

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.11 - phase: 0
(309 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
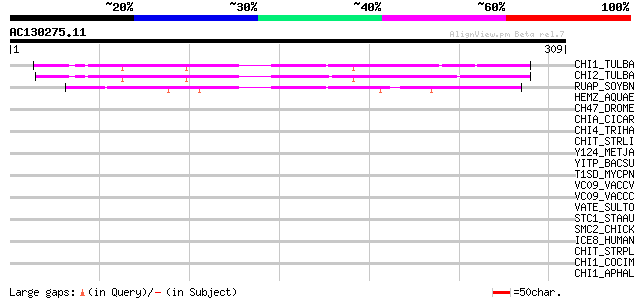
Sequences producing significant alignments: (bits) Value
CHI1_TULBA (Q9SLP4) Chitinase 1 precursor (EC 3.2.1.14) (Tulip b... 142 1e-33
CHI2_TULBA (Q7M443) Chitinase 2 (EC 3.2.1.14) (Tulip bulb chitin... 138 2e-32
RUAP_SOYBN (P39657) RuBisCO-associated protein 80 6e-15
HEMZ_AQUAE (O67083) Ferrochelatase (EC 4.99.1.1) (Protoheme ferr... 38 0.028
CH47_DROME (Q23997) Chitinase-like protein DS47 precursor 36 0.14
CHIA_CICAR (P36908) Acidic endochitinase precursor (EC 3.2.1.14) 35 0.24
CHI4_TRIHA (P48827) 42 kDa endochitinase precursor (EC 3.2.1.14) 34 0.53
CHIT_STRLI (P36909) Chitinase C precursor (EC 3.2.1.14) 33 0.69
Y124_METJA (Q57588) Hypothetical protein MJ0124 33 0.90
YITP_BACSU (O06751) Hypothetical protein yitP 32 1.5
T1SD_MYCPN (P75435) Putative type I restriction enzyme specifici... 32 1.5
VC09_VACCV (P17372) Protein C9 31 3.4
VC09_VACCC (P21042) Protein C9 31 3.4
VATE_SULTO (Q971B8) V-type ATP synthase subunit E (EC 3.6.3.14) ... 31 3.4
STC1_STAAU (P07767) Staphylocoagulase precursor 31 3.4
SMC2_CHICK (Q90988) Structural maintenance of chromosome 2 (Chro... 31 4.5
ICE8_HUMAN (Q14790) Caspase-8 precursor (EC 3.4.22.-) (CASP-8) (... 31 4.5
CHIT_STRPL (P11220) Chitinase 63 precursor (EC 3.2.1.14) 31 4.5
CHI1_COCIM (P54196) Endochitinase 1 precursor (EC 3.2.1.14) (Com... 31 4.5
CHI1_APHAL (P32470) Chitinase 1 precursor (EC 3.2.1.14) 31 4.5
>CHI1_TULBA (Q9SLP4) Chitinase 1 precursor (EC 3.2.1.14) (Tulip bulb
chitinase-1) (TBC-1)
Length = 314
Score = 142 bits (357), Expect = 1e-33
Identities = 95/285 (33%), Positives = 150/285 (52%), Gaps = 32/285 (11%)
Query: 14 IIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKG--KGEFYRTWNF 71
++FREYIG + D P IN D++ FHFIL + SG G F W+
Sbjct: 27 LVFREYIGSQFNDVKFSDVP---INPDVD-FHFILAFAIDYTSGSSPTPTNGNFKPFWDT 82
Query: 72 NNFSPAKVAKLKKDHKNVKVIISIGG--FGAENP-FNPKEIESWSTKAKQSIKKLINEYQ 128
NN SP++VA +K+ H NVKV +S+GG G +N F+P + SW A S+ ++I +Y
Sbjct: 83 NNLSPSQVAAVKRTHSNVKVSLSLGGDSVGGKNVFFSPSSVSSWVENAVSSLTRIIKQYH 142
Query: 129 EYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAP 188
+DGIDI+YE+ +P+ F+ CIG+L+ +LKK ++ + VSIAP
Sbjct: 143 -----------------LDGIDIDYEHFKGDPNTFAECIGQLVTRLKK-NEVVSFVSIAP 184
Query: 189 TE--LLKPHYHKLYWANKDIINWVDYKFYNQTV-SSADELVNLYNKLLNEYGTDVKLLPG 245
+ ++ HY L+ I++V+++FY + +S ++ + + + + Y K+L
Sbjct: 185 FDDAQVQSHYQALWEKYGHQIDYVNFQFYAYSARTSVEQFLKYFEEQSSNYHGG-KVLVS 243
Query: 246 VSTDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSN 290
STD F + C L + L GIFVW+A+DS M +N
Sbjct: 244 FSTDSSGGLKPDNG-FFRACSILKKQGKLHGIFVWSADDSLMSNN 287
>CHI2_TULBA (Q7M443) Chitinase 2 (EC 3.2.1.14) (Tulip bulb
chitinase-2) (TBC-2)
Length = 275
Score = 138 bits (348), Expect = 2e-32
Identities = 96/283 (33%), Positives = 146/283 (50%), Gaps = 30/283 (10%)
Query: 15 IFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKG--KGEFYRTWNFN 72
+FREYIG + D P IN +++ FHFIL + SG G F W+ N
Sbjct: 2 LFREYIGAQFNDVKFSDVP---INPNVD-FHFILAFAIDYTSGSSPTPTNGNFNPFWDTN 57
Query: 73 NFSPAKVAKLKKDHKNVKVIISIGG--FGAENPF-NPKEIESWSTKAKQSIKKLINEYQE 129
N SP++VA +K+ + NVKV +S+GG G E F NP + SW A S+ K+I +Y
Sbjct: 58 NLSPSQVAAIKRTYNNVKVSVSLGGNSVGGERVFFNPSSVSSWVDNAVSSLTKIIKQYH- 116
Query: 130 YSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAPT 189
+DGIDI+YE+ +P+ F+ CIG+L+ +LKK+ + VSIAP
Sbjct: 117 ----------------LDGIDIDYEHFKGDPNTFAECIGQLVTRLKKNG-VVSFVSIAPF 159
Query: 190 E--LLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTDVKLLPGVS 247
+ ++ HY L+ I++V+++FY + + E Y ++ K+L S
Sbjct: 160 DDAQVQSHYQALWEKYGHQIDYVNFQFYVYSSRMSVEQFLKYFEMQRSNYPGGKVLVSFS 219
Query: 248 TDPDSNTNMTRDVFIKGCKSLLESESLPGIFVWNANDSAMPSN 290
TD +S R+ F C L + L GIFVW+A+DS M ++
Sbjct: 220 TD-NSGGLKPRNGFFDACSILKKQGKLHGIFVWSADDSLMSND 261
>RUAP_SOYBN (P39657) RuBisCO-associated protein
Length = 283
Score = 80.1 bits (196), Expect = 6e-15
Identities = 70/272 (25%), Positives = 122/272 (44%), Gaps = 42/272 (15%)
Query: 32 FPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNFSPAKVAKLKKDHK---- 87
F ++I ++I EF L R+ Y G+ G+F W+ +P + K KK ++
Sbjct: 17 FLNQVIPENITEFQVTLSLARD-YDGNNSTNGKFIPYWDTEKVTPEVIKKFKKKYEPTAL 75
Query: 88 NVKVIISIGGFGAENPF---NPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDD 144
VKV++SIG + PF + E+W ++A S+K +I Y
Sbjct: 76 RVKVLVSIGNKNKQFPFTIGSDSNSEAWVSEATASLKSIIKTYN---------------- 119
Query: 145 IIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAPT-ELLKPHYHKLYWAN 203
+DGID++YE N +F + +G L+R LK+ +K I + S A + + ++ L +A
Sbjct: 120 -LDGIDVSYEDIAANEADFVNSVGGLVRNLKQ-NKLITVASFATSADAANNKFYNLLYAE 177
Query: 204 KD-------IINWVDYKFYNQTVSSADELVNLYNKLL---NEYGTDVKLLPGVSTDPDSN 253
++WV + T S A+ + +L K+L NEY L ST +
Sbjct: 178 YATFFDTVVFLSWVGF-----TPSRANPVASLEEKILAVANEYKAVKAFLVAYSTVAEDW 232
Query: 254 TNMTRDVFIKGCKSLLESESLPGIFVWNANDS 285
N + +F LL + ++ G + +D+
Sbjct: 233 ANFSPPIFFITLHGLLGNSAVKGASIKVISDA 264
>HEMZ_AQUAE (O67083) Ferrochelatase (EC 4.99.1.1) (Protoheme
ferro-lyase) (Heme synthetase)
Length = 309
Score = 38.1 bits (87), Expect = 0.028
Identities = 51/209 (24%), Positives = 84/209 (39%), Gaps = 42/209 (20%)
Query: 16 FREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNFS 75
++ +G++ +KD +E++ + I E IL L YS G FN F
Sbjct: 85 YKVVVGMRYWKPYIKDALSELLKEGINEV--ILLPLYPQYSKTTTGSA-------FNEFE 135
Query: 76 PAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSI--------------- 120
+K A LK DH VK I +P I++W+ + KQS+
Sbjct: 136 RSKKA-LKADHIKVKKIEHFYD-------HPLYIKAWAEQIKQSVEKPEEYHFLFSAHSL 187
Query: 121 -KKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCI----GELIRKLK 175
KKLI E Y + + T + ++ ++ Y + + F + E+IR L
Sbjct: 188 PKKLIEEGDPYQEQTEKTVKLIMENF---PEVEYTLAYQSKVGFGKWLEPSTDEVIRNLI 244
Query: 176 KSSKSIKLVSIAPTELLKPHYHKLYWANK 204
K K +K + + P + H LY +K
Sbjct: 245 K--KEVKKLLVIPISFVSEHSETLYELDK 271
>CH47_DROME (Q23997) Chitinase-like protein DS47 precursor
Length = 452
Score = 35.8 bits (81), Expect = 0.14
Identities = 35/125 (28%), Positives = 56/125 (44%), Gaps = 35/125 (28%)
Query: 61 GKGEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSI 120
GKG YRT V +LK+ + NVK+++S+GG K+IE +
Sbjct: 89 GKG-LYRT----------VTRLKRKYPNVKILLSVGG--------DKDIE-----LDKDA 124
Query: 121 KKLINEYQEYSKDSSSTDECHCDDII---------DGIDINYEYSNCNPDEFSSCIGELI 171
K+L N+Y E + S T + + DG+D+ +++ P + S IG L
Sbjct: 125 KELPNKYLELLE--SPTGRTRFVNTVYSLVKTYGFDGLDVAWQFPKNKPKKVHSGIGSLW 182
Query: 172 RKLKK 176
+ KK
Sbjct: 183 KGFKK 187
>CHIA_CICAR (P36908) Acidic endochitinase precursor (EC 3.2.1.14)
Length = 293
Score = 35.0 bits (79), Expect = 0.24
Identities = 46/207 (22%), Positives = 81/207 (38%), Gaps = 25/207 (12%)
Query: 87 KNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDDII 146
K +KV++S+GG S+S + + L N +ST D ++
Sbjct: 93 KGIKVLLSLGGGAG----------SYSLNSAEEATTLANYLWNNFLGGTSTSRPLGDAVL 142
Query: 147 DGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAPTELLKPHYHKLYWANKDI 206
DGID + E + D EL + L S+ +S AP + P H +
Sbjct: 143 DGIDFDIESGGQHYD-------ELAKALNGFSQQKVYLSAAP-QCPYPDAHLDSAIQTGL 194
Query: 207 INWVDYKFYNQ-----TVSSADELVNLYNKLLNEYGTDVKL-LPGVSTDPDSNTNMTRDV 260
++V +FYN + + + LVN +N+ + V L +P S + DV
Sbjct: 195 FDYVWVQFYNNPQCQYSNGNINNLVNAWNQWTSSQAKQVFLGVPASDAAAPSGGLIPADV 254
Query: 261 FI-KGCKSLLESESLPGIFVWNANDSA 286
+ ++ S G+ +W+ + A
Sbjct: 255 LTSQVLPAIKTSPKYGGVMIWDRFNDA 281
>CHI4_TRIHA (P48827) 42 kDa endochitinase precursor (EC 3.2.1.14)
Length = 423
Score = 33.9 bits (76), Expect = 0.53
Identities = 56/229 (24%), Positives = 94/229 (40%), Gaps = 42/229 (18%)
Query: 33 PAEMINDDIEEFHFILGTLRE---VYSGDGKGKGEFYR---TWN---FNNFSPAK-VAKL 82
PA+++ D+ + L+ V SGD E + +WN N + K + K+
Sbjct: 56 PADLVASDVTHVIYSFMNLQADGTVISGDTYADYEKHYADDSWNDVGTNAYGCVKQLFKV 115
Query: 83 KKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHC 142
KK ++ +KV++SIGG+ +WST + N + ++K + + +
Sbjct: 116 KKANRGLKVLLSIGGW------------TWSTNFPSAASTDANR-KNFAKTAITFMK--- 159
Query: 143 DDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAP---------TELLK 193
D DGIDI++EY + + S+ I L+ K +S + AP K
Sbjct: 160 DWGFDGIDIDWEYP-ADATQASNMI--LLLKEVRSQRDAYAAQYAPGYHFLLTIAAPAGK 216
Query: 194 PHYHKLYWAN----KDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGT 238
+Y KL A+ D IN + Y + NL+N N T
Sbjct: 217 DNYSKLRLADLGQVLDYINLMAYDYAGSFSPLTGHDANLFNNPSNPNAT 265
>CHIT_STRLI (P36909) Chitinase C precursor (EC 3.2.1.14)
Length = 619
Score = 33.5 bits (75), Expect = 0.69
Identities = 29/125 (23%), Positives = 56/125 (44%), Gaps = 25/125 (20%)
Query: 70 NFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQE 129
NFN ++ KLK + N+K++ S GG+ F P +++ + AK S L+
Sbjct: 318 NFN-----QLRKLKAKYPNIKILYSFGGWTWSGGF-PDAVKNPAAFAK-SCHDLV----- 365
Query: 130 YSKDSSSTDECHCDDIIDGIDINYEYSN-----CNPDEFSSCIGELIRKLKKSSKSIKLV 184
++ D+ DGID+++EY N C+ + +++ ++ L+
Sbjct: 366 --------EDPRWADVFDGIDLDWEYPNACGLSCDETSAPNAFSSMMKAMRAEFGQDYLI 417
Query: 185 SIAPT 189
+ A T
Sbjct: 418 TAAVT 422
>Y124_METJA (Q57588) Hypothetical protein MJ0124
Length = 1075
Score = 33.1 bits (74), Expect = 0.90
Identities = 58/257 (22%), Positives = 96/257 (36%), Gaps = 30/257 (11%)
Query: 22 VKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNF-----SP 76
VK+ KNLK IND E+ + TL+ + E + F
Sbjct: 751 VKESLKNLK------IND--EDLSIDVNTLKTLTKNKDFNNNELKEKLDLIAFYAEDGKN 802
Query: 77 AKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEYSKD-SS 135
A++ KL D K V + G + F ++IE S IKKL + + K
Sbjct: 803 ARILKLIDDLKAVIKLYKALGSYPQKIFYIEDIELLSFIYAYLIKKLKPKKKSNRKFWEE 862
Query: 136 STDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAPTELLKPH 195
H ++D + + E N NPD+ + E I K + I +L
Sbjct: 863 LISFIHNKMLVDDLTV-IEEINLNPDDLDKILKENIGKREIKRAVANYYFILKNSILDKQ 921
Query: 196 YHKLYWANKDII--------NWV----DYKFYNQTVSSADELVNLYNKLLNEYGTDVKLL 243
+ +Y K+I+ +W+ D K Y + + EL N Y+K + + ++
Sbjct: 922 HDPIY---KEILERLERLRRDWIMKRIDDKIYLNAIKNLMELKNNYDKKIKGKSSIERIK 978
Query: 244 PGVSTDPDSNTNMTRDV 260
+ST N +D+
Sbjct: 979 ESISTYIGENILKDQDI 995
>YITP_BACSU (O06751) Hypothetical protein yitP
Length = 178
Score = 32.3 bits (72), Expect = 1.5
Identities = 17/67 (25%), Positives = 35/67 (51%), Gaps = 3/67 (4%)
Query: 148 GIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAPTELLKPHYHKLYWANKDII 207
G+++ YS +++++C + + K +K+++L + AP LL Y Y +DI+
Sbjct: 95 GVEMGVIYSQVGKEQWTACKEKDLGACKSKAKTVQLKTPAPRPLLCGSY---YLTKEDIV 151
Query: 208 NWVDYKF 214
W K+
Sbjct: 152 PWSYSKY 158
>T1SD_MYCPN (P75435) Putative type I restriction enzyme specificity
protein MPN343 (S protein) (S.MpnORFDP) (H91_orf330)
Length = 330
Score = 32.3 bits (72), Expect = 1.5
Identities = 31/127 (24%), Positives = 51/127 (39%), Gaps = 13/127 (10%)
Query: 65 FYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLI 124
FYR FN V K+K K + + A PK + + +++ K S K +
Sbjct: 208 FYRNGKFNASQDCGVLKVKNKKICTKFLSFLLKIEA-----PKFVHNLASRPKLSQKVMA 262
Query: 125 N--------EYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKK 176
E QE D E C+D+++GI E D + + + +++ KK
Sbjct: 263 EIELSFPPLEIQEKIADILFAFEKLCNDLVEGIPAEIELRKKQLDYYQNFLFNWVQEQKK 322
Query: 177 SSKSIKL 183
+S S L
Sbjct: 323 NSLSTNL 329
>VC09_VACCV (P17372) Protein C9
Length = 634
Score = 31.2 bits (69), Expect = 3.4
Identities = 20/72 (27%), Positives = 34/72 (46%), Gaps = 8/72 (11%)
Query: 169 ELIRKLKKSSKSIKLVSIAPTELLKPHY------HKLYWANKDIINWVD--YKFYNQTVS 220
E I +K ++ + I T++L Y K Y N +W + YKFYNQ +
Sbjct: 532 EFITDCEKEIADMRQIKINGTDMLTVMYMLNKPTKKRYVNNPIFTDWANKQYKFYNQIIY 591
Query: 221 SADELVNLYNKL 232
+A++L+ K+
Sbjct: 592 NANKLIEQSKKI 603
>VC09_VACCC (P21042) Protein C9
Length = 634
Score = 31.2 bits (69), Expect = 3.4
Identities = 20/72 (27%), Positives = 34/72 (46%), Gaps = 8/72 (11%)
Query: 169 ELIRKLKKSSKSIKLVSIAPTELLKPHY------HKLYWANKDIINWVD--YKFYNQTVS 220
E I +K ++ + I T++L Y K Y N +W + YKFYNQ +
Sbjct: 532 EFITDCEKEIADMRQIKINGTDMLTVMYMLNKPTKKRYVNNPIFTDWANKQYKFYNQIIY 591
Query: 221 SADELVNLYNKL 232
+A++L+ K+
Sbjct: 592 NANKLIEQSKKI 603
>VATE_SULTO (Q971B8) V-type ATP synthase subunit E (EC 3.6.3.14)
(V-type ATPase subunit E)
Length = 194
Score = 31.2 bits (69), Expect = 3.4
Identities = 35/145 (24%), Positives = 61/145 (41%), Gaps = 17/145 (11%)
Query: 38 NDDIEEFHFILGTLREVYSGDGKGKGEFYRTWNFNNFSPAKVAKLKKDHKNVKVIISIGG 97
N EEF IL + ++ + E YR ++ AK+ L + N ++ I
Sbjct: 20 NKITEEFKKILSEMNQIID---EAYAEVYREYS------AKITDLVNKN-NDRIRGEIAK 69
Query: 98 FGAENP-FNPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYS 156
EN KE++ W K++ KK + E+ + + ++ DG I
Sbjct: 70 MEIENKRLISKEMDYWIENVKENAKKSLYEFVKTDNYKKGLESIISREVSDGSII----- 124
Query: 157 NCNPDEFSSCIGELIRKLKKSSKSI 181
C+P + S I ++I+K K S K +
Sbjct: 125 YCSPSDQKS-ISDIIKKKKISCKIV 148
>STC1_STAAU (P07767) Staphylocoagulase precursor
Length = 658
Score = 31.2 bits (69), Expect = 3.4
Identities = 37/190 (19%), Positives = 76/190 (39%), Gaps = 27/190 (14%)
Query: 90 KVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINE---YQEYSKDSSSTDECHCDDII 146
K IIS+G + + +W KA + K ++ E SK ++ + + II
Sbjct: 3 KQIISLGALAVAS-----SLFTWDNKADAIVTKDYSKESRVNEKSKKGATVSDYYYWKII 57
Query: 147 DGIDI----------NYEYSNCNPDEFSSCIGELIRKLKKSSKSIKLVSIAPTELLKPHY 196
D ++ NY+Y + + L+ ++ + + I EL K Y
Sbjct: 58 DSLEAQFTGAIDLLENYKYGD---PIYKEAKDRLMTRVLGEDQYLLKKKIDEYELYKKWY 114
Query: 197 HKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLLNEYGTDVKLLPGVST-----DPD 251
K N +++ + Y YN T++ +++ N + ++ +VK + + D D
Sbjct: 115 -KSSNKNTNMLTFHKYNLYNLTMNEYNDIFNSLKDAVYQFNKEVKEIEHKNVDLKQFDKD 173
Query: 252 SNTNMTRDVF 261
T++V+
Sbjct: 174 GEDKATKEVY 183
>SMC2_CHICK (Q90988) Structural maintenance of chromosome 2
(Chromosome scaffold protein ScII)
Length = 1189
Score = 30.8 bits (68), Expect = 4.5
Identities = 26/108 (24%), Positives = 51/108 (47%), Gaps = 14/108 (12%)
Query: 77 AKVAKLKK-DHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEYSKDSS 135
AK AK++K +N ++ +SI E+ N + E+ A ++ KL+ EY+ + +
Sbjct: 891 AKSAKIEKYREQNNELQLSINAL--EHDINKYQQET--ADASSTLDKLLKEYKWIASEK- 945
Query: 136 STDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRKLKKSSKSIKL 183
++ D Y++ NP E + +L+ K +K KS+ +
Sbjct: 946 --------ELFGQADTTYDFEANNPKETGQKLQKLLTKKEKLEKSLNM 985
>ICE8_HUMAN (Q14790) Caspase-8 precursor (EC 3.4.22.-) (CASP-8)
(ICE-like apoptotic protease 5) (MORT1-associated CED-3
homolog) (MACH) (FADD-homologous ICE/CED-3-like
protease) (FADD-like ICE) (FLICE) (Apoptotic cysteine
protease) (Apoptotic protease Mch
Length = 479
Score = 30.8 bits (68), Expect = 4.5
Identities = 13/34 (38%), Positives = 25/34 (73%)
Query: 118 QSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDI 151
+S+ K+IN+Y+E+SK+ SS+ E D+ +G ++
Sbjct: 169 KSLLKIINDYEEFSKERSSSLEGSPDEFSNGEEL 202
>CHIT_STRPL (P11220) Chitinase 63 precursor (EC 3.2.1.14)
Length = 610
Score = 30.8 bits (68), Expect = 4.5
Identities = 27/125 (21%), Positives = 56/125 (44%), Gaps = 25/125 (20%)
Query: 70 NFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQE 129
NFN ++ LK ++ ++K++ S GG+ F P +++ + AK S L+
Sbjct: 319 NFN-----QLRNLKAEYPHIKILYSFGGWTWSGGF-PDAVKNPAAFAK-SCHDLV----- 366
Query: 130 YSKDSSSTDECHCDDIIDGIDINYEYSN-----CNPDEFSSCIGELIRKLKKSSKSIKLV 184
++ D+ DGID+++EY N C+ + +++ ++ L+
Sbjct: 367 --------EDPRWADVFDGIDLDWEYPNACGLSCDETSAPNAFSSMMKAMRAEFGQDYLI 418
Query: 185 SIAPT 189
+ A T
Sbjct: 419 TAAVT 423
>CHI1_COCIM (P54196) Endochitinase 1 precursor (EC 3.2.1.14)
(Complement-fixation antigen) (CF-antigen) (CF-AG)
Length = 426
Score = 30.8 bits (68), Expect = 4.5
Identities = 22/75 (29%), Positives = 34/75 (45%), Gaps = 18/75 (24%)
Query: 82 LKKDHKNVKVIISIGGFGAENPF-NPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDEC 140
LKK+++N+K ++SIGG+ F P E K + KL+ +
Sbjct: 114 LKKNNRNLKTLLSIGGWTYSPNFKTPASTEEGRKKFADTSLKLMKDLG------------ 161
Query: 141 HCDDIIDGIDINYEY 155
DGIDI++EY
Sbjct: 162 -----FDGIDIDWEY 171
>CHI1_APHAL (P32470) Chitinase 1 precursor (EC 3.2.1.14)
Length = 423
Score = 30.8 bits (68), Expect = 4.5
Identities = 24/74 (32%), Positives = 35/74 (46%), Gaps = 16/74 (21%)
Query: 82 LKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECH 141
LKK ++N+KV++SIGG+ F P S +T+ K + KD
Sbjct: 115 LKKQNRNMKVMLSIGGWTWSTNF-PAAASSAATR-----KTFAQSAVGFMKDWG------ 162
Query: 142 CDDIIDGIDINYEY 155
DGIDI++EY
Sbjct: 163 ----FDGIDIDWEY 172
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,977,635
Number of Sequences: 164201
Number of extensions: 1798136
Number of successful extensions: 4143
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 4114
Number of HSP's gapped (non-prelim): 51
length of query: 309
length of database: 59,974,054
effective HSP length: 110
effective length of query: 199
effective length of database: 41,911,944
effective search space: 8340476856
effective search space used: 8340476856
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC130275.11


