

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127021.2 - phase: 0
(377 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
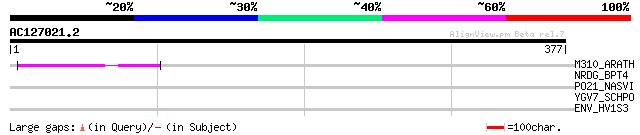
Score E
Sequences producing significant alignments: (bits) Value
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 44 9e-04
NRDG_BPT4 (P07075) Anaerobic ribonucleoside-triphosphate reducta... 35 0.41
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 33 1.2
YGV7_SCHPO (O43021) Oxysterol-binding protein homolog C354.07c 31 4.5
ENV_HV1S3 (P19549) Envelope polyprotein GP160 precursor [Contain... 30 7.7
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 43.5 bits (101), Expect = 9e-04
Identities = 30/98 (30%), Positives = 42/98 (42%), Gaps = 9/98 (9%)
Query: 6 GGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLR-EGGVTGSAWWRE 64
GGLG R L FN ALL K +R + L ++L SRY + G S WR
Sbjct: 55 GGLGFRDLGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRS 114
Query: 65 IVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWV 102
I + G + + +G+G +T W D W+
Sbjct: 115 I--------IHGRELLSRGLLRTIGDGIHTKVWLDRWI 144
>NRDG_BPT4 (P07075) Anaerobic ribonucleoside-triphosphate reductase
activating protein (EC 1.97.1.4) (Class III anaerobic
ribonucleotide reductase small component)
Length = 156
Score = 34.7 bits (78), Expect = 0.41
Identities = 25/82 (30%), Positives = 39/82 (47%), Gaps = 13/82 (15%)
Query: 156 EIRNLLTNVTLQDTQPDVWLWQPNIGDGYTVRGVYQMLMRQEVHNHDVVSDAPWHKSVPL 215
EI NL++ V + + D+WLW GY + Q+ M + V DV+ D + K++P
Sbjct: 82 EISNLVSWVKARFPEKDIWLW-----TGYKFEDIKQLEMLKYV---DVIIDGKYEKNLPT 133
Query: 216 KVSICAWRLLRNR--WPTKDNL 235
K WR N+ W D +
Sbjct: 134 KK---LWRGSDNQRLWSNTDGV 152
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 33.1 bits (74), Expect = 1.2
Identities = 20/56 (35%), Positives = 26/56 (45%), Gaps = 1/56 (1%)
Query: 210 HKSVPLKVSICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHC 265
H ++P V I N PT + RG TN LC GCG ET+ H++ C
Sbjct: 809 HVAIPASVWIKYHHTRINALPTLMRM-SRGRRTNGNALCRAGCGLPETLYHVVQQC 863
>YGV7_SCHPO (O43021) Oxysterol-binding protein homolog C354.07c
Length = 399
Score = 31.2 bits (69), Expect = 4.5
Identities = 17/48 (35%), Positives = 23/48 (47%), Gaps = 4/48 (8%)
Query: 135 EDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQPNIGD 182
E+ E W RR WEE + RN L L+ +P W++ IGD
Sbjct: 340 EEAEGAVWARRYFKWEEH-DSDARNALAQAVLEVIEPGFWIY---IGD 383
>ENV_HV1S3 (P19549) Envelope polyprotein GP160 precursor [Contains:
Exterior membrane glycoprotein (GP120); Transmembrane
glycoprotein (GP41)]
Length = 852
Score = 30.4 bits (67), Expect = 7.7
Identities = 26/80 (32%), Positives = 33/80 (40%), Gaps = 8/80 (10%)
Query: 75 EGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSIHKDLSVGEMHALGWG 134
EGG + S RL NGF FW D + + RL DL + V + GW
Sbjct: 731 EGGGERDRDRSTRLVNGFLALFWDDL---RSLCLFSYHRLTDLLLIVARIVELLGRRGW- 786
Query: 135 EDGEAWR-WRRRLLAWEEEL 153
E + W LL W +EL
Sbjct: 787 ---EVLKYWWNLLLYWSQEL 803
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.141 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,667,124
Number of Sequences: 164201
Number of extensions: 2112024
Number of successful extensions: 4888
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 4884
Number of HSP's gapped (non-prelim): 6
length of query: 377
length of database: 59,974,054
effective HSP length: 112
effective length of query: 265
effective length of database: 41,583,542
effective search space: 11019638630
effective search space used: 11019638630
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC127021.2


