

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.5 - phase: 0 /pseudo
(980 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
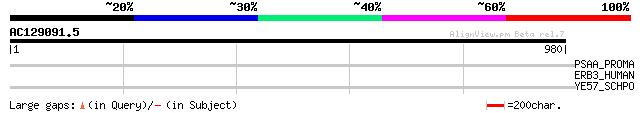
Score E
Sequences producing significant alignments: (bits) Value
PSAA_PROMA (Q9L4N4) Photosystem I P700 chlorophyll A apoprotein ... 33 3.7
ERB3_HUMAN (P21860) Receptor tyrosine-protein kinase erbB-3 prec... 33 3.7
YE57_SCHPO (O14173) Hypothetical protein C4D7.07c in chromosome I 33 4.9
>PSAA_PROMA (Q9L4N4) Photosystem I P700 chlorophyll A apoprotein A1
(PsaA)
Length = 773
Score = 33.1 bits (74), Expect = 3.7
Identities = 22/71 (30%), Positives = 31/71 (42%), Gaps = 4/71 (5%)
Query: 379 WESKVDISIPATDDKSKGIHFNPFDLPKYLGTL---ARVVFSAQGGEIAVAFFQGRVRIF 435
W SK+ P T +H + D +LG L +R +FSA G +AV F F
Sbjct: 42 WSSKLSKG-PKTTTWIWNLHADAHDFDTHLGDLEETSRKIFSAHFGHLAVVFIWMSAAFF 100
Query: 436 SGSNFEPVTNY 446
G+ F T +
Sbjct: 101 HGARFSNYTGW 111
>ERB3_HUMAN (P21860) Receptor tyrosine-protein kinase erbB-3 precursor
(EC 2.7.1.112) (c-erbB3) (Tyrosine kinase-type cell
surface receptor HER3)
Length = 1342
Score = 33.1 bits (74), Expect = 3.7
Identities = 44/151 (29%), Positives = 59/151 (38%), Gaps = 14/151 (9%)
Query: 775 GSKGGEEPSPGPVRLGNGNAGQGYSVEEVKVIFQVLLDLCRRTSGLQHPLPVSQVGSSNI 834
GS+ PS G + + GN G+ S +E V + C R L HP+P + S +
Sbjct: 1043 GSQSLLSPSSGYMPMNQGNLGE--SCQESAVSGSS--ERCPRPVSL-HPMPRGCLASESS 1097
Query: 835 QVQLHYIEGSYTVLPEVVEASLGPYMQNMPR-LGDA----DDTGLLLRELQLHPPAEEWH 889
+ ++ GS L E V PR GD+ LL L PP E
Sbjct: 1098 E---GHVTGSEAELQEKVSMCRSRSRSRSPRPRGDSAYHSQRHSLLTPVTPLSPPGLEEE 1154
Query: 890 QLNMFVRPCTDSNDTPKPFRSNPLDSRSLES 920
+N +V P T TP R L S L S
Sbjct: 1155 DVNGYVMPDTHLKGTPSS-REGTLSSVGLSS 1184
>YE57_SCHPO (O14173) Hypothetical protein C4D7.07c in chromosome I
Length = 600
Score = 32.7 bits (73), Expect = 4.9
Identities = 28/106 (26%), Positives = 46/106 (42%), Gaps = 12/106 (11%)
Query: 606 INPSALLPDPWQASEEILSNFDMEEMAVEPELTPCIQAYVDSV---IDLASHLITRLRHY 662
I+P +L W + ++S +D++ + + P + Y+D + I L SHL +H
Sbjct: 201 ISPEDVLLHFWDRN--LMSTYDLKIQDINDSVNPLLNTYLDFIEKDIYLVSHLPVSEKHP 258
Query: 663 AKIFRTLANQAVTVASGSTSGGVPNLTLNPSISGPSSLMLISINTG 708
I +L N++V NL NPS G + IN G
Sbjct: 259 GNIPISLVNKSVQAICSFAEH--YNLLRNPSYRG-----FLRINNG 297
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 115,740,043
Number of Sequences: 164201
Number of extensions: 4998359
Number of successful extensions: 13451
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 13447
Number of HSP's gapped (non-prelim): 7
length of query: 980
length of database: 59,974,054
effective HSP length: 120
effective length of query: 860
effective length of database: 40,269,934
effective search space: 34632143240
effective search space used: 34632143240
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC129091.5


