

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.4 - phase: 0
(362 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
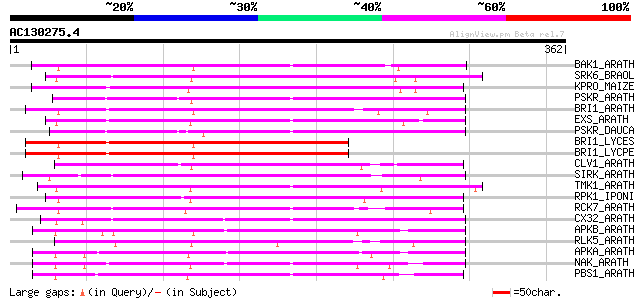
Score E
Sequences producing significant alignments: (bits) Value
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 197 3e-50
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 186 1e-46
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 180 6e-45
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 176 6e-44
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 175 2e-43
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 174 2e-43
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 172 1e-42
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 167 4e-41
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 166 9e-41
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 161 2e-39
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 160 4e-39
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 160 6e-39
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 157 4e-38
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 157 5e-38
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 154 4e-37
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 149 8e-36
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 149 1e-35
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 146 7e-35
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 145 2e-34
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 144 3e-34
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 197 bits (502), Expect = 3e-50
Identities = 124/294 (42%), Positives = 173/294 (58%), Gaps = 14/294 (4%)
Query: 15 RFTGQQLRIATDNYSN--LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
RF+ ++L++A+DN+SN +LG GGFG VYKG ++GT+VAVK L+ + + QF EV
Sbjct: 276 RFSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGELQFQTEV 335
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV---LGYEKLHEIAI 129
I H NL+RL GFC LVY YM NGS+ L + L + K IA+
Sbjct: 336 EMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWPKRQRIAL 395
Query: 130 GTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRG 189
G+ARG+AYLH+ C +IIH D+K NILLD+ F V DFGLAK + ++TH+T T RG
Sbjct: 396 GSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVT-TAVRG 454
Query: 190 TPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAG 249
T G+ APE + K DV+ +G++L E+I +R + + + + W K G
Sbjct: 455 TIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVK---G 511
Query: 250 LLG----EAMIVCGIEEKNK-EIAERMIKVALWCVQYRPELRPIMSVVVKMLEG 298
LL EA++ ++ K E E++I+VAL C Q P RP MS VV+MLEG
Sbjct: 512 LLKEKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRMLEG 565
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 186 bits (471), Expect = 1e-46
Identities = 117/297 (39%), Positives = 168/297 (56%), Gaps = 13/297 (4%)
Query: 24 ATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHF 81
AT+N+S N LG GGFG VYKG +G +AVK L +S + DE FM EV I R+ H
Sbjct: 524 ATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKRLSKTSVQGTDE-FMNEVTLIARLQHI 582
Query: 82 NLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETK--VLGYEKLHEIAIGTARGIAYLH 139
NLV++ G C E + L+YEY+ N SLD YLF +T+ L + + +I G ARG+ YLH
Sbjct: 583 NLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKTRRSKLNWNERFDITNGVARGLLYLH 642
Query: 140 EECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELW 199
++ + RIIH D+K NILLDKN PK++DFG+A+ R+ T GT GY +PE
Sbjct: 643 QDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFERDETEANTMKVVGTYGYMSPEYA 702
Query: 200 MPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGL---LGEAMI 256
M + K DV+SFG+++ EI+ ++N N + + +VW + G + + +I
Sbjct: 703 MYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYENDLLSYVWSRWKEGRALEIVDPVI 762
Query: 257 VCGIEEK----NKEIAERMIKVALWCVQYRPELRPIMSVVVKML-EGSLEIPKTFNP 308
V + + + + I++ L CVQ E RP MS VV M + EIP+ P
Sbjct: 763 VDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRPAMSSVVWMFGSEATEIPQPKPP 819
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 180 bits (456), Expect = 6e-45
Identities = 119/292 (40%), Positives = 163/292 (55%), Gaps = 12/292 (4%)
Query: 15 RFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGT 74
R++ ++L AT + LG G GTVYKG+ + VAVK L K E F AE+
Sbjct: 523 RYSYRELVKATRKFKVELGRGESGTVYKGVLEDDRHVAVKKLENVRQGK--EVFQAELSV 580
Query: 75 IGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE--TKVLGYEKLHEIAIGTA 132
IGRI+H NLVR++GFC E + LV EY+ NGSL LF E +L +E IA+G A
Sbjct: 581 IGRINHMNLVRIWGFCSEGSHRLLVSEYVENGSLANILFSEGGNILLDWEGRFNIALGVA 640
Query: 133 RGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPG 192
+G+AYLH EC +IH D+KP NILLD+ F PK+ DFGL K NR + ++ RGT G
Sbjct: 641 KGLAYLHHECLEWVIHCDVKPENILLDQAFEPKITDFGLVKLLNRGGSTQNVSHVRGTLG 700
Query: 193 YAAPELWMPFPITHKCDVYSFGMLLFEII-GRRRNLAIKNTESQEWFPIWVWKKKDAGLL 251
Y APE PIT K DVYS+G++L E++ G R + + T+ + + A L
Sbjct: 701 YIAPEWVSSLPITAKVDVYSYGVVLLELLTGTRVSELVGGTDEVHSMLRKLVRMLSAKLE 760
Query: 252 GE--AMIVCGIEEK-----NKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
GE + I ++ K N A +IK+A+ C++ RP M V+ L
Sbjct: 761 GEEQSWIDGYLDSKLNRPVNYVQARTLIKLAVSCLEEDRSKRPTMEHAVQTL 812
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 176 bits (447), Expect = 6e-44
Identities = 107/273 (39%), Positives = 150/273 (54%), Gaps = 7/273 (2%)
Query: 29 SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYG 88
+N++G GGFG VYK +G VA+K L G + I+ +F AEV T+ R H NLV L G
Sbjct: 737 ANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQ-IEREFEAEVETLSRAQHPNLVLLRG 795
Query: 89 FCFERNLIALVYEYMGNGSLDRYLFHETK----VLGYEKLHEIAIGTARGIAYLHEECQH 144
FCF +N L+Y YM NGSLD Y HE +L ++ IA G A+G+ YLHE C
Sbjct: 796 FCFYKNDRLLIYSYMENGSLD-YWLHERNDGPALLKWKTRLRIAQGAAKGLLYLHEGCDP 854
Query: 145 RIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPFPI 204
I+H DIK NILLD+NF +ADFGLA+ + TH++ T GT GY PE
Sbjct: 855 HILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVS-TDLVGTLGYIPPEYGQASVA 913
Query: 205 THKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLGEAMIVCGIEEKN 264
T+K DVYSFG++L E++ +R + + + WV K K E ++N
Sbjct: 914 TYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMKHESRASEVFDPLIYSKEN 973
Query: 265 KEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+ R++++A C+ P+ RP +V L+
Sbjct: 974 DKEMFRVLEIACLCLSENPKQRPTTQQLVSWLD 1006
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 175 bits (443), Expect = 2e-43
Identities = 114/298 (38%), Positives = 172/298 (57%), Gaps = 17/298 (5%)
Query: 11 EKPIR-FTGQQLRIATDNYSN--LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQ 67
EKP+R T L AT+ + N L+GSGGFG VYK I +G+ VA+K L S + D +
Sbjct: 865 EKPLRKLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQG-DRE 923
Query: 68 FMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV---LGYEKL 124
FMAE+ TIG+I H NLV L G+C + LVYE+M GSL+ L K L +
Sbjct: 924 FMAEMETIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTR 983
Query: 125 HEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITM 184
+IAIG+ARG+A+LH C IIH D+K N+LLD+N +V+DFG+A+ + +TH+++
Sbjct: 984 RKIAIGSARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSV 1043
Query: 185 TGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFP--IWV 242
+ GTPGY PE + F + K DVYS+G++L E++ +R T+S ++ +
Sbjct: 1044 STLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKR-----PTDSPDFGDNNLVG 1098
Query: 243 WKKKDAGLLGEAMIVCGIEEKNKEIAERM---IKVALWCVQYRPELRPIMSVVVKMLE 297
W K+ A L + + +++ + + +KVA+ C+ R RP M V+ M +
Sbjct: 1099 WVKQHAKLRISDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAMFK 1156
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 174 bits (442), Expect = 2e-43
Identities = 113/282 (40%), Positives = 165/282 (58%), Gaps = 12/282 (4%)
Query: 24 ATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHF 81
ATD++S N++G GGFGTVYK VAVK L + + + +FMAE+ T+G++ H
Sbjct: 913 ATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQG-NREFMAEMETLGKVKHP 971
Query: 82 NLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET---KVLGYEKLHEIAIGTARGIAYL 138
NLV L G+C LVYEYM NGSLD +L ++T +VL + K +IA+G ARG+A+L
Sbjct: 972 NLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGAARGLAFL 1031
Query: 139 HEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPEL 198
H IIH DIK NILLD +F PKVADFGLA+ + +H++ T GT GY PE
Sbjct: 1032 HHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVS-TVIAGTFGYIPPEY 1090
Query: 199 WMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPI-WVWKKKDAGLLGEAM-- 255
T K DVYSFG++L E++ + ES+ + W +K + G + +
Sbjct: 1091 GQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGNLVGWAIQKINQGKAVDVIDP 1150
Query: 256 IVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
++ + KN ++ R++++A+ C+ P RP M V+K L+
Sbjct: 1151 LLVSVALKNSQL--RLLQIAMLCLAETPAKRPNMLDVLKALK 1190
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 172 bits (436), Expect = 1e-42
Identities = 109/276 (39%), Positives = 148/276 (53%), Gaps = 9/276 (3%)
Query: 27 NYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRL 86
N +N++G GGFG VYK +GT VA+K L G + + +D +F AEV T+ R H NLV L
Sbjct: 744 NQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQ-MDREFQAEVETLSRAQHPNLVHL 802
Query: 87 YGFCFERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLH-----EIAIGTARGIAYLHEE 141
G+C +N L+Y YM NGSLD Y HE KV G L IA G A G+AYLH+
Sbjct: 803 LGYCNYKNDKLLIYSYMDNGSLD-YWLHE-KVDGPPSLDWKTRLRIARGAAEGLAYLHQS 860
Query: 142 CQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMP 201
C+ I+H DIK NILL F +ADFGLA+ +TH+T T GT GY PE
Sbjct: 861 CEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTHVT-TDLVGTLGYIPPEYGQA 919
Query: 202 FPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLGEAMIVCGIE 261
T+K DVYSFG++L E++ RR + + WV + K E +
Sbjct: 920 SVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWVLQMKTEKRESEIFDPFIYD 979
Query: 262 EKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+ + E ++++A C+ P+ RP +V LE
Sbjct: 980 KDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWLE 1015
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 167 bits (423), Expect = 4e-41
Identities = 96/217 (44%), Positives = 135/217 (61%), Gaps = 7/217 (3%)
Query: 11 EKPIR-FTGQQLRIATDNYSN--LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQ 67
EKP+R T L AT+ + N L+GSGGFG VYK +G++VA+K L S + D +
Sbjct: 870 EKPLRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQG-DRE 928
Query: 68 FMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV---LGYEKL 124
F AE+ TIG+I H NLV L G+C LVYEYM GSL+ L K+ L +
Sbjct: 929 FTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPAR 988
Query: 125 HEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITM 184
+IAIG ARG+A+LH C IIH D+K N+LLD+N +V+DFG+A+ + +TH+++
Sbjct: 989 RKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSV 1048
Query: 185 TGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEII 221
+ GTPGY PE + F + K DVYS+G++L E++
Sbjct: 1049 STLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELL 1085
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 166 bits (420), Expect = 9e-41
Identities = 96/217 (44%), Positives = 134/217 (61%), Gaps = 7/217 (3%)
Query: 11 EKPIR-FTGQQLRIATDNYSN--LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQ 67
EKP+R T L AT+ + N L+GSGGFG VYK +G++VA+K L S + D +
Sbjct: 870 EKPLRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQG-DRE 928
Query: 68 FMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV---LGYEKL 124
F AE+ TIG+I H NLV L G+C LVYEYM GSL+ L K L +
Sbjct: 929 FTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPAR 988
Query: 125 HEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITM 184
+IAIG ARG+A+LH C IIH D+K N+LLD+N +V+DFG+A+ + +TH+++
Sbjct: 989 RKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSV 1048
Query: 185 TGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEII 221
+ GTPGY PE + F + K DVYS+G++L E++
Sbjct: 1049 STLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELL 1085
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 161 bits (408), Expect = 2e-39
Identities = 101/280 (36%), Positives = 147/280 (52%), Gaps = 21/280 (7%)
Query: 30 NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGF 89
N++G GG G VY+G N VA+K L G + D F AE+ T+GRI H ++VRL G+
Sbjct: 696 NIIGKGGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRLLGY 755
Query: 90 CFERNLIALVYEYMGNGSLDRYLFHETK--VLGYEKLHEIAIGTARGIAYLHEECQHRII 147
++ L+YEYM NGSL L H +K L +E H +A+ A+G+ YLH +C I+
Sbjct: 756 VANKDTNLLLYEYMPNGSLGE-LLHGSKGGHLQWETRHRVAVEAAKGLCYLHHDCSPLIL 814
Query: 148 HYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPFPITHK 207
H D+K NILLD +F VADFGLAK M+ G+ GY APE + K
Sbjct: 815 HRDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAYTLKVDEK 874
Query: 208 CDVYSFGMLLFEIIGRRRNLA-----------IKNTESQEWFPIWVWKKKDAGLLGEAMI 256
DVYSFG++L E+I ++ + ++NTE + + + DA ++ A++
Sbjct: 875 SDVYSFGVVLLELIAGKKPVGEFGEGVDIVRWVRNTEEE------ITQPSDAAIV-VAIV 927
Query: 257 VCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
+ + K+A+ CV+ RP M VV ML
Sbjct: 928 DPRLTGYPLTSVIHVFKIAMMCVEEEAAARPTMREVVHML 967
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 160 bits (406), Expect = 4e-39
Identities = 104/300 (34%), Positives = 161/300 (53%), Gaps = 19/300 (6%)
Query: 9 EREKPIRFTGQQLRIA-----TDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKK 63
ER P++ + + + T+N+ ++G GGFG VY G+ NG VAVKVL S +
Sbjct: 552 ERNGPLKTAKRYFKYSEVVNITNNFERVIGKGGFGKVYHGVI-NGEQVAVKVLSEESAQG 610
Query: 64 IDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETK-VLGYE 122
E F AEV + R+HH NL L G+C E N + L+YEYM N +L YL + +L +E
Sbjct: 611 YKE-FRAEVDLLMRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSFILSWE 669
Query: 123 KLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHI 182
+ +I++ A+G+ YLH C+ I+H D+KP NILL++ K+ADFGL+++ + E +
Sbjct: 670 ERLKISLDAAQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQ 729
Query: 183 TMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWV 242
T G+ GY PE + + K DVYS G++L E+I + +A TE
Sbjct: 730 ISTVVAGSIGYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKTEKVH------ 783
Query: 243 WKKKDAGLLGEAMIVCGIEEKNKE-----IAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+L I ++++ +E A +M ++AL C ++ RP MS VV L+
Sbjct: 784 ISDHVRSILANGDIRGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVVMELK 843
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 160 bits (404), Expect = 6e-39
Identities = 109/302 (36%), Positives = 155/302 (51%), Gaps = 13/302 (4%)
Query: 19 QQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLR-GSSNKKIDEQFMAEVGTI 75
Q LR T+N+S N+LGSGGFG VYKG +GT +AVK + G K +F +E+ +
Sbjct: 579 QVLRSVTNNFSSDNILGSGGFGVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIAVL 638
Query: 76 GRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET----KVLGYEKLHEIAIGT 131
++ H +LV L G+C + N LVYEYM G+L R+LF + K L +++ +A+
Sbjct: 639 TKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRLTLALDV 698
Query: 132 ARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTP 191
ARG+ YLH IH D+KP NILL + KVADFGL + I T GT
Sbjct: 699 ARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIE-TRIAGTF 757
Query: 192 GYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIW---VWKKKDA 248
GY APE + +T K DVYSFG++L E+I R++L E W ++ K+A
Sbjct: 758 GYLAPEYAVTGRVTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSWFKRMYINKEA 817
Query: 249 GLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEI--PKTF 306
++E+ + ++A C P RP M V +L +E+ P
Sbjct: 818 SFKKAIDTTIDLDEETLASVHTVAELAGHCCAREPYQRPDMGHAVNILSSLVELWKPSDQ 877
Query: 307 NP 308
NP
Sbjct: 878 NP 879
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 157 bits (397), Expect = 4e-38
Identities = 100/282 (35%), Positives = 150/282 (52%), Gaps = 10/282 (3%)
Query: 24 ATDNYSN--LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHF 81
AT+N ++ ++G G GT+YK S + AVK L + K + E+ TIG++ H
Sbjct: 812 ATENLNDKYVIGKGAHGTIYKATLSPDKVYAVKKLVFTGIKNGSVSMVREIETIGKVRHR 871
Query: 82 NLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET---KVLGYEKLHEIAIGTARGIAYL 138
NL++L F + ++Y YM NGSL L HET K L + H IA+GTA G+AYL
Sbjct: 872 NLIKLEEFWLRKEYGLILYTYMENGSLHDIL-HETNPPKPLDWSTRHNIAVGTAHGLAYL 930
Query: 139 HEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPEL 198
H +C I+H DIKP NILLD + P ++DFG+AK ++ T I +GT GY APE
Sbjct: 931 HFDCDPAIVHRDIKPMNILLDSDLEPHISDFGIAKLLDQSATSIPSNTVQGTIGYMAPEN 990
Query: 199 WMPFPITHKCDVYSFGMLLFEIIGRRRNLAIK---NTESQEWF-PIWVWKKKDAGLLGEA 254
+ + DVYS+G++L E+I R++ L T+ W +W + ++ +
Sbjct: 991 AFTTVKSRESDVYSYGVVLLELITRKKALDPSFNGETDIVGWVRSVWTQTGEIQKIVDPS 1050
Query: 255 MIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
++ I+ E + +AL C + + RP M VVK L
Sbjct: 1051 LLDELIDSSVMEQVTEALSLALRCAEKEVDKRPTMRDVVKQL 1092
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 157 bits (396), Expect = 5e-38
Identities = 115/313 (36%), Positives = 170/313 (53%), Gaps = 35/313 (11%)
Query: 5 LNDMEREKPIR-FTGQQLRIATDNYSN--LLGSGGFGTVYKGIFSN-GTMVAVKVLRGSS 60
LND K + FT Q+L AT N+ + LG GGFG V+KG +VA+K L +
Sbjct: 79 LNDQVTGKKAQTFTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNG 138
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF---HETK 117
+ I E F+ EV T+ H NLV+L GFC E + LVYEYM GSL+ +L K
Sbjct: 139 VQGIRE-FVVEVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKK 197
Query: 118 VLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAK-NCN 176
L + +IA G ARG+ YLH+ +I+ D+K NILL +++ PK++DFGLAK +
Sbjct: 198 PLDWNTRMKIAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPS 257
Query: 177 RENTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQE 236
+ TH++ T GT GY AP+ M +T K D+YSFG++L E+I R+ AI NT++
Sbjct: 258 GDKTHVS-TRVMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRK--AIDNTKT-- 312
Query: 237 WFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIK-------------VALWCVQYRP 283
+KD L+G A + ++ + +++ ++ CVQ +P
Sbjct: 313 --------RKDQNLVGWARPLFKDRRNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQP 364
Query: 284 ELRPIMSVVVKML 296
+RP++S VV L
Sbjct: 365 TMRPVVSDVVLAL 377
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 154 bits (388), Expect = 4e-37
Identities = 105/291 (36%), Positives = 158/291 (54%), Gaps = 17/291 (5%)
Query: 21 LRIATDNYS--NLLGSGGFGTVYKGIFS----------NGTMVAVKVLRGSSNKKIDEQF 68
L+ AT N+ ++LG GGFG VY+G +G +VA+K L S + E +
Sbjct: 79 LKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRLNSESVQGFAE-W 137
Query: 69 MAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLHEIA 128
+EV +G + H NLV+L G+C E + LVYE+M GSL+ +LF ++ +I
Sbjct: 138 RSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLESHLFRRNDPFPWDLRIKIV 197
Query: 129 IGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAK-NCNRENTHITMTGG 187
IG ARG+A+LH Q +I+ D K NILLD N+ K++DFGLAK E +H+T T
Sbjct: 198 IGAARGLAFLH-SLQREVIYRDFKASNILLDSNYDAKLSDFGLAKLGPADEKSHVT-TRI 255
Query: 188 RGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKD 247
GT GYAAPE + K DV++FG++L EI+ K QE W+ +
Sbjct: 256 MGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTAHNTKRPRGQESLVDWLRPELS 315
Query: 248 AGLLGEAMIVCGIE-EKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+ ++ GI+ + ++A M ++ L C++ P+ RP M VV++LE
Sbjct: 316 NKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDPKNRPHMKEVVEVLE 366
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 149 bits (377), Expect = 8e-36
Identities = 105/301 (34%), Positives = 164/301 (53%), Gaps = 24/301 (7%)
Query: 16 FTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGS--SNKKIDE----- 66
FT +L+ AT N+ ++LG GGFG+V+KG T+ A K G + KK+++
Sbjct: 57 FTFAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTASKPGTGVVIAVKKLNQDGWQG 116
Query: 67 --QFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV---LGY 121
+++AEV +G+ H NLV+L G+C E LVYE+M GSL+ +LF L +
Sbjct: 117 HQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGSYFQPLSW 176
Query: 122 EKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTH 181
++A+G A+G+A+LH + +I+ D K NILLD + K++DFGLAK+ +
Sbjct: 177 TLRLKVALGAAKGLAFLHN-AETSVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPTGDKS 235
Query: 182 ITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRR----NLAIKNTESQEW 237
T GT GYAAPE +T K DVYS+G++L E++ RR N + EW
Sbjct: 236 HVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSGRRAVDKNRPPGEQKLVEW 295
Query: 238 F-PIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
P+ K+K ++ + ++ + E A ++ +AL C+ + +LRP M+ VV L
Sbjct: 296 ARPLLANKRKLFRVIDNRL----QDQYSMEEACKVATLALRCLTFEIKLRPNMNEVVSHL 351
Query: 297 E 297
E
Sbjct: 352 E 352
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 149 bits (376), Expect = 1e-35
Identities = 107/290 (36%), Positives = 148/290 (50%), Gaps = 33/290 (11%)
Query: 30 NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQ---------FMAEVGTIGRIHH 80
N++G G G VYK G +VAVK L S DE F AEV T+G I H
Sbjct: 687 NVIGFGSSGKVYKVELRGGEVVAVKKLNKSVKGGDDEYSSDSLNRDVFAAEVETLGTIRH 746
Query: 81 FNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETK---VLGYEKLHEIAIGTARGIAY 137
++VRL+ C + LVYEYM NGSL L + K VLG+ + IA+ A G++Y
Sbjct: 747 KSIVRLWCCCSSGDCKLLVYEYMPNGSLADVLHGDRKGGVVLGWPERLRIALDAAEGLSY 806
Query: 138 LHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAK--NCNRENTHITMTGGRGTPGYAA 195
LH +C I+H D+K NILLD ++ KVADFG+AK + T M+G G+ GY A
Sbjct: 807 LHHDCVPPIVHRDVKSSNILLDSDYGAKVADFGIAKVGQMSGSKTPEAMSGIAGSCGYIA 866
Query: 196 PELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLGEAM 255
PE + K D+YSFG++L E++ + + T+S+ + K A + A+
Sbjct: 867 PEYVYTLRVNEKSDIYSFGVVLLELVTGK-----QPTDSE------LGDKDMAKWVCTAL 915
Query: 256 IVCGIEE--------KNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
CG+E K KE ++I + L C P RP M VV ML+
Sbjct: 916 DKCGLEPVIDPKLDLKFKEEISKVIHIGLLCTSPLPLNRPSMRKVVIMLQ 965
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 146 bits (369), Expect = 7e-35
Identities = 107/303 (35%), Positives = 165/303 (54%), Gaps = 28/303 (9%)
Query: 16 FTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSN----------GTMVAVKVLRGSSNKK 63
F+ +L+ AT N+ ++LG GGFG V+KG G ++AVK L +
Sbjct: 56 FSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGLVIAVKKLN-QDGWQ 114
Query: 64 IDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE---TKVLG 120
++++AEV +G+ H +LV+L G+C E LVYE+M GSL+ +LF + L
Sbjct: 115 GHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGLYFQPLS 174
Query: 121 YEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENT 180
++ ++A+G A+G+A+LH + R+I+ D K NILLD + K++DFGLAK+ +
Sbjct: 175 WKLRLKVALGAAKGLAFLHSS-ETRVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPIGDK 233
Query: 181 HITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTES-----Q 235
T GT GYAAPE +T K DVYSFG++L E++ RR + KN S
Sbjct: 234 SHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVD-KNRPSGERNLV 292
Query: 236 EWF-PIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVK 294
EW P V K+K ++ + ++ + E A ++ ++L C+ +LRP MS VV
Sbjct: 293 EWAKPYLVNKRKIFRVIDNRL----QDQYSMEEACKVATLSLRCLTTEIKLRPNMSEVVS 348
Query: 295 MLE 297
LE
Sbjct: 349 HLE 351
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 145 bits (366), Expect = 2e-34
Identities = 101/308 (32%), Positives = 164/308 (52%), Gaps = 38/308 (12%)
Query: 16 FTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSN----------GTMVAVKVLRGSSNKK 63
F+ +L+ AT N+ +++G GGFG V+KG G ++AVK L +
Sbjct: 56 FSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLNQEGFQG 115
Query: 64 IDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET---KVLG 120
+++AE+ +G++ H NLV+L G+C E LVYE+M GSL+ +LF + L
Sbjct: 116 -HREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFYQPLS 174
Query: 121 YEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNR-EN 179
+ +A+G ARG+A+LH Q ++I+ D K NILLD N+ K++DFGLA++ +N
Sbjct: 175 WNTRVRMALGAARGLAFLHN-AQPQVIYRDFKASNILLDSNYNAKLSDFGLARDGPMGDN 233
Query: 180 THITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRR----NLAIKNTESQ 235
+H++ T GT GYAAPE ++ K DVYSFG++L E++ RR N +
Sbjct: 234 SHVS-TRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVGEHNLV 292
Query: 236 EWFPIWVWKKK------DAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIM 289
+W ++ K+ D L G+ + + ++ +AL C+ + RP M
Sbjct: 293 DWARPYLTNKRRLLRVMDPRLQGQYSLTRAL---------KIAVLALDCISIDAKSRPTM 343
Query: 290 SVVVKMLE 297
+ +VK +E
Sbjct: 344 NEIVKTME 351
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 144 bits (364), Expect = 3e-34
Identities = 102/298 (34%), Positives = 155/298 (51%), Gaps = 28/298 (9%)
Query: 16 FTGQQLRIATDNY--SNLLGSGGFGTVYKG-IFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
F ++L AT N+ LG GGFG VYKG + S G +VAVK L + + + +F+ EV
Sbjct: 74 FAFRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQL-DRNGLQGNREFLVEV 132
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFH---ETKVLGYEKLHEIAI 129
+ +HH NLV L G+C + + LVYE+M GSL+ +L + + L + +IA
Sbjct: 133 LMLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAA 192
Query: 130 GTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAK-NCNRENTHITMTGGR 188
G A+G+ +LH++ +I+ D K NILLD+ F+PK++DFGLAK + +H++ T
Sbjct: 193 GAAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVS-TRVM 251
Query: 189 GTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWV------ 242
GT GY APE M +T K DVYSFG++ E+I R+ + + ++ W
Sbjct: 252 GTYGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPLFND 311
Query: 243 ----WKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKML 296
K D L G + + VA C+Q + RP+++ VV L
Sbjct: 312 RRKFIKLADPRLKGRF---------PTRALYQALAVASMCIQEQAATRPLIADVVTAL 360
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,320,030
Number of Sequences: 164201
Number of extensions: 1959034
Number of successful extensions: 7690
Number of sequences better than 10.0: 1648
Number of HSP's better than 10.0 without gapping: 1103
Number of HSP's successfully gapped in prelim test: 545
Number of HSP's that attempted gapping in prelim test: 4287
Number of HSP's gapped (non-prelim): 1864
length of query: 362
length of database: 59,974,054
effective HSP length: 111
effective length of query: 251
effective length of database: 41,747,743
effective search space: 10478683493
effective search space used: 10478683493
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC130275.4


