

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
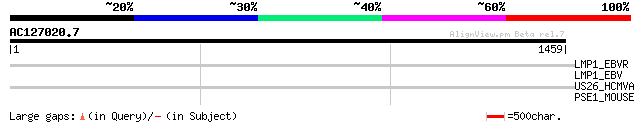
Query= AC127020.7 + phase: 0 /pseudo
(1459 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
LMP1_EBVR (P13198) Latent membrane protein 1 (LMP-1) (Protein p63) 35 1.5
LMP1_EBV (P03230) Latent membrane protein 1 (LMP-1) (Protein p63... 33 5.7
US26_HCMVA (P09699) Hypothetical protein HHLF5 33 7.5
PSE1_MOUSE (P97371) Proteasome activator complex subunit 1 (Prot... 33 7.5
>LMP1_EBVR (P13198) Latent membrane protein 1 (LMP-1) (Protein p63)
Length = 386
Score = 35.0 bits (79), Expect = 1.5
Identities = 42/133 (31%), Positives = 66/133 (49%), Gaps = 19/133 (14%)
Query: 488 LFMIVIFL*TAPGEAGVLLY-FGVIL*IVILLIF---QEITLLLGLLIVLRVL*DLLVIM 543
LF + I + G A ++LY F ++L I+IL+IF +++ LG L +L LL+I
Sbjct: 37 LFWLYIIMSNWTGGALLVLYAFALMLVIIILIIFIFRRDLLCPLGALCLL-----LLMIT 91
Query: 544 IILIAVVKMPRGIFYDNFHLNLQG----LGAFLAILMTSSIQVRKGGTPLDLLGLLTAFV 599
++LIA+ + Y L + G LG ++ +L I R G T LL AF
Sbjct: 92 LLLIALWNLHGQALYLGIVLFIFGCLLVLGLWIYLL---EILWRLGATIWQLLAFFLAF- 147
Query: 600 KLFLTLVYLMSRL 612
FL ++ L+ L
Sbjct: 148 --FLDIILLIIAL 158
>LMP1_EBV (P03230) Latent membrane protein 1 (LMP-1) (Protein p63)
[Contains: Protein p25]
Length = 386
Score = 33.1 bits (74), Expect = 5.7
Identities = 40/130 (30%), Positives = 63/130 (47%), Gaps = 13/130 (10%)
Query: 488 LFMIVIFL*TAPGEAGVLLY-FGVIL*IVILLIF---QEITLLLGLLIVLRVL*DLLVIM 543
LF + I + G A ++LY F ++L I+IL+IF +++ LG L +L LL+I
Sbjct: 37 LFWLYIVMSDWTGGALLVLYSFALMLIIIILIIFIFRRDLLCPLGALCIL-----LLMIT 91
Query: 544 IILIAVVKMPRGIFYDNFHLNLQGLGAFLAI-LMTSSIQVRKGGTPLDLLGLLTAFVKLF 602
++LIA+ + + L + G L I + + R G T LL AF F
Sbjct: 92 LLLIALWNLHGQALFLGIVLFIFGCLLVLGIWIYLLEMLWRLGATIWQLLAFFLAF---F 148
Query: 603 LTLVYLMSRL 612
L L+ L+ L
Sbjct: 149 LDLILLIIAL 158
>US26_HCMVA (P09699) Hypothetical protein HHLF5
Length = 603
Score = 32.7 bits (73), Expect = 7.5
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 5/65 (7%)
Query: 44 VLADREIQFAYFSERMSRAWKPGKRVTITKSVADRYLFQ----FHHKVDDARVLE-EGPW 98
+L DR +F YF W+ V + +V ++Q FH+ VDDA LE G
Sbjct: 292 ILVDRACEFFYFDVSRREIWRLADSVDMLLTVGLLKIYQAGRRFHYAVDDAERLEVPGRC 351
Query: 99 LYDNF 103
++NF
Sbjct: 352 PHENF 356
>PSE1_MOUSE (P97371) Proteasome activator complex subunit 1
(Proteasome activator 28-alpha subunit) (PA28alpha)
(PA28a) (Activator of multicatalytic protease subunit 1)
(11S regulator complex alpha subunit) (REG-alpha)
Length = 249
Score = 32.7 bits (73), Expect = 7.5
Identities = 20/66 (30%), Positives = 32/66 (48%), Gaps = 9/66 (13%)
Query: 236 PYLKPVTQRVGTAATNRWLQDPIPAVTPSHNSDVAAASVGRKTQTAGTGVVTNFHDRMTA 295
P +K VT+++ T WLQ IP + +N VA Q ++TN H ++
Sbjct: 119 PEIKDVTEQLNLVTT--WLQLQIPRIEDGNNFGVAV-------QEKVFELMTNLHTKLEG 169
Query: 296 FQTQLT 301
F TQ++
Sbjct: 170 FHTQIS 175
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.351 0.158 0.526
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 142,938,666
Number of Sequences: 164201
Number of extensions: 5487323
Number of successful extensions: 20780
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 20776
Number of HSP's gapped (non-prelim): 8
length of query: 1459
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1336
effective length of database: 39,777,331
effective search space: 53142514216
effective search space used: 53142514216
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 72 (32.3 bits)
Medicago: description of AC127020.7


