

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.7 + phase: 0
(359 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
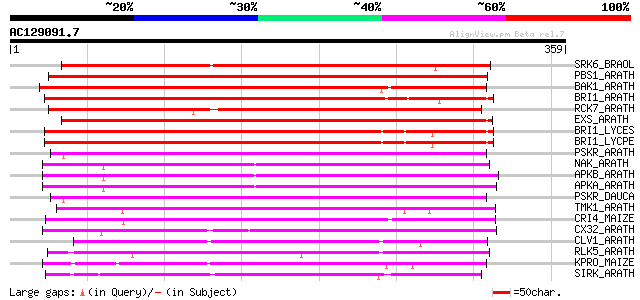
Score E
Sequences producing significant alignments: (bits) Value
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 244 2e-64
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 233 4e-61
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 230 5e-60
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 228 1e-59
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 226 7e-59
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 225 1e-58
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 221 2e-57
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 220 5e-57
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 217 3e-56
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 209 1e-53
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 209 1e-53
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 207 3e-53
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 203 5e-52
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 199 9e-51
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 194 3e-49
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 188 2e-47
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 185 2e-46
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 183 5e-46
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 182 9e-46
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 180 4e-45
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 244 bits (623), Expect = 2e-64
Identities = 127/287 (44%), Positives = 185/287 (64%), Gaps = 10/287 (3%)
Query: 34 ATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVKHSN 93
AT+N+ NK+G+GGFG VY+G L DG++IAVK LS S QG EF+ E+ ++ ++H N
Sbjct: 524 ATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKRLSKTSVQGTDEFMNEVTLIARLQHIN 583
Query: 94 LVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLAYLH 153
LV+++G CI+G + ++YEY+EN +L + L GK S K+ W ER I G A+GL YLH
Sbjct: 584 LVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKTRRS-KLNWNERFDITNGVARGLLYLH 642
Query: 154 EELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST-RIAGTTGYLAPEYA 212
++ I+HRD+K SN+LLDK+ PKI DFGMA++F D T +T ++ GT GY++PEYA
Sbjct: 643 QDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFERDETEANTMKVVGTYGYMSPEYA 702
Query: 213 LGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVDPEM 272
+ G ++K+DV+SFGV++LEI+SGK + LL + W +E + L +VDP +
Sbjct: 703 MYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYENDLLSYVWSRWKEGRALEIVDPVI 762
Query: 273 EE--------FPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKE 311
+ F +EV+K I++ L C Q A RP M+ VV M E
Sbjct: 763 VDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRPAMSSVVWMFGSE 809
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 233 bits (595), Expect = 4e-61
Identities = 125/288 (43%), Positives = 179/288 (61%), Gaps = 4/288 (1%)
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKD-GRKIAVKPLSVGSKQGVREFLTEIK 84
F+ +EL+ AT N+H +G GGFG VY+G L G+ +AVK L QG REFL E+
Sbjct: 74 FAFRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVL 133
Query: 85 TLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIG 144
LS + H NLV L+G+C G R +VYE++ G+L L + W R I G
Sbjct: 134 MLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAG 193
Query: 145 TAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP-DDITHISTRIAGT 203
AKGL +LH++ +++RD K+SN+LLD+ F+PK+ DFG+AKL P D +H+STR+ GT
Sbjct: 194 AAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRVMGT 253
Query: 204 TGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQL-HEEE 262
GY APEYA+ GQLT K+DVYSFGV+ LE+I+G+ + + ++L+ WA L ++
Sbjct: 254 YGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPLFNDRR 313
Query: 263 KWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLS 309
K++ L DP ++ FP + + + + VA C Q A RPL+ VV LS
Sbjct: 314 KFIKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVVTALS 361
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 230 bits (586), Expect = 5e-60
Identities = 122/294 (41%), Positives = 189/294 (63%), Gaps = 6/294 (2%)
Query: 20 LDNIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVR-E 78
L ++ FS +EL +A+DN+ N +GRGGFG VY+G L DG +AVK L QG +
Sbjct: 271 LGQLKRFSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGELQ 330
Query: 79 FLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRER 138
F TE++ +S H NL+ L GFC+ R +VY Y+ NG++ + L + + W +R
Sbjct: 331 FQTEVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWPKR 390
Query: 139 STICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST 198
I +G+A+GLAYLH+ I+HRD+KA+N+LLD++F +GDFG+AKL TH++T
Sbjct: 391 QRIALGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTT 450
Query: 199 RIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSS---SRTNWDGSHKSLLEWA 255
+ GT G++APEY G+ ++K DV+ +GV++LE+I+G+ + +R D LL+W
Sbjct: 451 AVRGTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLAND-DDVMLLDWV 509
Query: 256 WQLHEEEKWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L +E+K ALVD +++ + ++EV + I+VAL CTQ++ RP M++VV ML
Sbjct: 510 KGLLKEKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRML 563
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 228 bits (582), Expect = 1e-59
Identities = 127/295 (43%), Positives = 188/295 (63%), Gaps = 7/295 (2%)
Query: 23 IRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTE 82
+R + +L AT+ +H + IG GGFG VY+ LKDG +A+K L S QG REF+ E
Sbjct: 868 LRKLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQGDREFMAE 927
Query: 83 IKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTIC 142
++T+ +KH NLV L+G+C G R +VYE+++ G+L L K VK+ W R I
Sbjct: 928 METIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTRRKIA 987
Query: 143 IGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIS-TRIA 201
IG+A+GLA+LH + HI+HRD+K+SNVLLD++ ++ DFGMA+L TH+S + +A
Sbjct: 988 IGSARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLA 1047
Query: 202 GTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEE 261
GT GY+ PEY + + K DVYS+GV++LE+++GK + + D +L+ W Q H +
Sbjct: 1048 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSP-DFGDNNLVGWVKQ-HAK 1105
Query: 262 EKWLALVDPE-MEEFP--EKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQ 313
+ + DPE M+E P E E+++++KVA+ C A RRP M QV+ M KEIQ
Sbjct: 1106 LRISDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAMF-KEIQ 1159
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 226 bits (576), Expect = 7e-59
Identities = 118/288 (40%), Positives = 184/288 (62%), Gaps = 12/288 (4%)
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTL-KDGRKIAVKPLSVGSKQGVREFLTEIK 84
F+ +EL+ AT N+ +G GGFG V++GT+ K + +A+K L QG+REF+ E+
Sbjct: 91 FTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIREFVVEVL 150
Query: 85 TLSNVKHSNLVELVGFCIQGPNRTVVYEYVENG----NLHTALLGKKSLSVKMKWRERST 140
TLS H NLV+L+GFC +G R +VYEY+ G +LH GKK L W R
Sbjct: 151 TLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLD----WNTRMK 206
Query: 141 ICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPD-DITHISTR 199
I G A+GL YLH+ +T +++RD+K SN+LL +D+ PK+ DFG+AK+ P D TH+STR
Sbjct: 207 IAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVSTR 266
Query: 200 IAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLH 259
+ GT GY AP+YA+ GQLT K+D+YSFGV++LE+I+G+ + ++L+ WA L
Sbjct: 267 VMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQNLVGWARPLF 326
Query: 260 EEEK-WLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVV 305
++ + + +VDP ++ ++P + + + + ++ C Q RP+++ VV
Sbjct: 327 KDRRNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVV 374
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 225 bits (574), Expect = 1e-58
Identities = 120/281 (42%), Positives = 173/281 (60%), Gaps = 3/281 (1%)
Query: 34 ATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVKHSN 93
ATD++ N IG GGFGTVY+ L + +AVK LS QG REF+ E++TL VKH N
Sbjct: 913 ATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQGNREFMAEMETLGKVKHPN 972
Query: 94 LVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLAYLH 153
LV L+G+C + +VYEY+ NG+L L + + + W +R I +G A+GLA+LH
Sbjct: 973 LVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGAARGLAFLH 1032
Query: 154 EELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTGYLAPEYAL 213
HI+HRDIKASN+LLD DF PK+ DFG+A+L +H+ST IAGT GY+ PEY
Sbjct: 1033 HGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTVIAGTFGYIPPEYGQ 1092
Query: 214 GGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSH-KSLLEWAWQLHEEEKWLALVDPEM 272
+ T K DVYSFGV++LE+++GK + ++ S +L+ WA Q + K + ++DP +
Sbjct: 1093 SARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGNLVGWAIQKINQGKAVDVIDPLL 1152
Query: 273 EEFPEK-EVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEI 312
K ++ +++A+ C +RP M V+ L KEI
Sbjct: 1153 VSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKAL-KEI 1192
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 221 bits (563), Expect = 2e-57
Identities = 121/295 (41%), Positives = 183/295 (62%), Gaps = 7/295 (2%)
Query: 23 IRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTE 82
+R + +L AT+ +H + +G GGFG VY+ LKDG +A+K L S QG REF E
Sbjct: 873 LRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAE 932
Query: 83 IKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTIC 142
++T+ +KH NLV L+G+C G R +VYEY++ G+L L +K + +K+ W R I
Sbjct: 933 METIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIA 992
Query: 143 IGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIS-TRIA 201
IG A+GLA+LH HI+HRD+K+SNVLLD++ ++ DFGMA+L TH+S + +A
Sbjct: 993 IGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLA 1052
Query: 202 GTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEE 261
GT GY+ PEY + + K DVYS+GV++LE+++GK + + D +L+ W +LH +
Sbjct: 1053 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPT-DSADFGDNNLVGWV-KLHAK 1110
Query: 262 EKWLALVDPEM---EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQ 313
K + D E+ + E E+++++KVA C +RP M QV+ M KEIQ
Sbjct: 1111 GKITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMF-KEIQ 1164
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 220 bits (560), Expect = 5e-57
Identities = 121/295 (41%), Positives = 182/295 (61%), Gaps = 7/295 (2%)
Query: 23 IRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTE 82
+R + +L AT+ +H + +G GGFG VY+ LKDG +A+K L S QG REF E
Sbjct: 873 LRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAE 932
Query: 83 IKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTIC 142
++T+ +KH NLV L+G+C G R +VYEY++ G+L L +K +K+ W R I
Sbjct: 933 METIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIA 992
Query: 143 IGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIS-TRIA 201
IG A+GLA+LH HI+HRD+K+SNVLLD++ ++ DFGMA+L TH+S + +A
Sbjct: 993 IGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLA 1052
Query: 202 GTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEE 261
GT GY+ PEY + + K DVYS+GV++LE+++GK + + D +L+ W +LH +
Sbjct: 1053 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPT-DSADFGDNNLVGWV-KLHAK 1110
Query: 262 EKWLALVDPEM---EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQ 313
K + D E+ + E E+++++KVA C +RP M QV+ M KEIQ
Sbjct: 1111 GKITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMF-KEIQ 1164
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 217 bits (553), Expect = 3e-56
Identities = 117/288 (40%), Positives = 170/288 (58%), Gaps = 6/288 (2%)
Query: 27 SDKELSL-----ATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLT 81
+DKELS +T+++ N IG GGFG VY+ TL DG+K+A+K LS Q REF
Sbjct: 718 NDKELSYDDLLDSTNSFDQANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQIEREFEA 777
Query: 82 EIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTI 141
E++TLS +H NLV L GFC +R ++Y Y+ENG+L L + +KW+ R I
Sbjct: 778 EVETLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGPALLKWKTRLRI 837
Query: 142 CIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIA 201
G AKGL YLHE HI+HRDIK+SN+LLD++FN + DFG+A+L TH+ST +
Sbjct: 838 AQGAAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVSTDLV 897
Query: 202 GTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEE 261
GT GY+ PEY T K DVYSFGV++LE+++ K + L+ W ++ E
Sbjct: 898 GTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMKHE 957
Query: 262 EKWLALVDPEM-EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
+ + DP + + +KE+ + +++A C ++RP Q+V L
Sbjct: 958 SRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLVSWL 1005
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 209 bits (531), Expect = 1e-53
Identities = 113/302 (37%), Positives = 181/302 (59%), Gaps = 14/302 (4%)
Query: 22 NIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKD----------GRKIAVKPLSVG 71
N+++FS EL AT N+ + +G GGFG V++G + + G IAVK L+
Sbjct: 52 NLKNFSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLNQE 111
Query: 72 SKQGVREFLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSV 131
QG RE+L EI L + H NLV+L+G+C++ +R +VYE++ G+L L + +
Sbjct: 112 GFQGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFYQ 171
Query: 132 KMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP- 190
+ W R + +G A+GLA+LH Q +++RD KASN+LLD ++N K+ DFG+A+ P
Sbjct: 172 PLSWNTRVRMALGAARGLAFLHNAQPQ-VIYRDFKASNILLDSNYNAKLSDFGLARDGPM 230
Query: 191 DDITHISTRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKS 250
D +H+STR+ GT GY APEY G L+ K+DVYSFGV++LE++SG+ + N +
Sbjct: 231 GDNSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVGEHN 290
Query: 251 LLEWAW-QLHEEEKWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L++WA L + + L ++DP ++ ++ +K +AL C A+ RP M ++V +
Sbjct: 291 LVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAKSRPTMNEIVKTM 350
Query: 309 SK 310
+
Sbjct: 351 EE 352
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 209 bits (531), Expect = 1e-53
Identities = 117/308 (37%), Positives = 179/308 (57%), Gaps = 14/308 (4%)
Query: 22 NIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKD----------GRKIAVKPLSVG 71
N++ F+ EL AT N+ + +G GGFG+V++G + + G IAVK L+
Sbjct: 53 NLKSFTFAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTASKPGTGVVIAVKKLNQD 112
Query: 72 SKQGVREFLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSV 131
QG +E+L E+ L H NLV+L+G+C++ +R +VYE++ G+L L + S
Sbjct: 113 GWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGSYFQ 172
Query: 132 KMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP- 190
+ W R + +G AKGLA+LH T +++RD K SN+LLD ++N K+ DFG+AK P
Sbjct: 173 PLSWTLRLKVALGAAKGLAFLHNAETS-VIYRDFKTSNILLDSEYNAKLSDFGLAKDGPT 231
Query: 191 DDITHISTRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKS 250
D +H+STRI GT GY APEY G LT K+DVYS+GV++LE++SG+ + N +
Sbjct: 232 GDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSGRRAVDKNRPPGEQK 291
Query: 251 LLEWAWQ-LHEEEKWLALVDPEM-EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L+EWA L + K ++D + +++ +E K +AL C + RP M +VV L
Sbjct: 292 LVEWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRCLTFEIKLRPNMNEVVSHL 351
Query: 309 SKEIQLND 316
LN+
Sbjct: 352 EHIQTLNE 359
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 207 bits (528), Expect = 3e-53
Identities = 115/307 (37%), Positives = 180/307 (58%), Gaps = 14/307 (4%)
Query: 22 NIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKD----------GRKIAVKPLSVG 71
N++ FS EL AT N+ + +G GGFG V++G + + G IAVK L+
Sbjct: 52 NLKSFSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGLVIAVKKLNQD 111
Query: 72 SKQGVREFLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSV 131
QG +E+L E+ L H +LV+L+G+C++ +R +VYE++ G+L L +
Sbjct: 112 GWQGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGLYFQ 171
Query: 132 KMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP- 190
+ W+ R + +G AKGLA+LH T+ +++RD K SN+LLD ++N K+ DFG+AK P
Sbjct: 172 PLSWKLRLKVALGAAKGLAFLHSSETR-VIYRDFKTSNILLDSEYNAKLSDFGLAKDGPI 230
Query: 191 DDITHISTRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKS 250
D +H+STR+ GT GY APEY G LT K+DVYSFGV++LE++SG+ + N ++
Sbjct: 231 GDKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRPSGERN 290
Query: 251 LLEWAW-QLHEEEKWLALVDPEM-EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L+EWA L + K ++D + +++ +E K ++L C + RP M++VV L
Sbjct: 291 LVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSEVVSHL 350
Query: 309 SKEIQLN 315
LN
Sbjct: 351 EHIQSLN 357
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 203 bits (517), Expect = 5e-52
Identities = 110/288 (38%), Positives = 168/288 (58%), Gaps = 6/288 (2%)
Query: 27 SDKELSL-----ATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLT 81
S+ ELSL +T +++ N IG GGFG VY+ TL DG K+A+K LS + Q REF
Sbjct: 727 SNNELSLDDILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQMDREFQA 786
Query: 82 EIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTI 141
E++TLS +H NLV L+G+C ++ ++Y Y++NG+L L K + W+ R I
Sbjct: 787 EVETLSRAQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLRI 846
Query: 142 CIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIA 201
G A+GLAYLH+ HI+HRDIK+SN+LL F + DFG+A+L TH++T +
Sbjct: 847 ARGAAEGLAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTHVTTDLV 906
Query: 202 GTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEE 261
GT GY+ PEY T K DVYSFGV++LE+++G+ + L+ W Q+ E
Sbjct: 907 GTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWVLQMKTE 966
Query: 262 EKWLALVDPEM-EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
++ + DP + ++ +E++ +++A C + RP Q+V L
Sbjct: 967 KRESEIFDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWL 1014
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 199 bits (506), Expect = 9e-51
Identities = 111/291 (38%), Positives = 169/291 (57%), Gaps = 7/291 (2%)
Query: 31 LSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVG--SKQGVREFLTEIKTLSN 88
L T+N+ N +G GGFG VY+G L DG KIAVK + G + +G EF +EI L+
Sbjct: 581 LRSVTNNFSSDNILGSGGFGVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIAVLTK 640
Query: 89 VKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVK-MKWRERSTICIGTAK 147
V+H +LV L+G+C+ G + +VYEY+ G L L +K + W++R T+ + A+
Sbjct: 641 VRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRLTLALDVAR 700
Query: 148 GLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTGYL 207
G+ YLH Q +HRD+K SN+LL D K+ DFG+ +L P+ I TRIAGT GYL
Sbjct: 701 GVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIETRIAGTFGYL 760
Query: 208 APEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEW--AWQLHEEEKWL 265
APEYA+ G++T K DVYSFGV+++E+I+G+ S + L+ W +++E +
Sbjct: 761 APEYAVTGRVTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSWFKRMYINKEASFK 820
Query: 266 ALVDP--EMEEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQL 314
+D +++E V ++A C +RP M V++LS ++L
Sbjct: 821 KAIDTTIDLDEETLASVHTVAELAGHCCAREPYQRPDMGHAVNILSSLVEL 871
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 194 bits (493), Expect = 3e-49
Identities = 111/296 (37%), Positives = 175/296 (58%), Gaps = 7/296 (2%)
Query: 24 RHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGS--KQGVREFLT 81
+ FS +EL AT + +++G+G F V++G L+DG +AVK S K+ +EF
Sbjct: 491 QEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASDVKKSSKEFHN 550
Query: 82 EIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKK-SLSVKMKWRERST 140
E+ LS + H++L+ L+G+C G R +VYE++ +G+L+ L GK +L ++ W R T
Sbjct: 551 ELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKKRLNWARRVT 610
Query: 141 ICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP-DDITHISTR 199
I + A+G+ YLH ++HRDIK+SN+L+D+D N ++ DFG++ L P D T +S
Sbjct: 611 IAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPADSGTPLSEL 670
Query: 200 IAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLH 259
AGT GYL PEY LT K+DVYSFGV++LEI+SG+ + ++ +++EWA L
Sbjct: 671 PAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFE--EGNIVEWAVPLI 728
Query: 260 EEEKWLALVDPEMEEFPEKEVIKYI-KVALFCTQAAARRRPLMTQVVDMLSKEIQL 314
+ A++DP + + E +K I VA C + + RP M +V L + L
Sbjct: 729 KAGDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKVTTALEHALAL 784
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 188 bits (477), Expect = 2e-47
Identities = 107/307 (34%), Positives = 180/307 (57%), Gaps = 17/307 (5%)
Query: 22 NIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLK----------DGRKIAVKPLSVG 71
N++ ++ +L AT N+ + +G+GGFG VY+G + G +A+K L+
Sbjct: 70 NLKVYNFLDLKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRLNSE 129
Query: 72 SKQGVREFLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSV 131
S QG E+ +E+ L + H NLV+L+G+C + +VYE++ G+L + L +
Sbjct: 130 SVQGFAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLESHLFRRND--- 186
Query: 132 KMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP- 190
W R I IG A+GLA+LH L + +++RD KASN+LLD +++ K+ DFG+AKL P
Sbjct: 187 PFPWDLRIKIVIGAARGLAFLHS-LQREVIYRDFKASNILLDSNYDAKLSDFGLAKLGPA 245
Query: 191 DDITHISTRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKS 250
D+ +H++TRI GT GY APEY G L K+DV++FGV++LEI++G ++ T +S
Sbjct: 246 DEKSHVTTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTAHNTKRPRGQES 305
Query: 251 LLEWAW-QLHEEEKWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L++W +L + + ++D ++ ++ K + ++ L C + + RP M +VV++L
Sbjct: 306 LVDWLRPELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDPKNRPHMKEVVEVL 365
Query: 309 SKEIQLN 315
LN
Sbjct: 366 EHIQGLN 372
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 185 bits (469), Expect = 2e-46
Identities = 102/277 (36%), Positives = 158/277 (56%), Gaps = 13/277 (4%)
Query: 42 NKIGRGGFGTVYQGTLKDGRKIAVKPL-SVGSKQGVREFLTEIKTLSNVKHSNLVELVGF 100
N IG+GG G VY+G++ + +A+K L G+ + F EI+TL ++H ++V L+G+
Sbjct: 696 NIIGKGGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRLLGY 755
Query: 101 CIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLAYLHEELTQHI 160
++YEY+ NG+L L G K ++W R + + AKGL YLH + + I
Sbjct: 756 VANKDTNLLLYEYMPNGSLGELLHGSKG--GHLQWETRHRVAVEAAKGLCYLHHDCSPLI 813
Query: 161 VHRDIKASNVLLDKDFNPKIGDFGMAKLFPDD-ITHISTRIAGTTGYLAPEYALGGQLTK 219
+HRD+K++N+LLD DF + DFG+AK D + + IAG+ GY+APEYA ++ +
Sbjct: 814 LHRDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAYTLKVDE 873
Query: 220 KADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKW-------LALVDPEM 272
K+DVYSFGV++LE+I+GK G ++ W EE +A+VDP +
Sbjct: 874 KSDVYSFGVVLLELIAGKKP--VGEFGEGVDIVRWVRNTEEEITQPSDAAIVVAIVDPRL 931
Query: 273 EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLS 309
+P VI K+A+ C + A RP M +VV ML+
Sbjct: 932 TGYPLTSVIHVFKIAMMCVEEEAAARPTMREVVHMLT 968
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 183 bits (465), Expect = 5e-46
Identities = 106/299 (35%), Positives = 166/299 (55%), Gaps = 18/299 (6%)
Query: 25 HFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVRE------ 78
HFS+ E++ D N IG G G VY+ L+ G +AVK L+ K G E
Sbjct: 673 HFSEHEIADCLDEK---NVIGFGSSGKVYKVELRGGEVVAVKKLNKSVKGGDDEYSSDSL 729
Query: 79 ----FLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMK 134
F E++TL ++H ++V L C G + +VYEY+ NG+L L G + V +
Sbjct: 730 NRDVFAAEVETLGTIRHKSIVRLWCCCSSGDCKLLVYEYMPNGSLADVLHGDRKGGVVLG 789
Query: 135 WRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAK---LFPD 191
W ER I + A+GL+YLH + IVHRD+K+SN+LLD D+ K+ DFG+AK +
Sbjct: 790 WPERLRIALDAAEGLSYLHHDCVPPIVHRDVKSSNILLDSDYGAKVADFGIAKVGQMSGS 849
Query: 192 DITHISTRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSL 251
+ IAG+ GY+APEY ++ +K+D+YSFGV++LE+++GK T+ + K +
Sbjct: 850 KTPEAMSGIAGSCGYIAPEYVYTLRVNEKSDIYSFGVVLLELVTGKQP--TDSELGDKDM 907
Query: 252 LEWAWQLHEEEKWLALVDPEMEEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSK 310
+W ++ ++DP+++ ++E+ K I + L CT RP M +VV ML +
Sbjct: 908 AKWVCTALDKCGLEPVIDPKLDLKFKEEISKVIHIGLLCTSPLPLNRPSMRKVVIMLQE 966
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 182 bits (463), Expect = 9e-46
Identities = 110/297 (37%), Positives = 162/297 (54%), Gaps = 14/297 (4%)
Query: 22 NIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLT 81
N R +S +EL AT + + ++GRG GTVY+G L+D R +AVK L +QG F
Sbjct: 520 NFRRYSYRELVKATRKFKV--ELGRGESGTVYKGVLEDDRHVAVKKLE-NVRQGKEVFQA 576
Query: 82 EIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTI 141
E+ + + H NLV + GFC +G +R +V EYVENG+L L + ++ + W R I
Sbjct: 577 ELSVIGRINHMNLVRIWGFCSEGSHRLLVSEYVENGSLANILFSEGG-NILLDWEGRFNI 635
Query: 142 CIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLF-PDDITHISTRI 200
+G AKGLAYLH E + ++H D+K N+LLD+ F PKI DFG+ KL T + +
Sbjct: 636 ALGVAKGLAYLHHECLEWVIHCDVKPENILLDQAFEPKITDFGLVKLLNRGGSTQNVSHV 695
Query: 201 AGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRT--NWDGSHKSLLEWAWQL 258
GT GY+APE+ +T K DVYS+GV++LE+++G S D H L + L
Sbjct: 696 RGTLGYIAPEWVSSLPITAKVDVYSYGVVLLELLTGTRVSELVGGTDEVHSMLRKLVRML 755
Query: 259 H-----EEEKWL-ALVDPEMEE-FPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
EE+ W+ +D ++ + IK+A+ C + +RP M V L
Sbjct: 756 SAKLEGEEQSWIDGYLDSKLNRPVNYVQARTLIKLAVSCLEEDRSKRPTMEHAVQTL 812
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 180 bits (457), Expect = 4e-45
Identities = 109/287 (37%), Positives = 165/287 (56%), Gaps = 14/287 (4%)
Query: 24 RHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEI 83
R+F E+ T+N+ IG+GGFG VY G + +G ++AVK LS S QG +EF E+
Sbjct: 562 RYFKYSEVVNITNNFE--RVIGKGGFGKVYHGVI-NGEQVAVKVLSEESAQGYKEFRAEV 618
Query: 84 KTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICI 143
L V H+NL LVG+C + + ++YEY+ N NL L GK+S + W ER I +
Sbjct: 619 DLLMRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSFI--LSWEERLKISL 676
Query: 144 GTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP-DDITHISTRIAG 202
A+GL YLH IVHRD+K +N+LL++ K+ DFG+++ F + IST +AG
Sbjct: 677 DAAQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQISTVVAG 736
Query: 203 TTGYLAPEYALGGQLTKKADVYSFGVLILEIISGK---SSSRTNWDGSHKSLLEWAWQLH 259
+ GYL PEY Q+ +K+DVYS GV++LE+I+G+ +SS+T + + +
Sbjct: 737 SIGYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKT----EKVHISDHVRSIL 792
Query: 260 EEEKWLALVDPEM-EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVV 305
+VD + E + K ++AL CT+ + +RP M+QVV
Sbjct: 793 ANGDIRGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVV 839
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,919,674
Number of Sequences: 164201
Number of extensions: 1828254
Number of successful extensions: 8699
Number of sequences better than 10.0: 1658
Number of HSP's better than 10.0 without gapping: 1243
Number of HSP's successfully gapped in prelim test: 415
Number of HSP's that attempted gapping in prelim test: 4638
Number of HSP's gapped (non-prelim): 1915
length of query: 359
length of database: 59,974,054
effective HSP length: 111
effective length of query: 248
effective length of database: 41,747,743
effective search space: 10353440264
effective search space used: 10353440264
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC129091.7


