

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127021.6 + phase: 0 /pseudo
(408 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
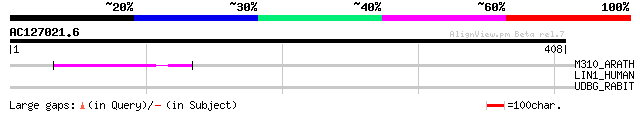
Score E
Sequences producing significant alignments: (bits) Value
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 51 5e-06
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 35 0.45
UDBG_RABIT (O19103) UDP-glucuronosyltransferase 2B16 precursor, ... 33 1.3
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 51.2 bits (121), Expect = 5e-06
Identities = 32/103 (31%), Positives = 48/103 (46%), Gaps = 9/103 (8%)
Query: 33 VYTFHIHRWPKIILNEITKIIRNFVWSGDCNQQKICTVAWNTMCRPK-DEGGLAIRDPFM 91
VY R K++ ++T + F WS N++KI VAW +C+ K D+GGL RD
Sbjct: 5 VYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRDLGW 64
Query: 92 VNNASLLHLTRKFLTSQDQWAQLCRHILLSNAKPKNHYIHSSI 134
N A L + + + H LLS ++ HSS+
Sbjct: 65 FNQALLAKQSFRIIHQP--------HTLLSRLLRSRYFPHSSM 99
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 34.7 bits (78), Expect = 0.45
Identities = 22/90 (24%), Positives = 39/90 (42%), Gaps = 7/90 (7%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHI--HRWPKIILNEITKIIRNFVW 58
+ IK + W S +GR +V I ++Y F+ + P E+ K F+W
Sbjct: 789 NEIKEDTNKWKNIPCSWVGRINIVKMAILPKVIYRFNAIPIKLPMTFFTELEKTTLKFIW 848
Query: 59 SGDCNQQKICTVAWNTMCRPKDEGGLAIRD 88
+ QK +A +T+ + GG+ + D
Sbjct: 849 N-----QKRAHIAKSTLSQKNKAGGITLPD 873
>UDBG_RABIT (O19103) UDP-glucuronosyltransferase 2B16 precursor,
microsomal (EC 2.4.1.17) (UDPGT) (Fragment)
Length = 523
Score = 33.1 bits (74), Expect = 1.3
Identities = 38/158 (24%), Positives = 58/158 (36%), Gaps = 31/158 (19%)
Query: 131 HSSIWIGLKSWVDVVIQHSNLEHW*WKICFILHSALSMSVSDYI--VDGEWNLPTYFHVK 188
H +GL + D QH NL H K I +MS SD++ + N P+Y K
Sbjct: 380 HGIPMVGLPLFAD---QHDNLAHMRAKGAAIRLDWKTMSSSDFLNALKTVINDPSY---K 433
Query: 189 DDVLTNKILQTTLPIKPVPDALQWLSTTDGHLTNKLTYLFVRGTCIKVPWHKLVWNTY-- 246
+ +T + P+KP+ A+ W+ H K ++V H L W Y
Sbjct: 434 EKAMTLSRIHHDQPMKPLDQAIFWIEFVMRHKGAK---------HLRVAAHDLTWFQYHS 484
Query: 247 ------------IPPTRVFICWRFIHNRLPTDENLRKR 272
I V CW ++ + +KR
Sbjct: 485 LDVIGFLLACLTITTYLVIKCWLLVYQNILMTGKKKKR 522
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.333 0.142 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,281,870
Number of Sequences: 164201
Number of extensions: 1961937
Number of successful extensions: 6234
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 6231
Number of HSP's gapped (non-prelim): 4
length of query: 408
length of database: 59,974,054
effective HSP length: 113
effective length of query: 295
effective length of database: 41,419,341
effective search space: 12218705595
effective search space used: 12218705595
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 67 (30.4 bits)
Medicago: description of AC127021.6


