

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.3 + phase: 0
(170 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
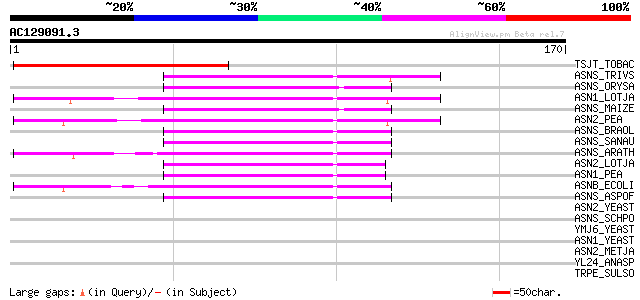
Score E
Sequences producing significant alignments: (bits) Value
TSJT_TOBAC (P24805) Stem-specific protein TSJT1 81 1e-15
ASNS_TRIVS (O24661) Asparagine synthetase [glutamine-hydrolyzing... 49 6e-06
ASNS_ORYSA (Q43011) Asparagine synthetase [glutamine-hydrolyzing... 48 1e-05
ASN1_LOTJA (P49092) Asparagine synthetase [glutamine-hydrolyzing... 47 2e-05
ASNS_MAIZE (P49094) Asparagine synthetase [glutamine-hydrolyzing... 47 3e-05
ASN2_PEA (P19252) Asparagine synthetase, root [glutamine-hydroly... 45 1e-04
ASNS_BRAOL (P49091) Asparagine synthetase [glutamine-hydrolyzing... 44 1e-04
ASNS_SANAU (O24338) Asparagine synthetase [glutamine-hydrolyzing... 44 2e-04
ASNS_ARATH (P49078) Asparagine synthetase [glutamine-hydrolyzing... 44 2e-04
ASN2_LOTJA (P49093) Asparagine synthetase [glutamine-hydrolyzing... 44 2e-04
ASN1_PEA (P19251) Asparagine synthetase, nodule [glutamine-hydro... 43 3e-04
ASNB_ECOLI (P22106) Asparagine synthetase B [glutamine-hydrolyzi... 43 4e-04
ASNS_ASPOF (P31752) Asparagine synthetase [glutamine-hydrolyzing... 42 5e-04
ASN2_YEAST (P49090) Asparagine synthetase [glutamine-hydrolyzing... 37 0.022
ASNS_SCHPO (P78753) Probable asparagine synthetase [glutamine-hy... 37 0.029
YMJ6_YEAST (Q04489) Hypothetical 59.5 kDa protein in VPS9-RAD10 ... 35 0.064
ASN1_YEAST (P49089) Asparagine synthetase [glutamine-hydrolyzing... 33 0.41
ASN2_METJA (Q58456) Putative asparagine synthetase [glutamine-hy... 30 3.5
YL24_ANASP (Q8YV57) Hypothetical WD-repeat protein all2124 29 6.0
TRPE_SULSO (Q06128) Anthranilate synthase component I (EC 4.1.3.27) 29 6.0
>TSJT_TOBAC (P24805) Stem-specific protein TSJT1
Length = 149
Score = 80.9 bits (198), Expect = 1e-15
Identities = 37/66 (56%), Positives = 46/66 (69%)
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G L N L K YGLS+ EAM ++EAY+ LRDR PYP DQV+K LEG FAF+++D K
Sbjct: 83 GALDNTFDLRKHYGLSRQATEAMIMVEAYKVLRDRAPYPPDQVIKELEGKFAFILFDSKA 142
Query: 62 GTVFAA 67
T+F A
Sbjct: 143 STLFLA 148
>ASNS_TRIVS (O24661) Asparagine synthetase [glutamine-hydrolyzing]
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase)
Length = 585
Score = 48.9 bits (115), Expect = 6e-06
Identities = 31/86 (36%), Positives = 47/86 (54%), Gaps = 2/86 (2%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D +D T AA + G LY G DGSV IS L+ + C ++F FP
Sbjct: 116 LDGMFSFVLLDSRDNTFIAARDAFGITSLYIGWGLDGSVWISSELKGLHDEC-ENFEVFP 174
Query: 108 TGCLFHSE-HGLLNFEHPTKKMKAMP 132
G ++ S+ G + +P +A+P
Sbjct: 175 PGHVYSSKTEGFRRWYNPPWFSEAIP 200
>ASNS_ORYSA (Q43011) Asparagine synthetase [glutamine-hydrolyzing]
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase)
Length = 590
Score = 47.8 bits (112), Expect = 1e-05
Identities = 27/70 (38%), Positives = 38/70 (53%), Gaps = 1/70 (1%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G FAFV+ D +D + AA + G LY G DGSV S ++ + C + F FP
Sbjct: 116 LDGMFAFVLLDTRDKSFIAARDAIGICPLYMGWGLDGSVWFSSEMKALSDDCER-FISFP 174
Query: 108 TGCLFHSEHG 117
G L+ S+ G
Sbjct: 175 PGHLYSSKTG 184
>ASN1_LOTJA (P49092) Asparagine synthetase [glutamine-hydrolyzing] 1
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase
1)
Length = 585
Score = 47.4 bits (111), Expect = 2e-05
Identities = 38/134 (28%), Positives = 61/134 (45%), Gaps = 11/134 (8%)
Query: 2 GNLHNLSQLNKQYGLS--KGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDH 59
G ++N +L KQ + G++ I Y + + L+G F+FV+ D
Sbjct: 75 GEIYNHEELRKQLPNHQFRTGSDCDVIAHLYEE-------HGENFMDMLDGIFSFVLLDT 127
Query: 60 KDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHS-EHGL 118
+D T A + G LY G DGSV IS ++ + C + F FP G L+ S E
Sbjct: 128 RDNTFIVARDAIGVTSLYIGWGLDGSVWISSEMKGLNDDC-EHFEVFPPGHLYSSRERAF 186
Query: 119 LNFEHPTKKMKAMP 132
+ +PT +++P
Sbjct: 187 RRWYNPTWFSESIP 200
>ASNS_MAIZE (P49094) Asparagine synthetase [glutamine-hydrolyzing]
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase)
Length = 585
Score = 46.6 bits (109), Expect = 3e-05
Identities = 26/70 (37%), Positives = 38/70 (54%), Gaps = 1/70 (1%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D +D + AA + G LY G DGSV S ++ + C + F FP
Sbjct: 116 LDGMFSFVLLDTRDKSFIAARDAIGICPLYMGWGLDGSVWFSSEMKALSDDCER-FITFP 174
Query: 108 TGCLFHSEHG 117
G L+ S+ G
Sbjct: 175 PGHLYSSKTG 184
>ASN2_PEA (P19252) Asparagine synthetase, root
[glutamine-hydrolyzing] (EC 6.3.5.4)
(Glutamine-dependent asparagine synthetase)
Length = 582
Score = 44.7 bits (104), Expect = 1e-04
Identities = 37/134 (27%), Positives = 60/134 (44%), Gaps = 11/134 (8%)
Query: 2 GNLHNLSQLNKQYG--LSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDH 59
G ++N L KQ + G++ I Y + + L+G F+FV D
Sbjct: 75 GEIYNHEDLRKQLSNHTFRTGSDCDVIAHLYEEY-------GEDFVDMLDGIFSFVPLDT 127
Query: 60 KDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHS-EHGL 118
+D + A + G LY G DGSV IS ++ + C + F FP G L+ S + G
Sbjct: 128 RDNSYIVARDAIGVTSLYIGWGLDGSVWISSEMKGLNDDC-EHFECFPPGHLYSSKDSGF 186
Query: 119 LNFEHPTKKMKAMP 132
+ +P+ +A+P
Sbjct: 187 RRWYNPSWYSEAIP 200
>ASNS_BRAOL (P49091) Asparagine synthetase [glutamine-hydrolyzing]
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase)
Length = 585
Score = 44.3 bits (103), Expect = 1e-04
Identities = 24/70 (34%), Positives = 38/70 (54%), Gaps = 1/70 (1%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D +D + A + G LY G DGS+ +S ++ + C + F FP
Sbjct: 116 LDGIFSFVLLDTRDNSFMVARDAVGVTSLYIGWGLDGSLWVSSEMKGLHEDC-EHFEAFP 174
Query: 108 TGCLFHSEHG 117
G L+ S+ G
Sbjct: 175 PGHLYSSKSG 184
>ASNS_SANAU (O24338) Asparagine synthetase [glutamine-hydrolyzing]
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase)
Length = 524
Score = 43.9 bits (102), Expect = 2e-04
Identities = 26/70 (37%), Positives = 39/70 (55%), Gaps = 1/70 (1%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D ++ + AA + G LY G DGSV IS ++ + C + F FP
Sbjct: 116 LDGIFSFVLLDSRNNSFVAARDAIGVTPLYIGWGLDGSVWISSEMKGLNDDC-EHFKFFP 174
Query: 108 TGCLFHSEHG 117
G L+ S+ G
Sbjct: 175 PGHLYSSKEG 184
>ASNS_ARATH (P49078) Asparagine synthetase [glutamine-hydrolyzing]
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase)
Length = 583
Score = 43.9 bits (102), Expect = 2e-04
Identities = 36/118 (30%), Positives = 54/118 (45%), Gaps = 10/118 (8%)
Query: 2 GNLHNLSQLNKQYGLSK--GGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDH 59
G ++N +L K+ K G++ I Y Y D V L+G F+FV+ D
Sbjct: 75 GEIYNHEELRKRLKNHKFRTGSDCEVIAHLYEE------YGVDFV-DMLDGIFSFVLLDT 127
Query: 60 KDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHG 117
+D + A + G LY G DGSV IS ++ + C + F FP G + S+ G
Sbjct: 128 RDNSFMVARDAIGVTSLYIGWGLDGSVWISSEMKGLNDDC-EHFETFPPGHFYSSKLG 184
>ASN2_LOTJA (P49093) Asparagine synthetase [glutamine-hydrolyzing] 2
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase
2)
Length = 585
Score = 43.5 bits (101), Expect = 2e-04
Identities = 25/68 (36%), Positives = 37/68 (53%), Gaps = 1/68 (1%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D +D + A + G LY G DGSV I+ L+ + C + F FP
Sbjct: 116 LDGIFSFVLLDTRDNSFLVARDAIGVTSLYIGYGLDGSVWIASELKGLNDDC-EHFELFP 174
Query: 108 TGCLFHSE 115
G L+ S+
Sbjct: 175 PGHLYSSK 182
>ASN1_PEA (P19251) Asparagine synthetase, nodule
[glutamine-hydrolyzing] (EC 6.3.5.4)
(Glutamine-dependent asparagine synthetase)
Length = 585
Score = 43.1 bits (100), Expect = 3e-04
Identities = 25/68 (36%), Positives = 37/68 (53%), Gaps = 1/68 (1%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D +D + A + G LY G DGSV I+ L+ + C + F FP
Sbjct: 116 LDGIFSFVLLDTRDNSFIVARDAIGVTSLYIGWGLDGSVWIASELKGLNDEC-EHFEVFP 174
Query: 108 TGCLFHSE 115
G L+ S+
Sbjct: 175 PGHLYSSK 182
>ASNB_ECOLI (P22106) Asparagine synthetase B [glutamine-hydrolyzing]
(EC 6.3.5.4)
Length = 553
Score = 42.7 bits (99), Expect = 4e-04
Identities = 30/119 (25%), Positives = 53/119 (44%), Gaps = 11/119 (9%)
Query: 2 GNLHNLSQLNKQYG---LSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYD 58
G ++N L +YG + G++ I+ Y+ ++GP + L L+G FAF +YD
Sbjct: 75 GEIYNHQALRAEYGDRYQFQTGSDCEVILALYQ---EKGP----EFLDDLQGMFAFALYD 127
Query: 59 HKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHG 117
+ G I LY G G + ++ ++ + C ++ FP G S+ G
Sbjct: 128 SEKDAYLIGRDHLGIIPLYMGYDEHGQLYVASEMKALVPVC-RTIKEFPAGSYLWSQDG 185
>ASNS_ASPOF (P31752) Asparagine synthetase [glutamine-hydrolyzing]
(EC 6.3.5.4) (AS)
Length = 589
Score = 42.4 bits (98), Expect = 5e-04
Identities = 25/70 (35%), Positives = 37/70 (52%), Gaps = 1/70 (1%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D ++ AA + G LY G DGSV +S ++ + C + F FP
Sbjct: 116 LDGMFSFVLLDTRNNCFVAARDAVGITPLYIGWGLDGSVWLSSEMKGLNDDC-EHFEVFP 174
Query: 108 TGCLFHSEHG 117
G L+ S G
Sbjct: 175 PGNLYSSRSG 184
>ASN2_YEAST (P49090) Asparagine synthetase [glutamine-hydrolyzing] 2
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase
2)
Length = 571
Score = 37.0 bits (84), Expect = 0.022
Identities = 24/72 (33%), Positives = 36/72 (49%), Gaps = 3/72 (4%)
Query: 46 KGLEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADG--SVVISENLELVKASCAKSF 103
K L+G FAF +YD K + AA G + LY G ++ +V + L+ + C S
Sbjct: 113 KYLDGMFAFCLYDSKKDRIVAARDPIGVVTLYMGRSSQSPETVYFASELKCLTDVC-DSI 171
Query: 104 APFPTGCLFHSE 115
FP G ++ SE
Sbjct: 172 ISFPPGHVYDSE 183
>ASNS_SCHPO (P78753) Probable asparagine synthetase
[glutamine-hydrolyzing] (EC 6.3.5.4)
(Glutamine-dependent asparagine synthetase)
Length = 556
Score = 36.6 bits (83), Expect = 0.029
Identities = 27/84 (32%), Positives = 40/84 (47%), Gaps = 7/84 (8%)
Query: 34 RDRGPYPADQVLKGLEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADG--SVVISEN 91
R+ GP A+ L+G F++V+YD V AA G LY G ++D + +
Sbjct: 107 REHGPACANM----LDGMFSWVLYDQDKDKVVAARDPIGITTLYQGFSSDSPDTAYFASE 162
Query: 92 LELVKASCAKSFAPFPTGCLFHSE 115
L+ + C K A FP G + SE
Sbjct: 163 LKALHPVCDKIIA-FPPGHYYDSE 185
>YMJ6_YEAST (Q04489) Hypothetical 59.5 kDa protein in VPS9-RAD10
intergenic region
Length = 525
Score = 35.4 bits (80), Expect = 0.064
Identities = 19/74 (25%), Positives = 38/74 (50%), Gaps = 2/74 (2%)
Query: 16 LSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKDGTVFAASGSDGHIG 75
+S+G N++++I + L++ G D V+K LEG +A+ IYD ++ G
Sbjct: 102 ISQGDNDSLYIASMLQNLKE-GMGVID-VIKSLEGEYAYTIYDVNSSKLYFGRDPIGRRS 159
Query: 76 LYWGIAADGSVVIS 89
L + + D + ++
Sbjct: 160 LSYSVTPDNELYVA 173
>ASN1_YEAST (P49089) Asparagine synthetase [glutamine-hydrolyzing] 1
(EC 6.3.5.4) (Glutamine-dependent asparagine synthetase
1)
Length = 571
Score = 32.7 bits (73), Expect = 0.41
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 3/72 (4%)
Query: 46 KGLEGSFAFVIYDHKDGTVFAASGSDGHIGLYWG--IAADGSVVISENLELVKASCAKSF 103
K L+G FA+ +YD K + AA G LY G A+ +V + L+ + C +
Sbjct: 113 KYLDGMFAWTLYDAKQDRIVAARDPIGITTLYMGRSSASPKTVYFASELKCLTDDC-DTI 171
Query: 104 APFPTGCLFHSE 115
FP G ++ S+
Sbjct: 172 TAFPPGHVYDSK 183
>ASN2_METJA (Q58456) Putative asparagine synthetase
[glutamine-hydrolyzing] 2 (EC 6.3.5.4)
Length = 515
Score = 29.6 bits (65), Expect = 3.5
Identities = 17/67 (25%), Positives = 33/67 (48%), Gaps = 8/67 (11%)
Query: 2 GNLHNLSQLNKQYGL-SKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
G ++N +L +++ L ++ G + I++ Y L +K G +AF I+D K
Sbjct: 93 GEIYNYLELKEKFNLETETGTDTEVILKLYNKL-------GFDCVKEFNGMWAFCIFDKK 145
Query: 61 DGTVFAA 67
G +F +
Sbjct: 146 KGLIFCS 152
>YL24_ANASP (Q8YV57) Hypothetical WD-repeat protein all2124
Length = 1683
Score = 28.9 bits (63), Expect = 6.0
Identities = 24/63 (38%), Positives = 37/63 (58%), Gaps = 6/63 (9%)
Query: 43 QVLKGLEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKS 102
+VL G G +A V + H DG++ A +G+DG+I L+ + DGS++ + L KA S
Sbjct: 1358 EVLAGNSGVYA-VSFLH-DGSIIATAGADGNIQLWH--SQDGSLL--KTLPGNKAIYGIS 1411
Query: 103 FAP 105
F P
Sbjct: 1412 FTP 1414
>TRPE_SULSO (Q06128) Anthranilate synthase component I (EC 4.1.3.27)
Length = 421
Score = 28.9 bits (63), Expect = 6.0
Identities = 27/98 (27%), Positives = 42/98 (42%), Gaps = 15/98 (15%)
Query: 15 GLSKGGNEAMFIIEAYR---TLRDRGPYPADQVLKGLEGSFAFVIYDHKDGTVF-----A 66
GL KGG +A R +RD P D +IYDH +G V+ +
Sbjct: 76 GLFKGGMIGYISYDAVRFWEKIRDLKPAAEDWPYAEFFTPDNIIIYDHNEGKVYVNADLS 135
Query: 67 ASGSDGHIGLYWGIAADGSV-------VISENLELVKA 97
+ G G IG + D S+ ++SE+LE +++
Sbjct: 136 SVGGCGDIGEFKVSFYDESLNKNSYERIVSESLEYIRS 173
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,059,096
Number of Sequences: 164201
Number of extensions: 837669
Number of successful extensions: 1532
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1516
Number of HSP's gapped (non-prelim): 21
length of query: 170
length of database: 59,974,054
effective HSP length: 102
effective length of query: 68
effective length of database: 43,225,552
effective search space: 2939337536
effective search space used: 2939337536
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC129091.3


