIQTVGENFSRRFVPTSFIVSAIALLITGDV----RRAMTMLLIAC 364
Query: 352 PCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFS 411
PCA+ LSTP AI A+ A G+L+KGG +LE R+ + FDKTGT+T G VT+
Sbjct: 365 PCAVGLSTPTAISAAIGNGARRGILIKGGSHLEQAGRVDAIVFDKTGTLTVGRPVVTNIV 424
Query: 412 A-GDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGT 470
A D E +L + +S E S HP+A A++ I P E + G G+
Sbjct: 425 AMHKDWEPEQVLAYAASSEIHSRHPLAEAVIRSTEERRISIPPH--EECEVLVGLGMRTW 482
Query: 471 IDGRDIYIGN----KRVGVRAICKRDNCEVQFQRPEISTKKNNCEETLVGVFSLVDACRS 526
DGR + +G+ + VR K + +R + + TLVG+ SL D R
Sbjct: 483 ADGRTLLLGSPSLLRAEKVRVSKKASEWVDKLRRQAETPLLLAVDGTLVGLISLRDEVRP 542
Query: 527 GALEAMEELKLLGVRS-VMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK 585
A + + +L+ G+R VMLTGD ++A V +L ID AE++P +K + +
Sbjct: 543 EAAQVLTKLRANGIRRIVMLTGDHPEIAQVVADEL--GIDEWRAEVMPEDKLAAVRELQD 600
Query: 586 EG-PIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK 644
+G + M+GDGINDAPALA ADIGI+MG++G+ +A ET++ L ++D+ ++ + L +
Sbjct: 601 DGYVVGMVGDGINDAPALAAADIGIAMGLAGTDVAVETADVALANDDLHRLLDVGDLGER 660
Query: 645 TTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
+ +N +S+ A L + G LA + + + V+ NS +++
Sbjct: 661 AVDVIRQNYGMSIAVNAAGLLIGAGGALSPVLAAILHNASSVAVVANSSRLIR 713
>CTPC_MYCLE (Q9CCL1) Probable cation-transporting P-type ATPase C
(EC 3.6.3.-)
Length = 725
Score = 215 bits (547), Expect = 4e-55
Identities = 196/707 (27%), Positives = 333/707 (46%), Gaps = 56/707 (7%)
Query: 21 ATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALN-TARLEASFRP-QGE 78
A VE + +GV+ V +V V + I A++ A + A P +
Sbjct: 41 AVTVEEAIAKCNGVRVVHAYPRTGSVVVWYSPRCCDRQSILAAISGAAHVAAELIPTRAP 100
Query: 79 TNNEKKCPDILTMACGL--LLALSFLKYIY------PPLGWLALGSVAI--GFPKVLFRA 128
+++ + ++L MA G L L +Y++ P L V I G+P +
Sbjct: 101 HSSDIRNIEVLRMAIGAAALTLLGVRRYVFARPLLLPTTSRLVASGVTIFTGYPFLR--- 157
Query: 129 IASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLET---RATHKALVAMS 185
++R + LV +A + L++ +++L +I ++L+ R T +A+ A+
Sbjct: 158 -GALRFGKTGTDALVSVATIASLILRENVVALAVLWLLNIGEYLQDLTLRRTRRAISALL 216
Query: 186 SLTNMAPQMAIV----AETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKML 241
S T + + A T +V + V++ + V AIP+DG V++G+ V++ +
Sbjct: 217 SGTQDTAWIRLTDGPQAGTEIQVPIGTVQIGDEVVVHEHVAIPVDGEVIDGEAVVNQSAI 276
Query: 242 TGESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*R 301
TGE+ PV+ + S V AG++ + + + +VGK + G+ V
Sbjct: 277 TGENLPVSVMAGSHVHAGSVVVRGRLMV------------RASAVGKHTTIGRIVTRVEE 324
Query: 302 SF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPV 361
+ + + F + VV A + + VLL CPCA+ L+TP
Sbjct: 325 AQHDRAPIQTVGENFSRCFVPTSFVVSAITLAITKD--VRRTMTVLLIACPCAVGLATPT 382
Query: 362 AIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSA-GDDINNET 420
AI A+ A G+L+KGG +LE R+ + FDKTGT+T G VT+ A D + E
Sbjct: 383 AISAAIGNGARRGILIKGGSHLEQAGRVDAILFDKTGTLTVGRPVVTNIVAMHKDWSPEQ 442
Query: 421 LLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGN 480
+L + +S E S HP+A A++ I P E + G G+ DGR + +G+
Sbjct: 443 VLAYAASSEIHSRHPLAEAVIRSTEERHISIPPH--EECEVLVGLGMRTWADGRTLLLGS 500
Query: 481 ------KRVGVRAICKR--DNCEVQFQRPEISTKKNNCEETLVGVFSLVDACRSGALEAM 532
++V V D Q + P + + TLVG+ SL D R A E +
Sbjct: 501 PSLLCAEKVKVSKTASEWVDKLRHQTETPLLFA----VDGTLVGLISLRDEVRPEAAEVL 556
Query: 533 EELKLLGVRS-VMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEG-PIA 590
+L+ GVR VMLTGD +A V ++L ID AE++P +K K++ + + EG +
Sbjct: 557 TKLRASGVRRIVMLTGDHPDIAKAVATEL--GIDEWRAEVMPEDKLKVVRDLQNEGYVVG 614
Query: 591 MIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLV 650
M+GDG+NDAPALA ADIGI+MG++G+ +A ET++ L ++D+ ++ + L + +
Sbjct: 615 MVGDGVNDAPALAAADIGIAMGLAGTDVAVETADVALANDDLNRLLDVRDLGGRAVEVIR 674
Query: 651 ENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
EN +S+ A L + G LA + + + V+ NS +++
Sbjct: 675 ENYGMSIAVNAAGLFIGAGGALSPVLAAVLHNASSVAVVANSSRLIR 721
>ATCU_YERPE (Q8ZCA7) Copper-transporting P-type ATPase (EC 3.6.3.4)
Length = 961
Score = 213 bits (543), Expect = 1e-54
Identities = 156/511 (30%), Positives = 259/511 (50%), Gaps = 39/511 (7%)
Query: 161 VIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMA-IVAETGERV-DVNDVKMNTILAVK 218
+II L ++ +E RA ++ A+ L ++AP A +V + GE+V + DV++ IL +
Sbjct: 418 MIIGLINLGHAMEQRARQRSSNALERLLDLAPPTAKLVTDDGEKVIPLADVQLGMILRLT 477
Query: 219 AGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRI 278
GD +P+DG +V+G+ +DE MLTGE P K +V AGT ++ T FR
Sbjct: 478 TGDRVPVDGEIVQGEVWMDEAMLTGEPIPQQKSVGDIVHAGT-------QVQDGTVQFRA 530
Query: 279 YKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSV----- 333
++G + + + +K ++ K K+ V + V+ +
Sbjct: 531 S-----AIGSQTTLARIIKLVRQAQSSKPEIGKLADRISAVFVPTVVVIAIVAGLIWYFF 585
Query: 334 ---PDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIK 390
P + +A VL+ CPCAL L+TP++I + +AA G+L++ D L+ S +
Sbjct: 586 GPQPQLVYTLVVATTVLIIACPCALGLATPMSIISGVGRAAEFGVLVRDADALQQASNLD 645
Query: 391 TVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAGALVDYARLHSIK 450
T+ FDKTGT+T G V + ++ + L W +++E+ S+HP+A A++ A ++
Sbjct: 646 TLVFDKTGTLTEGHPQVVAIHTFNGVSEQQALGWAAALETGSNHPLARAILQRAEGLTL- 704
Query: 451 PVPENVENFQNFPGEGIFGTIDGRDIYIGNKR------VGVRAICKRDNCEVQFQRPEIS 504
F+ G G+ G +DG + +GN R + R + + + +
Sbjct: 705 ---ATASQFRTLRGLGVSGEVDGIPLLLGNNRLLEEQQIDTRELQSLIQQQAESGATPVI 761
Query: 505 TKKNNCEETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAI 564
N L+ S+ D R ++ A++ L LG VMLTGD+ A + + I
Sbjct: 762 LTANGKPAALL---SIRDPLREDSIGALQRLHQLGYSLVMLTGDNPITANAIAKEA--GI 816
Query: 565 DIVHAELLPHEKAKLIENFKKEG-PIAMIGDGINDAPALATADIGISMGISGSALANETS 623
D V A +LP KA I+ + G +AMIGDGINDAPALA AD+GI+MG GS +A ET+
Sbjct: 817 DRVIAGVLPDGKADAIKQLQAAGHKVAMIGDGINDAPALAQADVGIAMG-GGSDIAIETA 875
Query: 624 NAILMSNDIRKVPEAIRLARKTTRKLVENVI 654
LM + + V +A+ L++ T R + +N++
Sbjct: 876 AITLMRHSLYGVVDAVELSKATLRNMKQNLL 906
>CTPA_MYCLE (P46839) Cation-transporting P-type ATPase A (EC
3.6.3.-)
Length = 780
Score = 201 bits (512), Expect = 5e-51
Identities = 156/545 (28%), Positives = 262/545 (47%), Gaps = 40/545 (7%)
Query: 146 AVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETGER-- 203
A+ G+ A+ G+ +F+ + + +H ++ ++ A A++ G
Sbjct: 185 ALLGSDAIYFEVAAGITVFVLAGKYYTARAKSHASIALLALAALSAKDAAVLQPDGSEMV 244
Query: 204 VDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINM 263
+ N++ V+ G I DG+V++G V +TGE+ PV + V GT+ +
Sbjct: 245 IPANELNEQQRFVVRPGQTIAADGLVIDGSATVSMSPITGEAKPVRVNPGAQVIGGTVVL 304
Query: 264 NAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYSC 323
N ++I + +VG V+ ++ Q + ++ V C
Sbjct: 305 NG--RLIVEAA----------AVGDETQLAGMVRLVEQAQQQNANAQRLADRIASVFVPC 352
Query: 324 IAVVPAALSV--------PDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGL 375
+ V A +V PD F A+ VL+ CPCAL L+TP A+ A + A G+
Sbjct: 353 VFAVAALTAVGWLIAGSGPDRV--FSAAIAVLVIACPCALGLATPTAMMVASGRGAQLGI 410
Query: 376 LLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHP 435
LLKG + E + TV FDKTGT+T G+ V+ +A +L +++ES S H
Sbjct: 411 LLKGHESFEATRAVDTVVFDKTGTLTTGQLKVSAVTAAPGWQANEVLQMAATVESASEHA 470
Query: 436 VAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRVGVRAICKRDNCE 495
VA A+ + H E V NF+ PG G+ GT+ R + +G K + + C
Sbjct: 471 VALAIA-ASTTHR-----EPVANFRAVPGHGVSGTVAERAVRVG-KPSWIASRCNSTTLV 523
Query: 496 VQFQRPEISTKKN---NCEETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQV 552
+ E+ + + GV ++ DA ++ A +A+ L G R+ +LTGD+
Sbjct: 524 TARRNAELRGETAVFVEIDGEQCGVIAVADAVKASAADAVAALHDRGFRTALLTGDNPAS 583
Query: 553 AMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEG-PIAMIGDGINDAPALATADIGISM 611
A V S++ ID V A++LP +K +IE + G +AM+GDGIND PALA AD+G+++
Sbjct: 584 AAAVASRI--GIDEVIADILPEDKVDVIEQLRDRGHVVAMVGDGINDGPALARADLGMAI 641
Query: 612 GISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIAGY 671
G G+ +A ++ IL+ +++ VP + LA T R + N++ + G+ A + +A AG
Sbjct: 642 G-RGTDVAIGAADIILVRDNLDVVPITLDLAAATMRTIKFNMVWAFGYNIAAIPIAAAGL 700
Query: 672 --PLV 674
PLV
Sbjct: 701 LNPLV 705
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.137 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 90,026,408
Number of Sequences: 164201
Number of extensions: 3662894
Number of successful extensions: 11088
Number of sequences better than 10.0: 319
Number of HSP's better than 10.0 without gapping: 277
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 9616
Number of HSP's gapped (non-prelim): 1044
length of query: 833
length of database: 59,974,054
effective HSP length: 119
effective length of query: 714
effective length of database: 40,434,135
effective search space: 28869972390
effective search space used: 28869972390
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC130275.7


