

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
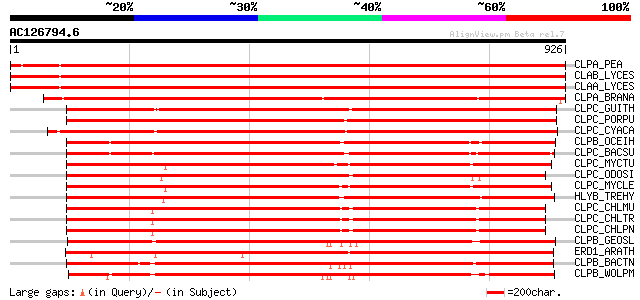
Query= AC126794.6 - phase: 0
(926 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CLPA_PEA (P35100) ATP-dependent Clp protease ATP-binding subunit... 1617 0.0
CLAB_LYCES (P31542) ATP-dependent Clp protease ATP-binding subun... 1564 0.0
CLAA_LYCES (P31541) ATP-dependent Clp protease ATP-binding subun... 1551 0.0
CLPA_BRANA (P46523) ATP-dependent Clp protease ATP-binding subun... 1447 0.0
CLPC_GUITH (O78410) ATP-dependent Clp protease ATP-binding subun... 1229 0.0
CLPC_PORPU (P51332) ATP-dependent Clp protease ATP-binding subun... 1206 0.0
CLPC_CYACA (Q9TM05) ATP-dependent Clp protease ATP-binding subun... 1187 0.0
CLPB_OCEIH (Q8EU05) Chaperone clpB 969 0.0
CLPC_BACSU (P37571) Negative regulator of genetic competence clp... 948 0.0
CLPC_MYCTU (O06286) Probable ATP-dependent Clp protease ATP-bind... 931 0.0
CLPC_ODOSI (P49574) ATP-dependent Clp protease ATP-binding subun... 925 0.0
CLPC_MYCLE (P24428) Probable ATP-dependent Clp protease ATP-bind... 925 0.0
HLYB_TREHY (Q54316) Hemolysin B 818 0.0
CLPC_CHLMU (Q9PKA8) Probable ATP-dependent Clp protease ATP-bind... 803 0.0
CLPC_CHLTR (O84288) Probable ATP-dependent Clp protease ATP-bind... 798 0.0
CLPC_CHLPN (Q9Z8A6) Probable ATP-dependent Clp protease ATP-bind... 793 0.0
CLPB_GEOSL (Q74FF1) Chaperone clpB 749 0.0
ERD1_ARATH (P42762) ERD1 protein, chloroplast precursor 748 0.0
CLPB_BACTN (Q89YY3) Chaperone clpB 723 0.0
CLPB_WOLPM (Q73IE4) Chaperone clpB 709 0.0
>CLPA_PEA (P35100) ATP-dependent Clp protease ATP-binding subunit
clpA homolog, chloroplast precursor
Length = 922
Score = 1617 bits (4187), Expect = 0.0
Identities = 835/926 (90%), Positives = 877/926 (94%), Gaps = 4/926 (0%)
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 60
M+R LAQS++VPGLVAG + + +KG+ +S+RSV+ M RT+ R+S +SGLRT N L+
Sbjct: 1 MARVLAQSLSVPGLVAGHKD-SQHKGSGKSKRSVKTMCALRTSGLRMSGFSGLRTFNHLN 59
Query: 61 SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHN 120
+M+RPG DFHSKV + + RA R R + +AMFERFTEKAIKVIMLAQEEARRLGHN
Sbjct: 60 TMMRPGLDFHSKVSKAVSSRRA---RAKRFIPRAMFERFTEKAIKVIMLAQEEARRLGHN 116
Query: 121 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 180
FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR
Sbjct: 117 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 176
Query: 181 VLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGEGA 240
VLELS EEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGE A
Sbjct: 177 VLELSQEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGESA 236
Query: 241 DSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP 300
DSV ATVGSGSSNNK PTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP
Sbjct: 237 DSVTATVGSGSSNNKTPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP 296
Query: 301 CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKL 360
CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKL
Sbjct: 297 CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKL 356
Query: 361 MEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHI 420
MEEIKQSD+IILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHI
Sbjct: 357 MEEIKQSDDIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHI 416
Query: 421 EKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISD 480
EKDP LERRFQPVKVPEPTV ETIQILKGLRERYEIHHKLRYTDEAL+AAA+LS+QYISD
Sbjct: 417 EKDPDLERRFQPVKVPEPTVDETIQILKGLRERYEIHHKLRYTDEALIAAAQLSYQYISD 476
Query: 481 RFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRD 540
RFLPDKAIDL+DEAGSRVRLQHAQLPEEA+ L+KEVR+IVKEK+E VRNQ+FEKAGELRD
Sbjct: 477 RFLPDKAIDLVDEAGSRVRLQHAQLPEEAKELDKEVRKIVKEKEEYVRNQDFEKAGELRD 536
Query: 541 KEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDESDR 600
KEMDLK QISALIEK KEM+KAE+E D G +VTEVDIQHIV+SWTGIPVDKVS DESDR
Sbjct: 537 KEMDLKAQISALIEKGKEMSKAETETADEGPIVTEVDIQHIVSSWTGIPVDKVSADESDR 596
Query: 601 LLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKAL 660
LLKMEDTLHKRIIGQ EAV+AISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKAL
Sbjct: 597 LLKMEDTLHKRIIGQDEAVQAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKAL 656
Query: 661 ASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDE 720
A+YYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDE
Sbjct: 657 AAYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDE 716
Query: 721 IEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDY 780
IEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDY
Sbjct: 717 IEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDY 776
Query: 781 DEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLK 840
DEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVF+RLK
Sbjct: 777 DEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFQRLK 836
Query: 841 TKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVD 900
TKEIEL VTERFR+RVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVD
Sbjct: 837 TKEIELQVTERFRDRVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVD 896
Query: 901 ADSDGNVIVLNGSTGAPDSLPDALPV 926
DSDG VIVLNGS+G P+SLP+AL +
Sbjct: 897 VDSDGKVIVLNGSSGTPESLPEALSI 922
>CLAB_LYCES (P31542) ATP-dependent Clp protease ATP-binding subunit
clpA homolog CD4B, chloroplast precursor
Length = 923
Score = 1564 bits (4049), Expect = 0.0
Identities = 813/927 (87%), Positives = 859/927 (91%), Gaps = 5/927 (0%)
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 60
M+RAL QS ++P VAG R N G+ +++R+V M+ +++S L ++GLR N++D
Sbjct: 1 MARALVQSTSIPSSVAGERTTKFN-GSGKTKRAVTMLCNAQSSSLTLRDFTGLRGCNAID 59
Query: 61 SMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHN 120
+++R G+ SKV R RG R V KAMFERFTEKAIKVIMLAQEEARRLGHN
Sbjct: 60 TLVRSGETLQSKVAAATYVRRP---RGCRFVPKAMFERFTEKAIKVIMLAQEEARRLGHN 116
Query: 121 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 180
FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR
Sbjct: 117 FVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKR 176
Query: 181 VLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGEGA 240
VLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADP+NIRTQVIRMVGE
Sbjct: 177 VLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPSNIRTQVIRMVGESN 236
Query: 241 DSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP 300
++VGA+VG G+S KMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP
Sbjct: 237 EAVGASVGGGTSGQKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNP 296
Query: 301 CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKL 360
CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKL
Sbjct: 297 CLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKL 356
Query: 361 MEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHI 420
MEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHI
Sbjct: 357 MEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHI 416
Query: 421 EKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISD 480
EKDPALERRFQPVKVPEPTV ETIQILKGLRERYEIHHKLRYTDE LVAAA+LS+QYISD
Sbjct: 417 EKDPALERRFQPVKVPEPTVDETIQILKGLRERYEIHHKLRYTDEDLVAAAQLSYQYISD 476
Query: 481 RFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRD 540
RFLPDKAIDLIDEAGSRVRL+HAQLPEEA+ LEKE+RQI KEK+EAVR Q+FEKAGELRD
Sbjct: 477 RFLPDKAIDLIDEAGSRVRLRHAQLPEEAKELEKELRQITKEKNEAVRGQDFEKAGELRD 536
Query: 541 KEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDESDR 600
+EMDLK QI+ALI+KNKE++KAESEA D G LVTE DIQHIV+SWTGIPV+KVS DESDR
Sbjct: 537 REMDLKAQITALIDKNKEVSKAESEAADTGPLVTEADIQHIVSSWTGIPVEKVSTDESDR 596
Query: 601 LLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKAL 660
LLKME+TLH RIIGQ EAV+AISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKAL
Sbjct: 597 LLKMEETLHTRIIGQDEAVKAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKAL 656
Query: 661 ASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDE 720
A+YYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDE
Sbjct: 657 AAYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDE 716
Query: 721 IEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDY 780
IEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLD
Sbjct: 717 IEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDL 776
Query: 781 DEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLK 840
DEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLK
Sbjct: 777 DEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLK 836
Query: 841 TKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVD 900
KEIEL VTERFR+RVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLA EIKEGDSVIVD
Sbjct: 837 VKEIELQVTERFRDRVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLANEIKEGDSVIVD 896
Query: 901 ADSDGNVIVLNGSTGAP-DSLPDALPV 926
DSDGNV VLNGS+G P D P+ +PV
Sbjct: 897 VDSDGNVTVLNGSSGTPSDPAPEPIPV 923
>CLAA_LYCES (P31541) ATP-dependent Clp protease ATP-binding subunit
clpA homolog CD4A, chloroplast precursor
Length = 926
Score = 1551 bits (4015), Expect = 0.0
Identities = 812/929 (87%), Positives = 859/929 (92%), Gaps = 7/929 (0%)
Query: 1 MSRALAQSINVPGLVAGRRHVNNNKGAARSRRSVRMMFTTRTASPRLSSYSGLRTLNSLD 60
M+RAL QS N+ VAG R N G+ + +R+VRM+ + S RL++++GLR N+LD
Sbjct: 2 MARALVQSTNILPSVAGERAGQFN-GSRKDQRTVRMLCNVKCCSSRLNNFAGLRGCNALD 60
Query: 61 SML-RPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGH 119
++L + G+ HSKV R RG R V KAMFERFTEKAIKVIMLAQEEARRLGH
Sbjct: 61 TLLVKSGETLHSKVAAATFVRRP---RGCRFVPKAMFERFTEKAIKVIMLAQEEARRLGH 117
Query: 120 NFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAK 179
NFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGF+AVEIPFTPRAK
Sbjct: 118 NFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFIAVEIPFTPRAK 177
Query: 180 RVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGEG 239
RVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGE
Sbjct: 178 RVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGES 237
Query: 240 ADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNN 299
+++VGA+VG G+S KMPTLEEYGTNLTKLAEEGKLDPVVGRQ QIERVTQILGRRTKNN
Sbjct: 238 SEAVGASVGGGTSGLKMPTLEEYGTNLTKLAEEGKLDPVVGRQAQIERVTQILGRRTKNN 297
Query: 300 PCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKK 359
PCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKK
Sbjct: 298 PCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKK 357
Query: 360 LMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKH 419
LMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKH
Sbjct: 358 LMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKH 417
Query: 420 IEKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYIS 479
IEKDPALERRFQPVKVPEP+V ETIQILKGLRERYEIHHKL YTDEA+ AAA+LSHQYIS
Sbjct: 418 IEKDPALERRFQPVKVPEPSVDETIQILKGLRERYEIHHKLHYTDEAIEAAAKLSHQYIS 477
Query: 480 DRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELR 539
DRFLPDKAIDLIDEAGSRVRL+HAQLPEEAR LEKE+RQI KEK+EAVR Q+FEKAGELR
Sbjct: 478 DRFLPDKAIDLIDEAGSRVRLRHAQLPEEARELEKELRQITKEKNEAVRGQDFEKAGELR 537
Query: 540 DKEMDLKTQISALIEKNKEMNKAESEAGD-VGALVTEVDIQHIVASWTGIPVDKVSVDES 598
D+EMDLK QISALI+KNKE +KAESEAGD G +VTE DIQHIV+SWTGIPV+KVS DES
Sbjct: 538 DREMDLKAQISALIDKNKEKSKAESEAGDAAGPIVTEADIQHIVSSWTGIPVEKVSTDES 597
Query: 599 DRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAK 658
DRLLKME+TLH R+IGQ EAV+AISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAK
Sbjct: 598 DRLLKMEETLHTRVIGQDEAVKAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAK 657
Query: 659 ALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLF 718
+LA+YYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLF
Sbjct: 658 SLATYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLF 717
Query: 719 DEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDL 778
DEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDL
Sbjct: 718 DEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDL 777
Query: 779 DYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFER 838
D+DEKDSSYNRIKSLVTEELKQYFRPEFLNRL EMIVFRQLTKLEVKEIADIMLKEVF R
Sbjct: 778 DFDEKDSSYNRIKSLVTEELKQYFRPEFLNRLSEMIVFRQLTKLEVKEIADIMLKEVFVR 837
Query: 839 LKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVI 898
LK KEIEL VTERFR+RVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLA EIKEGDSVI
Sbjct: 838 LKNKEIELQVTERFRDRVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAGEIKEGDSVI 897
Query: 899 VDADSDGNVIVLNGSTGAP-DSLPDALPV 926
VD DSDGNV VLNG++GAP DS P+ + V
Sbjct: 898 VDVDSDGNVTVLNGTSGAPSDSAPEPILV 926
>CLPA_BRANA (P46523) ATP-dependent Clp protease ATP-binding subunit
clpA homolog, chloroplast precursor (Fragment)
Length = 874
Score = 1447 bits (3745), Expect = 0.0
Identities = 758/874 (86%), Positives = 803/874 (91%), Gaps = 10/874 (1%)
Query: 57 NSLDSMLRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARR 116
N LD++ R Q F KV + + KG RG V KAMFERFTEKAIKVIMLAQEEARR
Sbjct: 6 NVLDTLGRSRQSFGGKVRQAMNVPKGKGSRG---VVKAMFERFTEKAIKVIMLAQEEARR 62
Query: 117 LGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTP 176
LGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTP
Sbjct: 63 LGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTP 122
Query: 177 RAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMV 236
RAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADP+NIRTQVIRMV
Sbjct: 123 RAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPSNIRTQVIRMV 182
Query: 237 GEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRT 296
GE + V A VG GS NKMPTLEEYGTNLTKLAEEGKLDPVVGR PQIERV QILGRRT
Sbjct: 183 GEN-NEVTANVGGGSGTNKMPTLEEYGTNLTKLAEEGKLDPVVGRHPQIERVVQILGRRT 241
Query: 297 KNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEER 356
KNNPCLIGEPGVGKTAIAEGLAQRIA+G V ET EGKKVITLDMGLL AGTKYRGEFEER
Sbjct: 242 KNNPCLIGEPGVGKTAIAEGLAQRIASGVVRETSEGKKVITLDMGLLAAGTKYRGEFEER 301
Query: 357 LKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEY 416
+KKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEY
Sbjct: 302 VKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEY 361
Query: 417 RKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQ 476
RKHIEKDPALERRFQPVKVPEPTV ETIQILKGLRERYEIHHKLRYTDE+LVAAA+LS+Q
Sbjct: 362 RKHIEKDPALERRFQPVKVPEPTVDETIQILKGLRERYEIHHKLRYTDESLVAAAQLSYQ 421
Query: 477 YISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAG 536
YISDRFLPD+AIDL+DEAGSRVRL+HAQ+PEEAR LEKE+RQI KE +EAVR Q+FEKAG
Sbjct: 422 YISDRFLPDRAIDLMDEAGSRVRLRHAQVPEEARELEKELRQITKE-NEAVRGQDFEKAG 480
Query: 537 ELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVD 596
LRD+E++L+ ++SA+ K KEM+KAESE GD G +VTE DIQHIV+SWTGI V+KVS D
Sbjct: 481 TLRDREIELRAEVSAIQAKGKEMSKAESETGDEGPMVTESDIQHIVSSWTGILVEKVSTD 540
Query: 597 ESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSEL 656
ESD LLKME+TLHKR+IGQ EAV+AISRAIRRARVGLKNPNRPIASFIF GPTGVGKSEL
Sbjct: 541 ESDLLLKMEETLHKRVIGQDEAVKAISRAIRRARVGLKNPNRPIASFIFFGPTGVGKSEL 600
Query: 657 AKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVV 716
AKALA+YYFG EEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTE QLTEAVRRRPYTVV
Sbjct: 601 AKALAAYYFGCEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEPPQLTEAVRRRPYTVV 660
Query: 717 LFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGF 776
LFDEIEKAHPDVFNMMLQILEDGRLT+SKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGF
Sbjct: 661 LFDEIEKAHPDVFNMMLQILEDGRLTNSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGF 720
Query: 777 DLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVF 836
DLDY EKDSSYNRIKSLVT+ELKQYFRPEFLNRLDEMI+FRQLTKLEVKEIADI+L+E+F
Sbjct: 721 DLDY-EKDSSYNRIKSLVTQELKQYFRPEFLNRLDEMILFRQLTKLEVKEIADILLQELF 779
Query: 837 ERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDS 896
ERLK KE+EL VTERF+ERVVDEGYNPSYGARPLRRAIMRLLEDSM EKMLAREIKEGDS
Sbjct: 780 ERLKKKEVELQVTERFKERVVDEGYNPSYGARPLRRAIMRLLEDSMEEKMLAREIKEGDS 839
Query: 897 VIVDADSDGNVIVLNGSTGAP----DSLPDALPV 926
VIVD DS+G V VLNG +G P + D+LPV
Sbjct: 840 VIVDVDSEGKVTVLNGGSGTPTTSLEEQEDSLPV 873
>CLPC_GUITH (O78410) ATP-dependent Clp protease ATP-binding subunit
clpA homolog
Length = 819
Score = 1229 bits (3179), Expect = 0.0
Identities = 630/818 (77%), Positives = 722/818 (88%), Gaps = 7/818 (0%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMG+NLKDAR
Sbjct: 1 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGVNLKDAR 60
Query: 155 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVA 214
VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIG+EHLLLGL+REGEGVA
Sbjct: 61 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGTEHLLLGLIREGEGVA 120
Query: 215 ARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGK 274
ARVLENL D T +RTQVIR++G+ A+ V AT +G + K PTLEE+G+NLT+ A EGK
Sbjct: 121 ARVLENLALDLTKVRTQVIRLLGDTAE-VSAT--NGQTKGKTPTLEEFGSNLTQKAAEGK 177
Query: 275 LDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKK 334
LDPV+GRQ +IERV QILGRRTKNNP LIGEPGVGKTAIAEGLAQRI N DVP+ +E K+
Sbjct: 178 LDPVIGRQKEIERVIQILGRRTKNNPILIGEPGVGKTAIAEGLAQRINNRDVPDILEDKR 237
Query: 335 VITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAAN 394
V+TLD+GLLVAGTKYRGEFEERLKK+++EI+ ++ +IL IDEVHTLIGAGAAEGAIDAAN
Sbjct: 238 VVTLDIGLLVAGTKYRGEFEERLKKIIDEIRVANNVILVIDEVHTLIGAGAAEGAIDAAN 297
Query: 395 ILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERY 454
ILKPALARGE+QCIGATTL+EYRKHIEKD ALERRFQPV V EP+V ETI+IL GLR+RY
Sbjct: 298 ILKPALARGEMQCIGATTLEEYRKHIEKDSALERRFQPVMVGEPSVEETIEILYGLRDRY 357
Query: 455 EIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEK 514
E HHKL +DEAL AAA+ + QYI+DRFLPDKAIDLIDEAGSRVRL ++QLP AR L+K
Sbjct: 358 EKHHKLVISDEALSAAAKFADQYIADRFLPDKAIDLIDEAGSRVRLMNSQLPPAARELDK 417
Query: 515 EVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVT 574
E+R+I+K+KDEAVR+Q+FE AG+LRD+EM++K QI+A+ K + E ++VT
Sbjct: 418 ELREILKQKDEAVRSQDFETAGQLRDREMEIKAQIAAIAHSKKGDEENTKEV----SVVT 473
Query: 575 EVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLK 634
E DI IVA+WTGIPV+K++ ES++LL+ME+TLH RIIGQ EAV A+S+AIRRARVGLK
Sbjct: 474 EEDIAQIVAAWTGIPVNKMTRSESEKLLQMEETLHGRIIGQDEAVVAVSKAIRRARVGLK 533
Query: 635 NPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPG 694
NPNRPIASFIFSGPTGVGK+EL KALASY+FGSEEAM+RLDMSE+MERHTVSKLIGSPPG
Sbjct: 534 NPNRPIASFIFSGPTGVGKTELTKALASYFFGSEEAMVRLDMSEYMERHTVSKLIGSPPG 593
Query: 695 YVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNT 754
YVGY EGGQLTE+VRRRPYTVVLFDEIEK HPDVFN++LQILEDGRLTDSKGRTVDFKNT
Sbjct: 594 YVGYNEGGQLTESVRRRPYTVVLFDEIEKGHPDVFNLLLQILEDGRLTDSKGRTVDFKNT 653
Query: 755 LLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMI 814
LLI+TSNVGS VIEKGG +GFDL D+ +S Y RIK+LV EELKQYFRPEFLNRLDE+I
Sbjct: 654 LLILTSNVGSKVIEKGGGGLGFDLSEDQTESQYGRIKALVNEELKQYFRPEFLNRLDEII 713
Query: 815 VFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAI 874
VFRQLTK EV EIA+IMLKEVF R+ K I+L VT RF+ +++EGYNP YGARPLRRA+
Sbjct: 714 VFRQLTKDEVGEIAEIMLKEVFTRISEKGIQLEVTARFKTHLINEGYNPIYGARPLRRAV 773
Query: 875 MRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLNG 912
MRLLED+++E+ LA +IKEGD+ +VD D DG V VL G
Sbjct: 774 MRLLEDTLSEEFLAEKIKEGDTAVVDVDDDGKVKVLLG 811
>CLPC_PORPU (P51332) ATP-dependent Clp protease ATP-binding subunit
clpA homolog
Length = 821
Score = 1206 bits (3119), Expect = 0.0
Identities = 612/818 (74%), Positives = 719/818 (87%), Gaps = 4/818 (0%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGL+GEGTGIAA+VLKSM +NLKDAR
Sbjct: 1 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLVGEGTGIAAQVLKSMNVNLKDAR 60
Query: 155 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVA 214
VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIG+EHLL+GL+REGEGVA
Sbjct: 61 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGTEHLLMGLVREGEGVA 120
Query: 215 ARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGK 274
ARVLENL D ++IR +VI+M+GE A++ + + + +K PTLEE+G+NLT++A EG
Sbjct: 121 ARVLENLAVDVSSIRAEVIQMLGENAEANVSGSNATQARSKTPTLEEFGSNLTQMAIEGG 180
Query: 275 LDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKK 334
LDPVVGRQ +IERV QILGRRTKNNP LIGEPGVGKTAIAEGLAQRIAN DVP +E K
Sbjct: 181 LDPVVGRQKEIERVIQILGRRTKNNPVLIGEPGVGKTAIAEGLAQRIANRDVPSILEDKL 240
Query: 335 VITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAAN 394
VITLD+GLLVAGTKYRGEFEERLK++M+EIK +D +IL IDEVHTLIGAGAAEGAIDAAN
Sbjct: 241 VITLDVGLLVAGTKYRGEFEERLKRIMDEIKSADNVILVIDEVHTLIGAGAAEGAIDAAN 300
Query: 395 ILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERY 454
+LKPALARGELQCIGATTL+EYRKHIEKDPALERRF PV V EP+V ETI+IL GLR+RY
Sbjct: 301 LLKPALARGELQCIGATTLEEYRKHIEKDPALERRFHPVVVGEPSVEETIEILFGLRDRY 360
Query: 455 EIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEK 514
E HH+L +D AL AAA+ ++QYISDRFLPDKAIDLIDEAGSRVRL ++QLP AR L+K
Sbjct: 361 EKHHQLTMSDGALAAAAKYANQYISDRFLPDKAIDLIDEAGSRVRLLNSQLPPAARELDK 420
Query: 515 EVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVT 574
E+R ++K KDEA+R Q++E A + R +EM++K QI+A+ + K E + +VT
Sbjct: 421 ELRAVLKTKDEAIRAQKYETAEQYRAREMEIKAQIAAIAQSKKN----EPDLNLEDPVVT 476
Query: 575 EVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLK 634
E DI IVA+WTGIPV K++ ES++L++ME+TLH RIIGQ EAV A+SRAIRRARVGLK
Sbjct: 477 EDDIAEIVAAWTGIPVTKLTKSESEKLMQMEETLHGRIIGQDEAVIAVSRAIRRARVGLK 536
Query: 635 NPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPG 694
NPNRPIASFIFSGPTGVGK+EL KALASY+FGSE +MIRLDMSE+MERHTVSKLIGSPPG
Sbjct: 537 NPNRPIASFIFSGPTGVGKTELTKALASYFFGSEASMIRLDMSEYMERHTVSKLIGSPPG 596
Query: 695 YVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNT 754
YVGY+EGG LTEAVR++PYTV+LFDEIEKAHPD+FN++LQILEDGRLTD+KGRT+DFKNT
Sbjct: 597 YVGYSEGGYLTEAVRKKPYTVILFDEIEKAHPDIFNLLLQILEDGRLTDAKGRTIDFKNT 656
Query: 755 LLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMI 814
LLIMTSN+GS VIEKGG +GF+L D+ +S Y R++SLV EELKQYFRPEFLNRLDE+I
Sbjct: 657 LLIMTSNIGSKVIEKGGGSLGFELSEDQTESQYTRVRSLVNEELKQYFRPEFLNRLDEII 716
Query: 815 VFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAI 874
VFRQLTK EV+EIA++ML EVF R+K ++I+L+VTERF+ER+V+EGYNPSYGARPLRRA+
Sbjct: 717 VFRQLTKDEVREIAELMLNEVFARIKQQDIQLNVTERFKERLVEEGYNPSYGARPLRRAV 776
Query: 875 MRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVLNG 912
MRLLEDS+AE++L+ +IK GDS +VD ++G V VL G
Sbjct: 777 MRLLEDSLAEEVLSGKIKAGDSPVVDVTNEGEVKVLLG 814
>CLPC_CYACA (Q9TM05) ATP-dependent Clp protease ATP-binding subunit
clpA homolog
Length = 854
Score = 1187 bits (3072), Expect = 0.0
Identities = 599/852 (70%), Positives = 730/852 (85%), Gaps = 8/852 (0%)
Query: 63 LRPGQDFHSKVLTQIGTNRAKGGRGSRCVTKAMFERFTEKAIKVIMLAQEEARRLGHNFV 122
++P F +LT + KG K MFERFTEKA+KVIMLAQEEARRLGHNFV
Sbjct: 3 IQPNTFFLKAMLTLLNI---KGNNMCWAHKKNMFERFTEKAVKVIMLAQEEARRLGHNFV 59
Query: 123 GTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKRVL 182
GTEQILLG++GEGTG+AAK LKSMGI LKDAR+EVEKIIGRGSGFVA+EIPFTPRAK++L
Sbjct: 60 GTEQILLGILGEGTGLAAKALKSMGITLKDARIEVEKIIGRGSGFVAIEIPFTPRAKKIL 119
Query: 183 ELSLEEARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGEGADS 242
EL++EE+R L HNY+G+EHLLLGL++EGEGVAARVLENLG D +R+ +IRM+GE ++
Sbjct: 120 ELAIEESRILTHNYVGTEHLLLGLIKEGEGVAARVLENLGVDLPKLRSNIIRMIGETSE- 178
Query: 243 VGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCL 302
+VG+ S +K+PTLEE+GTNLT++A EGKLDPVVGR +IERV QILGRRTKNNP L
Sbjct: 179 --VSVGATSGRSKVPTLEEFGTNLTQMAVEGKLDPVVGRAKEIERVVQILGRRTKNNPVL 236
Query: 303 IGEPGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLME 362
IGEPGVGKTAIAEGLAQRI N +VP+T+E KKVITLD+ LLVAGTKYRGEFEERLKK+M+
Sbjct: 237 IGEPGVGKTAIAEGLAQRIINNEVPDTLEDKKVITLDVSLLVAGTKYRGEFEERLKKIMD 296
Query: 363 EIKQSDEIILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHIEK 422
EI+ +D +IL IDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTL+EYRKHIEK
Sbjct: 297 EIRMADNVILVIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLEEYRKHIEK 356
Query: 423 DPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISDRF 482
D ALERRFQPV V EPTV ETI+IL+GLR+RYE HH+L+ +D A+VAAA+LS QYI+DRF
Sbjct: 357 DAALERRFQPVMVEEPTVEETIEILRGLRDRYEAHHRLKISDSAIVAAAKLSDQYIADRF 416
Query: 483 LPDKAIDLIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKE 542
LPDKAIDL+DEA SRVRL + +LP A L++E+R I K K+E +R+ +FE+A + R++E
Sbjct: 417 LPDKAIDLVDEASSRVRLMNYKLPPSAEYLDEELRHIQKIKNELIRSGDFEEASQFRERE 476
Query: 543 MDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDESDRLL 602
+++K Q++AL++ KE E E +V E DI +IV+SWTGIPV K++ ES++LL
Sbjct: 477 IEVKVQMAALMKAKKEA--IEEELALNPPIVNEDDIANIVSSWTGIPVSKLTKSESEKLL 534
Query: 603 KMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKALAS 662
ME+TLH RI+GQ+EAV A+S+AIRRARVGLKNPNRPIASFIFSGPTGVGK+EL KA+AS
Sbjct: 535 HMEETLHSRIVGQNEAVIAVSKAIRRARVGLKNPNRPIASFIFSGPTGVGKTELTKAMAS 594
Query: 663 YYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIE 722
Y+FGSEEAM+RLDMSE+MERHTVSKLIGSPPGYVGY EGGQLTEAVR+RPYTVVLFDEIE
Sbjct: 595 YFFGSEEAMVRLDMSEYMERHTVSKLIGSPPGYVGYNEGGQLTEAVRKRPYTVVLFDEIE 654
Query: 723 KAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDE 782
KAHPDVFN++LQILEDGRLTDSKGRT+DFKNTLLIMTSN+GS VIEK G +GF+L+ +
Sbjct: 655 KAHPDVFNLLLQILEDGRLTDSKGRTIDFKNTLLIMTSNIGSKVIEKKGGGLGFELEENI 714
Query: 783 KDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTK 842
++ Y+R+++LV EELKQYFRPEFLNR+DE+IVFRQLTK EV++IA IML+E+FER+K +
Sbjct: 715 EELQYSRMRNLVNEELKQYFRPEFLNRVDEIIVFRQLTKDEVRDIAHIMLREIFERVKQQ 774
Query: 843 EIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDAD 902
I L VTERF+ +++EGYNPSYGARPLRRA++RLLEDS+AE++L+ +IKEGD+ ++D D
Sbjct: 775 GISLQVTERFKNLLIEEGYNPSYGARPLRRALVRLLEDSLAEEVLSGKIKEGDNAMIDVD 834
Query: 903 SDGNVIVLNGST 914
+ V +L G++
Sbjct: 835 ENKQVKILLGNS 846
>CLPB_OCEIH (Q8EU05) Chaperone clpB
Length = 809
Score = 969 bits (2505), Expect = 0.0
Identities = 493/817 (60%), Positives = 636/817 (77%), Gaps = 11/817 (1%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MF RFTE+A KV+ L+QEEA RLGHN +GTE ILLGL+ EG GIAAK L+S+G+ + +
Sbjct: 2 MFGRFTERAQKVLALSQEEAVRLGHNNIGTEHILLGLVREGEGIAAKALQSLGLEVSKIQ 61
Query: 155 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVA 214
EVEK+IG G I +TPRAK+V+ELS +EAR+LGH+Y+G+EH+LLGL+REGEGVA
Sbjct: 62 EEVEKLIGVGKQ-PTQSIHYTPRAKKVVELSQDEARKLGHSYVGTEHILLGLIREGEGVA 120
Query: 215 ARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGS-SNNKMPTLEEYGTNLTKLAEEG 273
ARVL NLG R QV++++G G SG SN PTL+ +LT A+EG
Sbjct: 121 ARVLNNLGVSLNKARQQVLQLLGSNESQAGRQGRSGQQSNASTPTLDSLARDLTVSAKEG 180
Query: 274 KLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGK 333
K+DPV+GR +IERV Q+L RRTKNNP LIGEPGVGKTA+AEGLAQ+I + +VPET+ K
Sbjct: 181 KIDPVIGRSKEIERVIQVLSRRTKNNPVLIGEPGVGKTAVAEGLAQQIIDNEVPETLRDK 240
Query: 334 KVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAA 393
+V+TLDMG +VAGTKYRGEFE+RLKK+MEEI+Q+ IILFIDE+HTLIGAG AEGAIDA+
Sbjct: 241 RVMTLDMGTVVAGTKYRGEFEDRLKKVMEEIRQAGNIILFIDELHTLIGAGGAEGAIDAS 300
Query: 394 NILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRER 453
NILKP+LARGELQCIGATTLDEYRK+IEKD ALERRFQP++V EPT+ ETIQIL GLR+R
Sbjct: 301 NILKPSLARGELQCIGATTLDEYRKYIEKDAALERRFQPIQVDEPTLEETIQILNGLRDR 360
Query: 454 YEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLE 513
YE HH++ TDEA+ AAA LS +YI+DRFLPDKAIDLIDEAGS+VRL+ +P + LE
Sbjct: 361 YEAHHRVTITDEAIEAAASLSDRYITDRFLPDKAIDLIDEAGSKVRLRSYTVPPNLKELE 420
Query: 514 KEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALV 573
+++ ++ KEKD AV++QEFEKA LRD E + ++ N+ + + G + V
Sbjct: 421 QKLDEVRKEKDAAVQSQEFEKAASLRDSEQRFREELET------TKNQWKEKQGQTDSEV 474
Query: 574 TEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGL 633
T DI +V++WTG+PV K++ DE+DRLL ME LH R+IGQ EAV A+++AIRRAR GL
Sbjct: 475 TMEDIAAVVSTWTGVPVSKLTKDETDRLLNMEKILHDRVIGQSEAVNAVAKAIRRARAGL 534
Query: 634 KNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPP 693
K+P RPI SFIF GPTGVGK+ELA+ALA F E+AMIR+DMSE+MERH S+L+GSPP
Sbjct: 535 KDPKRPIGSFIFLGPTGVGKTELARALAEVMFADEDAMIRIDMSEYMERHATSRLVGSPP 594
Query: 694 GYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKN 753
GYVGY EGGQLTE VRR+PY+VVL DE+EKAHP+VFN++LQ+LEDGRLTDSKGR VDF+N
Sbjct: 595 GYVGYDEGGQLTEKVRRKPYSVVLLDEVEKAHPEVFNILLQVLEDGRLTDSKGRVVDFRN 654
Query: 754 TLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEM 813
T++IMTSNVG+S + K + +GF LD +EKD Y +KS V EELK+ FRPEFLNR+DE
Sbjct: 655 TVIIMTSNVGASEL-KRNKYVGFALDNEEKD--YKDMKSKVIEELKKAFRPEFLNRIDET 711
Query: 814 IVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRA 873
IVF L K +K+I +M++++ +RLK +++ LS+T++ E++ +EG++P YGARPLRR+
Sbjct: 712 IVFHSLEKEHMKDIVTLMVQQLQKRLKEQDLHLSLTDKAIEKIANEGFDPEYGARPLRRS 771
Query: 874 IMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVL 910
I + +ED ++E++L I++ V + ++ G IVL
Sbjct: 772 IQKNIEDLLSEELLRGAIEKEQQVKIGLNNKGEFIVL 808
>CLPC_BACSU (P37571) Negative regulator of genetic competence
clpC/mecB
Length = 810
Score = 948 bits (2451), Expect = 0.0
Identities = 479/815 (58%), Positives = 636/815 (77%), Gaps = 13/815 (1%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MF RFTE+A KV+ LAQEEA RLGHN +GTE ILLGL+ EG GIAAK L+++G+ + +
Sbjct: 2 MFGRFTERAQKVLALAQEEALRLGHNNIGTEHILLGLVREGEGIAAKALQALGLGSEKIQ 61
Query: 155 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVA 214
EVE +IGRG ++ I +TPRAK+V+ELS++EAR+LGH+Y+G+EH+LLGL+REGEGVA
Sbjct: 62 KEVESLIGRGQE-MSQTIHYTPRAKKVIELSMDEARKLGHSYVGTEHILLGLIREGEGVA 120
Query: 215 ARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGK 274
ARVL NLG R QV++++G ++ G++ +SN PTL+ +LT +A+E
Sbjct: 121 ARVLNNLGVSLNKARQQVLQLLG--SNETGSSAAGTNSNANTPTLDSLARDLTAIAKEDS 178
Query: 275 LDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKK 334
LDPV+GR +I+RV ++L RRTKNNP LIGEPGVGKTAIAEGLAQ+I N +VPE + K+
Sbjct: 179 LDPVIGRSKEIQRVIEVLSRRTKNNPVLIGEPGVGKTAIAEGLAQQIINNEVPEILRDKR 238
Query: 335 VITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEGAIDAAN 394
V+TLDMG +VAGTKYRGEFE+RLKK+M+EI+Q+ IILFIDE+HTLIGAG AEGAIDA+N
Sbjct: 239 VMTLDMGTVVAGTKYRGEFEDRLKKVMDEIRQAGNIILFIDELHTLIGAGGAEGAIDASN 298
Query: 395 ILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERY 454
ILKP+LARGELQCIGATTLDEYRK+IEKD ALERRFQP++V +P+V E+IQIL+GLR+RY
Sbjct: 299 ILKPSLARGELQCIGATTLDEYRKYIEKDAALERRFQPIQVDQPSVDESIQILQGLRDRY 358
Query: 455 EIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEK 514
E HH++ TD+A+ AA +LS +YISDRFLPDKAIDLIDEAGS+VRL+ P + LE+
Sbjct: 359 EAHHRVSITDDAIEAAVKLSDRYISDRFLPDKAIDLIDEAGSKVRLRSFTTPPNLKELEQ 418
Query: 515 EVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVT 574
++ ++ KEKD AV++QEFEKA LRD E L+ Q+ + KE + G + VT
Sbjct: 419 KLDEVRKEKDAAVQSQEFEKAASLRDTEQRLREQVEDTKKSWKE------KQGQENSEVT 472
Query: 575 EVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLK 634
DI +V+SWTG+PV K++ E+D+LL ME+ LH R+IGQ EAV A+++A+RRAR GLK
Sbjct: 473 VDDIAMVVSSWTGVPVSKIAQTETDKLLNMENILHSRVIGQDEAVVAVAKAVRRARAGLK 532
Query: 635 NPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPG 694
+P RPI SFIF GPTGVGK+ELA+ALA FG EE+MIR+DMSE+ME+H+ S+L+GSPPG
Sbjct: 533 DPKRPIGSFIFLGPTGVGKTELARALAESIFGDEESMIRIDMSEYMEKHSTSRLVGSPPG 592
Query: 695 YVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNT 754
YVGY EGGQLTE VRR+PY+VVL DEIEKAHPDVFN++LQ+LEDGRLTDSKGRTVDF+NT
Sbjct: 593 YVGYDEGGQLTEKVRRKPYSVVLLDEIEKAHPDVFNILLQVLEDGRLTDSKGRTVDFRNT 652
Query: 755 LLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMI 814
+LIMTSNVG+S + K + +GF++ ++ ++ +K V ELK+ FRPEF+NR+DE+I
Sbjct: 653 ILIMTSNVGASEL-KRNKYVGFNV--QDETQNHKDMKDKVMGELKRAFRPEFINRIDEII 709
Query: 815 VFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAI 874
VF L K + EI +M ++ +RLK +++ + +T+ + +V +EG + YGARPLRRAI
Sbjct: 710 VFHSLEKKHLTEIVSLMSDQLTKRLKEQDLSIELTDAAKAKVAEEGVDLEYGARPLRRAI 769
Query: 875 MRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIV 909
+ +ED ++E++L I +G +++D + DG +V
Sbjct: 770 QKHVEDRLSEELLRGNIHKGQHIVLDVE-DGEFVV 803
>CLPC_MYCTU (O06286) Probable ATP-dependent Clp protease ATP-binding
subunit
Length = 848
Score = 931 bits (2405), Expect = 0.0
Identities = 471/815 (57%), Positives = 628/815 (76%), Gaps = 15/815 (1%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MFERFT++A +V++LAQEEAR L HN++GTE ILLGLI EG G+AAK L+S+GI+L+ R
Sbjct: 1 MFERFTDRARRVVVLAQEEARMLNHNYIGTEHILLGLIHEGEGVAAKSLESLGISLEGVR 60
Query: 155 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVA 214
+VE+IIG+G + IPFTPRAK+VLELSL EA QLGHNYIG+EH+LLGL+REGEGVA
Sbjct: 61 SQVEEIIGQGQQAPSGHIPFTPRAKKVLELSLREALQLGHNYIGTEHILLGLIREGEGVA 120
Query: 215 ARVLENLGADPTNIRTQVIRMVG--EGADSVGATVGSGSSNNKMPT----LEEYGTNLTK 268
A+VL LGA+ T +R QVI+++ +G ++ A G + P+ L+++G NLT
Sbjct: 121 AQVLVKLGAELTRVRQQVIQLLSGYQGKEAAEAGTGGRGGESGSPSTSLVLDQFGRNLTA 180
Query: 269 LAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPE 328
A EGKLDPV+GR+ +IERV Q+L RRTKNNP LIGEPGVGKTA+ EGLAQ I +G+VPE
Sbjct: 181 AAMEGKLDPVIGREKEIERVMQVLSRRTKNNPVLIGEPGVGKTAVVEGLAQAIVHGEVPE 240
Query: 329 TIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEG 388
T++ K++ TLD+G LVAG++YRG+FEERLKK+++EI +IILFIDE+HTL+GAGAAEG
Sbjct: 241 TLKDKQLYTLDLGSLVAGSRYRGDFEERLKKVLKEINTRGDIILFIDELHTLVGAGAAEG 300
Query: 389 AIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILK 448
AIDAA+ILKP LARGELQ IGATTLDEYRK+IEKD ALERRFQPV+V EPTV TI+ILK
Sbjct: 301 AIDAASILKPKLARGELQTIGATTLDEYRKYIEKDAALERRFQPVQVGEPTVEHTIEILK 360
Query: 449 GLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEE 508
GLR+RYE HH++ TD A+VAAA L+ +YI+DRFLPDKAIDLIDEAG+R+R++ P +
Sbjct: 361 GLRDRYEAHHRVSITDAAMVAAATLADRYINDRFLPDKAIDLIDEAGARMRIRRMTAPPD 420
Query: 509 ARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGD 568
R ++++ + +EK+ A+ Q+FEKA LRD+E KT ++ E+ K+ + D
Sbjct: 421 LREFDEKIAEARREKESAIDAQDFEKAASLRDRE---KTLVAQRAEREKQWRSGDL---D 474
Query: 569 VGALVTEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRR 628
V A V + I ++ +WTGIPV K++ E+ RLL+ME+ LHKRIIGQ +AV+A+S+AIRR
Sbjct: 475 VVAEVDDEQIAEVLGNWTGIPVFKLTEAETTRLLRMEEELHKRIIGQEDAVKAVSKAIRR 534
Query: 629 ARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKL 688
R GLK+P RP SFIF+GP+GVGK+EL+KALA++ FG ++A+I++DM EF +R T S+L
Sbjct: 535 TRAGLKDPKRPSGSFIFAGPSGVGKTELSKALANFLFGDDDALIQIDMGEFHDRFTASRL 594
Query: 689 IGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRT 748
G+PPGYVGY EGGQLTE VRR+P++VVLFDEIEKAH +++N +LQ+LEDGRLTD +GRT
Sbjct: 595 FGAPPGYVGYEEGGQLTEKVRRKPFSVVLFDEIEKAHQEIYNSLLQVLEDGRLTDGQGRT 654
Query: 749 VDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLN 808
VDFKNT+LI TSN+G+S I K +G ++ Y R+K V +ELK++FRPEFLN
Sbjct: 655 VDFKNTVLIFTSNLGTSDISK---PVGLGFSKGGGENDYERMKQKVNDELKKHFRPEFLN 711
Query: 809 RLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGAR 868
R+D++IVF QLT+ E+ + D+M+ V +LK+K++ L +T+ + + G++P GAR
Sbjct: 712 RIDDIIVFHQLTREEIIRMVDLMISRVAGQLKSKDMALVLTDAAKALLAKRGFDPVLGAR 771
Query: 869 PLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADS 903
PLRR I R +ED ++EK+L E+ G V VD D+
Sbjct: 772 PLRRTIQREIEDQLSEKILFEEVGPGQVVTVDVDN 806
>CLPC_ODOSI (P49574) ATP-dependent Clp protease ATP-binding subunit
clpA homolog
Length = 885
Score = 925 bits (2391), Expect = 0.0
Identities = 464/831 (55%), Positives = 626/831 (74%), Gaps = 35/831 (4%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MFE+FTE AIKVIML+QEEARR+GHNFVGTEQ+LLG+IG+ GI A+ LK + LK AR
Sbjct: 1 MFEKFTEGAIKVIMLSQEEARRMGHNFVGTEQLLLGIIGQRHGIGARALKKQKVTLKKAR 60
Query: 155 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVA 214
E+E IGRG+GFVA EIPFTPRAKRVLE+++ E + LG N++G+EH+LL L+ E +GVA
Sbjct: 61 REIELYIGRGTGFVASEIPFTPRAKRVLEMAVHEGKDLGQNFVGTEHILLALISESDGVA 120
Query: 215 ARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSS--------NNKMPTLEEYGTNL 266
R L+ LG + +R ++ + E + + + + PTL+EY N+
Sbjct: 121 MRTLDKLGVNIPKLRNLILMYIEENQEEILRPLTQAEKFLLEREKKGSSTPTLDEYSENI 180
Query: 267 TKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDV 326
+K A +GKLDPV+GR +I V ++L RR KNNP LIGEPGVGKTA+AEGLAQ I
Sbjct: 181 SKEAVDGKLDPVIGRDKEIHEVIKVLARRRKNNPVLIGEPGVGKTAVAEGLAQLIIAEKA 240
Query: 327 PETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAA 386
P+ ++G ++ LD+G ++AGTKYRGEFEER+K+++EE++ IIL IDE+HTL+GAGAA
Sbjct: 241 PDFLDGNLLMALDLGSILAGTKYRGEFEERIKRIVEEVQNDSAIILVIDEIHTLVGAGAA 300
Query: 387 EGAIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQI 446
EGA+DAANILKPALARG+ +CIGATT+DEYRK+IE+DPALERRFQPV V EPTV TI+I
Sbjct: 301 EGAVDAANILKPALARGKFRCIGATTIDEYRKYIERDPALERRFQPVHVKEPTVGVTIEI 360
Query: 447 LKGLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLP 506
L GLR ++E HH L Y D+A+ AA L+ ++I+DRFLPDKAID++DEAGSRVRL++ +LP
Sbjct: 361 LLGLRSKFEEHHTLSYHDKAVEQAAILADKFIADRFLPDKAIDVLDEAGSRVRLENRRLP 420
Query: 507 EEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISAL---IEKNKEMNKAE 563
+ L KE++ +++K+E+++ +F+ A +L D EM+++T I + I N+ + A
Sbjct: 421 RGMKRLLKELQDTLRDKEESIKEHDFDIAKQLVDHEMEVRTHIRIMKQSILTNETLGLAR 480
Query: 564 SEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAIS 623
E V E D+ ++A WTGIPV+K+S ES RLL ME+TLH+R+IGQH A+ ++S
Sbjct: 481 KEID----TVLEGDVAEVIAGWTGIPVNKISDSESKRLLTMEETLHERLIGQHHAIVSVS 536
Query: 624 RAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERH 683
+AIRRARVGL+NP+RPIASFIF+GPTGVGK+EL KAL+ Y FG+E++MIRLDMSE+ME+H
Sbjct: 537 KAIRRARVGLRNPDRPIASFIFAGPTGVGKTELTKALSEYMFGNEDSMIRLDMSEYMEKH 596
Query: 684 TVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTD 743
TV+KLIGSPPGYVGY EGGQLTEAV+ +PY+VVL DE+EKAHPDVFN++LQIL+DGRLTD
Sbjct: 597 TVAKLIGSPPGYVGYNEGGQLTEAVQTKPYSVVLLDEVEKAHPDVFNLLLQILDDGRLTD 656
Query: 744 SKGRTVDFKNTLLIMTSNVGSSVIEKGG--------RRIGFDLDYDE-----------KD 784
SKGRT+DF+NT++IMT+N+G+ +IEK + F +D KD
Sbjct: 657 SKGRTIDFRNTMIIMTTNLGAKIIEKESGIKPKTKQDKPAFRIDESGCLGWEPTPEPIKD 716
Query: 785 SS-YNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKE 843
S+ + ++ LV EELK++FRPEFLNR+DE+IVF LTK ++ EI +M+K++ +RL+ KE
Sbjct: 717 SALFEKVTELVNEELKEFFRPEFLNRIDEIIVFNHLTKYDIWEICGLMVKQLQKRLEEKE 776
Query: 844 IELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEG 894
+ L V R + +EGY+P YGARPLRRA+MRLLED++A++ L++ + G
Sbjct: 777 LTLEVDVSVRNLLTEEGYDPVYGARPLRRAVMRLLEDTLAQQCLSKPLYPG 827
>CLPC_MYCLE (P24428) Probable ATP-dependent Clp protease ATP-binding
subunit
Length = 848
Score = 925 bits (2391), Expect = 0.0
Identities = 469/815 (57%), Positives = 623/815 (75%), Gaps = 15/815 (1%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MFERFT++A +V++LAQEEAR L HN++GTE ILLGLI EG G+AAK L S+GI+L+ R
Sbjct: 1 MFERFTDRARRVVVLAQEEARMLNHNYIGTEHILLGLIHEGEGVAAKSLDSLGISLEAVR 60
Query: 155 VEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVA 214
+VE IIG+G + IPFTPRAK+VLELSL EA QLGHNYIG+EH+LLGL+REGEGVA
Sbjct: 61 SQVEDIIGQGQQAPSGHIPFTPRAKKVLELSLREALQLGHNYIGTEHILLGLIREGEGVA 120
Query: 215 ARVLENLGADPTNIRTQVIRMVG--EGADSVGATVGSGSSNNKMPT----LEEYGTNLTK 268
A+VL LGA+ T +R QVI+++ +G ++ A G + P+ L+++G NLT
Sbjct: 121 AQVLVKLGAELTRVRQQVIQLLSGYQGKEAAEAGTGGRGGESGSPSTSLVLDQFGRNLTA 180
Query: 269 LAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPE 328
A E KLDPV+GR+ +IERV Q+L RRTKNNP LIGEPGVGKTA+ EGLAQ I +G+VPE
Sbjct: 181 AAMESKLDPVIGREKEIERVMQVLSRRTKNNPVLIGEPGVGKTAVVEGLAQAIVHGEVPE 240
Query: 329 TIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAEG 388
T++ K++ TLD+G LVAG++YRG+FEERLKK+++EI +IILFIDE+HTL+GAGAAEG
Sbjct: 241 TLKDKQLYTLDLGSLVAGSRYRGDFEERLKKVLKEINTRGDIILFIDELHTLVGAGAAEG 300
Query: 389 AIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILK 448
AIDAA+ILKP LARGELQ IGATTLDEYRK+IEKD ALERRFQPV+V EPTV TI+ILK
Sbjct: 301 AIDAASILKPKLARGELQTIGATTLDEYRKYIEKDAALERRFQPVQVGEPTVEHTIEILK 360
Query: 449 GLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEE 508
GLR+RYE HH++ TD A+VAAA L+ +YI+DRFLPDKAIDLIDEAG+R+R++ P +
Sbjct: 361 GLRDRYEAHHRVSITDSAMVAAATLADRYINDRFLPDKAIDLIDEAGARMRIRRMTAPPD 420
Query: 509 ARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAGD 568
R ++++ + +EK+ A+ Q+FEKA LRD+E L Q + E+ K+ + D
Sbjct: 421 LREFDEKIAEARREKESAIDAQDFEKAASLRDREKQLVAQRA---EREKQWRSGDL---D 474
Query: 569 VGALVTEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRR 628
V A V + I ++ +WTGIPV K++ E+ RLL+ME+ LHKRIIGQ +AV+A+S+AIRR
Sbjct: 475 VIAEVDDEQIAEVLGNWTGIPVFKLTEAETTRLLRMEEELHKRIIGQEDAVKAVSKAIRR 534
Query: 629 ARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKL 688
R GLK+P RP SFIF+GP+GVGK+EL+KALA++ FG ++A+I++DM EF +R T S+L
Sbjct: 535 TRAGLKDPKRPSGSFIFAGPSGVGKTELSKALANFLFGDDDALIQIDMGEFHDRFTASRL 594
Query: 689 IGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRT 748
G+PPGYVGY EGGQLTE VRR+P++VVLFDEIEKAH +++N +LQ+LEDGRLTD +GRT
Sbjct: 595 FGAPPGYVGYEEGGQLTEKVRRKPFSVVLFDEIEKAHQEIYNSLLQVLEDGRLTDGQGRT 654
Query: 749 VDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLN 808
VDFKNT+LI TSN+G+S I K +G ++ Y R+K V +ELK++FRPEFLN
Sbjct: 655 VDFKNTVLIFTSNLGTSDISK---PVGLGFTQGSGENDYERMKQKVNDELKKHFRPEFLN 711
Query: 809 RLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGAR 868
R+D++IVF QL++ E+ + D+M+ V +LK K++ L +T + + + G++P GAR
Sbjct: 712 RIDDIIVFHQLSRDEIIRMVDLMISRVANQLKVKDMTLELTNKAKALLAKRGFDPVLGAR 771
Query: 869 PLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADS 903
PLRR I R +ED ++EK+L E+ G V VD D+
Sbjct: 772 PLRRTIQREIEDQLSEKILFEEVGPGQVVTVDVDN 806
>HLYB_TREHY (Q54316) Hemolysin B
Length = 828
Score = 818 bits (2113), Expect = 0.0
Identities = 420/821 (51%), Positives = 599/821 (72%), Gaps = 17/821 (2%)
Query: 95 MFE-RFTEKAIKVIML-AQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKD 152
MF+ T KA KVI L AQEEA+RL H+ V E ILLGL+ E +A +VL + I+L
Sbjct: 1 MFQFHLTSKAKKVIELYAQEEAKRLNHDMVTPEHILLGLLYESEALATRVLMRLKIDLDR 60
Query: 153 ARVEVEKIIGRGSGF-VAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGE 211
++E+E + + S V +P PR ++++ S EEAR L HNYIG+EHLLLGLLRE
Sbjct: 61 LKLELESAMVKSSTTKVFGTLPTAPRVQKLISRSAEEARALSHNYIGTEHLLLGLLREES 120
Query: 212 GVAARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNN----KMPTLEEYGTNLT 267
G A VL ++G + T +R ++++M+G ++ + + +N K PTL+++ +LT
Sbjct: 121 GTAYNVLTSMGLELTILRQEILKMLGVAGSNISSMEQTSQEDNVKKVKTPTLDQFARDLT 180
Query: 268 KLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVP 327
K+A + LD V+GR+ ++ RV QIL RR KNNP L+GEPGVGKTAI EGLA++I DVP
Sbjct: 181 KMARDKALDRVIGRENEVMRVVQILSRRKKNNPILLGEPGVGKTAIVEGLAEKIVAADVP 240
Query: 328 ETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEIILFIDEVHTLIGAGAAE 387
+ + K+V+TLD+ +VAGTKYRGEFEER+K ++ EIK++ II+FIDE+HTLIGAG AE
Sbjct: 241 DILLKKRVLTLDLSSVVAGTKYRGEFEERIKNIVLEIKKASNIIIFIDELHTLIGAGGAE 300
Query: 388 GAIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQIL 447
GA+DAAN+LKPAL+RGE+QCIGATT++EY+K+IEKD AL RRFQP+ V EP++ +TI+IL
Sbjct: 301 GALDAANMLKPALSRGEIQCIGATTINEYKKYIEKDGALVRRFQPINVEEPSIEDTIEIL 360
Query: 448 KGLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPE 507
G++ +YE HHK++YTDEA+ AAA LS +YI +R LPDKAIDLIDEAGSR RL + P+
Sbjct: 361 NGIKGKYEEHHKVKYTDEAINAAAVLSKRYIFERHLPDKAIDLIDEAGSRARLLNMTRPQ 420
Query: 508 EARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISALIEKNKEMNKAESEAG 567
E + LEK++ ++ ++K V +Q FE A ++RD+ L+ ++S K+ K E
Sbjct: 421 EFKDLEKKIEELNQQKKRVVESQNFEDAAKIRDEITSLQEELS------KKEEKCREERE 474
Query: 568 DVGALVTEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIR 627
+ + E DI+H+++ T IP+ ++ ES RL+ ME+ LH++++GQ EA+ +IS+AIR
Sbjct: 475 KIETFIEEDDIRHVISEITNIPIKRLLNSESKRLIGMEEELHQKVVGQKEAISSISKAIR 534
Query: 628 RARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSK 687
R+R GLK RP+ SFIF GPTGVGK+ LAK L+ + FG +A+IR+DMSEFME+ VS+
Sbjct: 535 RSRAGLKTSKRPLGSFIFLGPTGVGKTALAKVLSEFMFGDSDALIRIDMSEFMEKFAVSR 594
Query: 688 LIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGR 747
LIG+PPGYVGY EGG LTE VRR+PY+++LFDEIEKAHPDV N++LQ+LE+G+LTD+ GR
Sbjct: 595 LIGAPPGYVGYEEGGGLTEKVRRKPYSLILFDEIEKAHPDVTNILLQVLEEGQLTDNFGR 654
Query: 748 TVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFL 807
VDF NT++I+TSN+G+ I KG +GF+ EKD+ N IK+ EELKQ F PEFL
Sbjct: 655 KVDFSNTIIIITSNLGARDIVKGS-SLGFNAVGSEKDA--NDIKNFALEELKQNFNPEFL 711
Query: 808 NRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGA 867
NR+D++IVF L+K ++K+I +IMLKE+ E +K + I ++++E + ++D+G++ YGA
Sbjct: 712 NRIDDIIVFHTLSKEDLKDIINIMLKELNEAIKERNIVINLSEEAKNYIIDKGFDKKYGA 771
Query: 868 RPLRRAIMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVI 908
R LRRAI + +ED ++ ++L I++GD++ VD +DG++I
Sbjct: 772 RSLRRAIQKEIEDYVSTEILFGNIEDGDTINVDR-NDGSLI 811
>CLPC_CHLMU (Q9PKA8) Probable ATP-dependent Clp protease ATP-binding
subunit
Length = 870
Score = 803 bits (2074), Expect = 0.0
Identities = 413/824 (50%), Positives = 581/824 (70%), Gaps = 35/824 (4%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MFE+FT +A +VI LA++EA+RL HN++GTE ILLGL+ G G+A VL+++G++ A+
Sbjct: 17 MFEKFTNRAKQVIKLAKKEAQRLNHNYLGTEHILLGLLKLGQGVAVNVLRTLGVDFDTAK 76
Query: 155 VEVEKIIGRGSGFVAVEIP-FTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGV 213
EVE++IG G P T R K+ E + EEA L HNY+G+EHLLLG+L + +GV
Sbjct: 77 NEVERLIGYGPEIQVYGDPALTGRVKKSFESANEEASILEHNYVGTEHLLLGILNQADGV 136
Query: 214 AARVLENLGADPTNIRTQVIRMV------------------------GEGADSVGATVGS 249
A +VLENL DP IR ++++ + + S+G
Sbjct: 137 ALQVLENLHIDPKEIRKEILKELETFNLQLPPSSSITPRNTNSSSSSSSKSSSLGGHSLG 196
Query: 250 GSSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVG 309
G K+ L+ YG +LT++ +E +LDPV+GR ++ER+ IL RR KNNP LIGE GVG
Sbjct: 197 GDKPEKLSALKAYGYDLTEMFKEARLDPVIGRSAEVERLILILCRRRKNNPVLIGEAGVG 256
Query: 310 KTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDE 369
KTAI EGLAQ+I +G+VPE + K++ITLD+ L++AGTKYRG+FEER+K +M+E+++
Sbjct: 257 KTAIVEGLAQKIVSGEVPEALRKKRLITLDLALMIAGTKYRGQFEERIKAVMDEVRKHGN 316
Query: 370 IILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERR 429
I+LFIDE+HT++GAGAAEGAIDA++ILKPALARGE+QCIGATTLDEYRKHIEKD ALERR
Sbjct: 317 ILLFIDELHTIVGAGAAEGAIDASHILKPALARGEIQCIGATTLDEYRKHIEKDAALERR 376
Query: 430 FQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAID 489
FQ + V P+V ET++IL+GL+++YE HH + TDEALVAAA+LS QY+ RFLPDKAID
Sbjct: 377 FQKIVVQPPSVDETVEILRGLKKKYEEHHNVFITDEALVAAAKLSDQYVHGRFLPDKAID 436
Query: 490 LIDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQI 549
L+DEAG+RVR+ P E LE E+ + K++A+ QE+EKA LRD+E L+ ++
Sbjct: 437 LLDEAGARVRVNTMGQPSELLRLEAEIESTKQAKEQAIGTQEYEKAASLRDEEKKLREKL 496
Query: 550 SALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLH 609
S + ++ E NK E + V E + +V+ TGIP +++ ES++LL +E+TL
Sbjct: 497 SNM-KQQWESNKEEHQVP-----VDEEAVAQVVSVQTGIPAARLTEAESEKLLMLENTLQ 550
Query: 610 KRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEE 669
K++IGQ +AV +I RAIRR+R G+K+PNRP+ SF+F GPTGVGK+ LA+ +A FG E+
Sbjct: 551 KKVIGQDQAVASICRAIRRSRTGIKDPNRPMGSFLFLGPTGVGKTLLAQQIAVEMFGGED 610
Query: 670 AMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVF 729
++I++DMSE+ME+ +K++GSPPGYVG+ EGG LTE VRRRPY VVLFDEIEKAHPD+
Sbjct: 611 SLIQVDMSEYMEKFAATKMMGSPPGYVGHEEGGHLTEQVRRRPYCVVLFDEIEKAHPDIM 670
Query: 730 NMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNR 789
++MLQILE GRLTDS GR +DF+NT++IMTSN+G+ +I K G IGF L Y
Sbjct: 671 DLMLQILEQGRLTDSFGRKIDFRNTIIIMTSNLGADLIRKTG-EIGFGL---RSHMDYGV 726
Query: 790 IKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVT 849
IK + +K++ +PEF+NRLDE ++F+ L K + EI + + ++ RL+ ++ L++
Sbjct: 727 IKEKIDAAVKKHLKPEFINRLDESVIFKPLEKEALSEIIHLEINKLGSRLQNHQMALNIP 786
Query: 850 ERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKE 893
+ +VD+G++P GARPLRR + + LED +AE +L ++
Sbjct: 787 DSVVSFLVDKGHSPEMGARPLRRVVEQYLEDPLAEMLLKESCRQ 830
>CLPC_CHLTR (O84288) Probable ATP-dependent Clp protease ATP-binding
subunit
Length = 854
Score = 798 bits (2062), Expect = 0.0
Identities = 408/823 (49%), Positives = 580/823 (69%), Gaps = 34/823 (4%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MFE+FT +A +VI LA++EA+RL HN++GTE ILLGL+ G G+A VL+++G++ A+
Sbjct: 1 MFEKFTNRAKQVIKLAKKEAQRLNHNYLGTEHILLGLLKLGQGVAVNVLRTLGVDFDTAK 60
Query: 155 VEVEKIIGRGSGFVAVEIP-FTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGV 213
EVE++IG G P T R K+ E + EEA L HNY+G+EHLLLG+L + +GV
Sbjct: 61 HEVERLIGYGPEIQVCGDPALTGRVKKSFESANEEAALLEHNYVGTEHLLLGILNQSDGV 120
Query: 214 AARVLENLGADPTNIRTQVIRMV-----------------------GEGADSVGATVGSG 250
A +VLENL DP IR ++++ + + + +G G
Sbjct: 121 ALQVLENLHVDPKEIRKEILKELETFNLQLPPSSSITPRNTNSSSSSKSSSPLGGHTLGG 180
Query: 251 SSNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGK 310
K+ L+ YG +LT++ +E +LDPV+GR ++ER+ IL RR KNNP L+GE GVGK
Sbjct: 181 DKPEKLSALKAYGYDLTEMFKESRLDPVIGRSAEVERLILILCRRRKNNPVLVGEAGVGK 240
Query: 311 TAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEI 370
TAI EGLAQ+I +G+VPE + K++ITLD+ L++AGTKYRG+FEER+K +M+E+++ I
Sbjct: 241 TAIVEGLAQKIVSGEVPEALRKKRLITLDLALMIAGTKYRGQFEERIKAVMDEVRKHGNI 300
Query: 371 ILFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRF 430
+LFIDE+HT++GAGAAEGAIDA++ILKPALARGE+QCIGATTLDEYRKHIEKD ALERRF
Sbjct: 301 LLFIDELHTIVGAGAAEGAIDASHILKPALARGEIQCIGATTLDEYRKHIEKDAALERRF 360
Query: 431 QPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDL 490
Q + V P+V ET++IL+GL+++YE HH + TDEALVAAA+LS QY+ RFLPDKAIDL
Sbjct: 361 QKIVVQPPSVDETVEILRGLKKKYEEHHNVFITDEALVAAAKLSDQYVHGRFLPDKAIDL 420
Query: 491 IDEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQIS 550
+DEAG+RVR+ P + LE E+ + + K++A+ QE+EKA LRD+E L+ ++
Sbjct: 421 LDEAGARVRVNTMGQPSDLVRLEAEIEKTKQAKEQAIGTQEYEKAASLRDEEKKLREKLG 480
Query: 551 ALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHK 610
+ ++ E NK E + V E + +V+ TGIP +++ ES++LL +E TL K
Sbjct: 481 NM-KQQWESNKEEHQVP-----VDEEAVAQVVSVQTGIPAARLTEAESEKLLTLETTLQK 534
Query: 611 RIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEA 670
++IGQ +AV +I RAIRR+R G+K+PNRP+ SF+F GPTGVGK+ LA+ +A FG E++
Sbjct: 535 KVIGQSQAVASICRAIRRSRTGIKDPNRPMGSFLFLGPTGVGKTLLAQQIAIEMFGGEDS 594
Query: 671 MIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFN 730
+I++DMSE+ME+ +K++GSPPGYVG+ EGG LTE VRRRPY VVLFDEIEKAHPD+ +
Sbjct: 595 LIQVDMSEYMEKFAATKMMGSPPGYVGHEEGGHLTEQVRRRPYCVVLFDEIEKAHPDIMD 654
Query: 731 MMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRI 790
+MLQILE GRLTDS GR +DF+NT++IMTSN+G+ +I K G IGF L Y I
Sbjct: 655 LMLQILEQGRLTDSFGRKIDFRNTIIIMTSNLGADLIRKSG-EIGFGL---RSHMDYAVI 710
Query: 791 KSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTE 850
K + +K++ +PEF+NRLDE ++F+ L K + EI + + ++ RL+ +++L++ +
Sbjct: 711 KEKIDAAVKKHLKPEFINRLDESVIFKPLEKEALSEIIHLEINKLGSRLQNYQMDLNIPD 770
Query: 851 RFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKE 893
+V +G++P GARPLRR + + LED +AE +L ++
Sbjct: 771 SVISFLVTKGHSPEMGARPLRRVVEQYLEDPLAEMLLKESCRQ 813
>CLPC_CHLPN (Q9Z8A6) Probable ATP-dependent Clp protease ATP-binding
subunit
Length = 845
Score = 793 bits (2048), Expect = 0.0
Identities = 412/822 (50%), Positives = 576/822 (69%), Gaps = 34/822 (4%)
Query: 95 MFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDAR 154
MFE+FT +A +VI LA++EA+RL HN++GTE ILLGL+ G G+A VL+++GI+ AR
Sbjct: 1 MFEKFTNRAKQVIKLAKKEAQRLNHNYLGTEHILLGLLKLGQGVAVNVLRNLGIDFDTAR 60
Query: 155 VEVEKIIGRGSGFVAVEIP-FTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGV 213
EVE++IG G P T R K+ E + EEA L HNY+G+EHLLLG+L + + V
Sbjct: 61 QEVERLIGYGPEIQVYGDPALTGRVKKSFESANEEASLLEHNYVGTEHLLLGILHQSDSV 120
Query: 214 AARVLENLGADPTNIRTQVIRMV----------------------GEGADSVGATVGSGS 251
A +VLENL DP +R ++++ + +G ++GS
Sbjct: 121 ALQVLENLHIDPREVRKEILKELETFNLQLPPSSSSSSSSSRSNPSSSKSPLGHSLGS-D 179
Query: 252 SNNKMPTLEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKT 311
N K+ L+ YG +LT++ E KLDPV+GR ++ER+ IL RR KNNP LIGE GVGKT
Sbjct: 180 KNEKLSALKAYGYDLTEMVRESKLDPVIGRSSEVERLILILCRRRKNNPVLIGEAGVGKT 239
Query: 312 AIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSDEII 371
AI EGLAQ+I +VP+ + K++ITLD+ L++AGTKYRG+FEER+K +M+E+++ I+
Sbjct: 240 AIVEGLAQKIILNEVPDALRKKRLITLDLALMIAGTKYRGQFEERIKAVMDEVRKHGNIL 299
Query: 372 LFIDEVHTLIGAGAAEGAIDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQ 431
LFIDE+HT++GAGAAEGAIDA+NILKPALARGE+QCIGATT+DEYRKHIEKD ALERRFQ
Sbjct: 300 LFIDELHTIVGAGAAEGAIDASNILKPALARGEIQCIGATTIDEYRKHIEKDAALERRFQ 359
Query: 432 PVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLI 491
+ V P+V ETI+IL+GL+++YE HH + T+EAL AAA LS QY+ RFLPDKAIDL+
Sbjct: 360 KIVVHPPSVDETIEILRGLKKKYEEHHNVFITEEALKAAATLSDQYVHGRFLPDKAIDLL 419
Query: 492 DEAGSRVRLQHAQLPEEARGLEKEVRQIVKEKDEAVRNQEFEKAGELRDKEMDLKTQISA 551
DEAG+RVR+ P + LE E+ K++A+ QE+EKA LRD+E L+ ++ +
Sbjct: 420 DEAGARVRVNTMGQPTDLMKLEAEIENTKLAKEQAIGTQEYEKAAGLRDEEKKLRERLQS 479
Query: 552 LIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKR 611
+ ++ E +K E + V E + +V+ TGIP +++ ES++LLK+EDTL ++
Sbjct: 480 M-KQEWENHKEEHQVP-----VDEEAVAQVVSLQTGIPSARLTEAESEKLLKLEDTLRRK 533
Query: 612 IIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAM 671
+IGQ++AV +I RAIRR+R G+K+PNRP SF+F GPTGVGKS LA+ +A FG E+A+
Sbjct: 534 VIGQNDAVTSICRAIRRSRTGIKDPNRPTGSFLFLGPTGVGKSLLAQQIAIEMFGGEDAL 593
Query: 672 IRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNM 731
I++DMSE+ME+ +K++GSPPGYVG+ EGG LTE VRRRPY VVLFDEIEKAHPD+ ++
Sbjct: 594 IQVDMSEYMEKFAATKMMGSPPGYVGHEEGGHLTEQVRRRPYCVVLFDEIEKAHPDIMDL 653
Query: 732 MLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIK 791
MLQILE GRLTDS GR VDF++ ++IMTSN+G+ +I K G IGF L + Y I+
Sbjct: 654 MLQILEQGRLTDSFGRKVDFRHAIIIMTSNLGADLIRKSG-EIGFGL---KSHMDYKVIQ 709
Query: 792 SLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTER 851
+ +K++ +PEF+NRLDE ++FR L K + EI + + ++ RLK ++ L++ +
Sbjct: 710 EKIEHAMKKHLKPEFINRLDESVIFRPLEKESLSEIIHLEINKLDSRLKNYQMALNIPDS 769
Query: 852 FRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKE 893
+V +G++P GARPLRR I + LED +AE +L ++
Sbjct: 770 VISFLVTKGHSPEMGARPLRRVIEQYLEDPLAELLLKESCRQ 811
>CLPB_GEOSL (Q74FF1) Chaperone clpB
Length = 865
Score = 749 bits (1935), Expect = 0.0
Identities = 409/877 (46%), Positives = 580/877 (65%), Gaps = 82/877 (9%)
Query: 97 ERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMG---INLKDA 153
E+ T K + + AQ+ A R G+ + E +L+ L+ + G+ A +++ +G L+ A
Sbjct: 5 EKMTIKTQEALAGAQQLAARQGNGSIEPEHLLVSLLEQEGGLIAPIIQKVGGAPAALRSA 64
Query: 154 RVEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGV 213
+ K + + SG A + +P R+L+ + EA + ++ +EHLLLG + +
Sbjct: 65 ADVLVKRLPQVSGATA-QAYLSPALNRILDAAQREADTMKDEFVSTEHLLLGFFADRQCA 123
Query: 214 AARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEG 273
AAR L + G N+ ++ + G G V + + L +Y +LT LA +G
Sbjct: 124 AARALLDAGVSRDNVLAALMEIRG------GERVTDQNPEDTYQALAKYARDLTDLARQG 177
Query: 274 KLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGK 333
KLDPV+GR +I RV Q+L RRTKNNP LIGEPGVGKTAI EGLAQRI +GDVPET++ K
Sbjct: 178 KLDPVIGRDDEIRRVLQVLSRRTKNNPVLIGEPGVGKTAIVEGLAQRIVSGDVPETLKDK 237
Query: 334 KVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSD-EIILFIDEVHTLIGAGAAEGAIDA 392
+++ LDMG L+AG KYRGEFEERLK ++ E+ +S+ ++ILFIDE+HTL+GAGAAEGA+DA
Sbjct: 238 RLVALDMGALIAGAKYRGEFEERLKAVIREVAKSEGKVILFIDELHTLVGAGAAEGAMDA 297
Query: 393 ANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRE 452
+N+LKPALARGEL CIGATTL+EYRK+IEKD ALERRFQ V EP+V +TI IL+GL+E
Sbjct: 298 SNMLKPALARGELHCIGATTLNEYRKYIEKDAALERRFQQVYTGEPSVEDTIAILRGLKE 357
Query: 453 RYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGL 512
+YE +H +R D A++AAA LS +YI+DRFLPDKAIDLIDEA SR+R++ +P E +
Sbjct: 358 KYENYHGIRIKDSAIIAAATLSDRYITDRFLPDKAIDLIDEAASRLRIEIDSMPTEIDEV 417
Query: 513 EKEVRQIVKEKDEAVRN--------------QEFE----KAGELR---DKEMDLKTQISA 551
E+ + Q+ EK +R E E K+ EL+ +E D+ ++S+
Sbjct: 418 ERRIIQLEIEKQALLRESQDPHSLERLKKLADELEELKAKSAELKGHWQREKDIIGRVSS 477
Query: 552 ----LIEKNKEMNKAESEA----------GDVGALVTEVD-------------------- 577
L EK +E KAE E G++ A+ E+
Sbjct: 478 LRQRLEEKREEAKKAEREGNLARTAEIRYGEIPAIEKEIADRSAELEDIRKEGKMLPEEV 537
Query: 578 ----IQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGL 633
+ IV+ WTGIPV ++ E+D+L+ MED L R++GQ EA+ ++ AIRRAR GL
Sbjct: 538 DGELVAEIVSRWTGIPVSRMMEGEADKLVHMEDRLITRVVGQDEALVLVANAIRRARSGL 597
Query: 634 KNPNRPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPP 693
+PNRPI SF+F GPTGVGK+E AKALA + F ++A++R+DMSE+ E+HTV++LIG+PP
Sbjct: 598 SDPNRPIGSFLFLGPTGVGKTETAKALAEFLFNDDQAIVRIDMSEYQEKHTVARLIGAPP 657
Query: 694 GYVGYTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKN 753
GYVGY EGGQLTEAVRRRPY++VLFDEIEKAHP+VFN++LQ+L+DGRLTD +GRTVDF+N
Sbjct: 658 GYVGYEEGGQLTEAVRRRPYSIVLFDEIEKAHPEVFNVLLQVLDDGRLTDGQGRTVDFRN 717
Query: 754 TLLIMTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEM 813
T++IMTSN+GS I++ G S Y R+K++VTE LK+ F+PEFLNR+DE+
Sbjct: 718 TVIIMTSNLGSQWIQQYG------------SSDYARMKAMVTETLKEGFKPEFLNRIDEI 765
Query: 814 IVFRQLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRA 873
+++ L ++K+I DI ++ + +RL + I L ++++ RE + EGY+P+YGARPL+R
Sbjct: 766 VIYHALPLEQIKKIVDIQVECLKQRLADRRIVLELSDKAREYLSREGYDPAYGARPLKRT 825
Query: 874 IMRLLEDSMAEKMLAREIKEGDSVIVDADSDGNVIVL 910
I R ++D +A +L + +EGD+V VD G +V+
Sbjct: 826 IQRKIQDPLALALLEGKFQEGDTVRVDLSVSGEELVI 862
>ERD1_ARATH (P42762) ERD1 protein, chloroplast precursor
Length = 945
Score = 748 bits (1930), Expect = 0.0
Identities = 413/849 (48%), Positives = 560/849 (65%), Gaps = 38/849 (4%)
Query: 94 AMFERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGE--------GTGIAA-KVLK 144
A+FERFTE+AI+ I+ +Q+EA+ LG + V T+ +LLGLI E G+GI K +
Sbjct: 78 AVFERFTERAIRAIIFSQKEAKSLGKDMVYTQHLLLGLIAEDRDPQGFLGSGITIDKARE 137
Query: 145 SMGINLKDARVEVEKIIGRGSGFV-AVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLL 203
++ +A + ++ + + + ++PF+ KRV E ++E +R + YI EH+
Sbjct: 138 AVWSIWDEANSDSKQEEASSTSYSKSTDMPFSISTKRVFEAAVEYSRTMDCQYIAPEHIA 197
Query: 204 LGLLREGEGVAARVLENLGADPTNIRTQVI-RMVGEGADS----------------VGAT 246
+GL +G A RVL+ LGA+ + + R+ GE A G
Sbjct: 198 VGLFTVDDGSAGRVLKRLGANMNLLTAAALTRLKGEIAKDGREPSSSSKGSFESPPSGRI 257
Query: 247 VGSGSSNNKMPT-LEEYGTNLTKLAEEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGE 305
GSG K LE++ +LT A EG +DPV+GR+ +++RV QIL RRTKNNP L+GE
Sbjct: 258 AGSGPGGKKAKNVLEQFCVDLTARASEGLIDPVIGREKEVQRVIQILCRRTKNNPILLGE 317
Query: 306 PGVGKTAIAEGLAQRIANGDVPETIEGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIK 365
GVGKTAIAEGLA IA P + K++++LD+GLL+AG K RGE E R+ L+ E+K
Sbjct: 318 AGVGKTAIAEGLAISIAEASAPGFLLTKRIMSLDIGLLMAGAKERGELEARVTALISEVK 377
Query: 366 QSDEIILFIDEVHTLIGAGAAE-----GAIDAANILKPALARGELQCIGATTLDEYRKHI 420
+S ++ILFIDEVHTLIG+G +D AN+LKP+L RGELQCI +TTLDE+R
Sbjct: 378 KSGKVILFIDEVHTLIGSGTVGRGNKGSGLDIANLLKPSLGRGELQCIASTTLDEFRSQF 437
Query: 421 EKDPALERRFQPVKVPEPTVPETIQILKGLRERYEIHHKLRYTDEALVAAAELSHQYISD 480
EKD AL RRFQPV + EP+ + ++IL GLRE+YE HH +YT EA+ AA LS +YI+D
Sbjct: 438 EKDKALARRFQPVLINEPSEEDAVKILLGLREKYEAHHNCKYTMEAIDAAVYLSSRYIAD 497
Query: 481 RFLPDKAIDLIDEAGSRVRLQHAQLPEEARG--LEKEVRQIVKEKDEAVRNQEFEKAGEL 538
RFLPDKAIDLIDEAGSR R++ + +E L K +E E +
Sbjct: 498 RFLPDKAIDLIDEAGSRARIEAFRKKKEDAICILSKPPNDYWQEIKTVQAMHEVVLSSRQ 557
Query: 539 RDKEMDLKTQISALIEKNKEMNKAESEAGDVGALVTEVDIQHIVASWTGIPVDKVSVDES 598
+ + D + S + + + A + D LV DI + + W+GIPV +++ DE
Sbjct: 558 KQDDGDAISDESGELVEESSLPPAAGD--DEPILVGPDDIAAVASVWSGIPVQQITADER 615
Query: 599 DRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRPIASFIFSGPTGVGKSELAK 658
L+ +ED L R++GQ EAV AISRA++R+RVGLK+P+RPIA+ +F GPTGVGK+EL K
Sbjct: 616 MLLMSLEDQLRGRVVGQDEAVAAISRAVKRSRVGLKDPDRPIAAMLFCGPTGVGKTELTK 675
Query: 659 ALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYTEGGQLTEAVRRRPYTVVLF 718
ALA+ YFGSEE+M+RLDMSE+MERHTVSKLIGSPPGYVG+ EGG LTEA+RRRP+TVVLF
Sbjct: 676 ALAANYFGSEESMLRLDMSEYMERHTVSKLIGSPPGYVGFEEGGMLTEAIRRRPFTVVLF 735
Query: 719 DEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTSNVGSSVIEKGGR-RIGFD 777
DEIEKAHPD+FN++LQ+ EDG LTDS+GR V FKN L+IMTSNVGS I KG IGF
Sbjct: 736 DEIEKAHPDIFNILLQLFEDGHLTDSQGRRVSFKNALIIMTSNVGSLAIAKGRHGSIGFI 795
Query: 778 LDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEIADIMLKEVFE 837
LD DE+ +SY +K+LV EELK YFRPE LNR+DE+++FRQL K ++ EI ++ML+++
Sbjct: 796 LDDDEEAASYTGMKALVVEELKNYFRPELLNRIDEIVIFRQLEKAQMMEILNLMLQDLKS 855
Query: 838 RLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLEDSMAEKMLAREIKEGDSV 897
RL + L V+E +E + +GY+P+YGARPLRR + ++ED ++E LA K GD+
Sbjct: 856 RLVALGVGLEVSEPVKELICKQGYDPAYGARPLRRTVTEIVEDPLSEAFLAGSFKPGDTA 915
Query: 898 IVDADSDGN 906
V D GN
Sbjct: 916 FVVLDDTGN 924
>CLPB_BACTN (Q89YY3) Chaperone clpB
Length = 862
Score = 723 bits (1867), Expect = 0.0
Identities = 397/868 (45%), Positives = 554/868 (63%), Gaps = 67/868 (7%)
Query: 96 FERFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARV 155
F FT K+ + + A + G + IL G++ G + + + +G+N + +
Sbjct: 3 FNNFTIKSQEAVQEAVNLVQSRGQQAIEPAHILHGVMKVGENVTNFIFQKLGMNGQQVAL 62
Query: 156 EVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREGEGVAA 215
V+K I E + + VL+ + + ++++G ++ EH+LL LL V+
Sbjct: 63 VVDKQIESLPKVSGGEPYLSRESNEVLQKATQYSKEMGDEFVSLEHILLALLTVKSTVST 122
Query: 216 RVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLAEEGKL 275
+L++ G +R + + G V S SS + +LE+Y NL + A GKL
Sbjct: 123 -ILKDAGMTEKELRNAINEL------RKGEKVTSQSSEDNYQSLEKYAINLNEAARSGKL 175
Query: 276 DPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETIEGKKV 335
DPV+GR +I RV QIL RRTKNNP LIGEPG GKTAI EGLA RI GDVPE ++ K+V
Sbjct: 176 DPVIGRDEEIRRVLQILSRRTKNNPILIGEPGTGKTAIVEGLAHRILRGDVPENLKNKQV 235
Query: 336 ITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSD-EIILFIDEVHTLIGAGAAEGAIDAAN 394
+LDMG LVAG KY+GEFEERLK ++ E+K+S+ +IILFIDE+HTL+GAG EGA+DAAN
Sbjct: 236 YSLDMGALVAGAKYKGEFEERLKSVVNEVKKSEGDIILFIDEIHTLVGAGKGEGAMDAAN 295
Query: 395 ILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKGLRERY 454
ILKPALARGEL+ IGATTLDEY+K+ EKD ALERRFQ V+V EP TI IL+GL+ERY
Sbjct: 296 ILKPALARGELRSIGATTLDEYQKYFEKDKALERRFQIVQVDEPDTLSTISILRGLKERY 355
Query: 455 EIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEARGLEK 514
E HH +R D+A++AA ELS +YI+DRFLPDKAIDL+DEA +++R++ +PEE + +
Sbjct: 356 ENHHHVRIKDDAIIAAVELSSRYITDRFLPDKAIDLMDEAAAKLRMEVDSVPEELDEISR 415
Query: 515 EVRQIVKEKDEAVRNQEFEK-------AGELRDKEMDLKTQ------ISALIEKNK---- 557
+++Q+ E++ R + K EL++ E K + + I++NK
Sbjct: 416 KIKQLEIEREAIKRENDKPKLEIIGKELAELKEVEKSFKAKWQSEKTLMDKIQQNKVEIE 475
Query: 558 ----EMNKAESEA-------------------------------GDVGALVTEV---DIQ 579
E KAE E GD + EV DI
Sbjct: 476 NLKFEAEKAEREGDYGKVAEIRYGKLQALDKEIEDTQEKLRGMQGDKAMIKEEVDAEDIA 535
Query: 580 HIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPNRP 639
+V+ WTGIPV K+ E D+LL +ED LH+R+IGQ EA+EA++ A+RR+R GL++P RP
Sbjct: 536 DVVSRWTGIPVSKMMQSEKDKLLHLEDELHQRVIGQDEAIEAVADAVRRSRAGLQDPKRP 595
Query: 640 IASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVGYT 699
I SFIF G TGVGK+ELAKALA + F E M R+DMSE+ E+H+VS+L+G+PPGYVGY
Sbjct: 596 IGSFIFLGTTGVGKTELAKALAEFLFDDESMMTRIDMSEYQEKHSVSRLVGAPPGYVGYD 655
Query: 700 EGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMT 759
EGGQLTEA+RR+PY+VVLFDEIEKAHPDVFN++LQ+L+DGRLTD+KGR V+FKNT++IMT
Sbjct: 656 EGGQLTEAIRRKPYSVVLFDEIEKAHPDVFNILLQVLDDGRLTDNKGRVVNFKNTIIIMT 715
Query: 760 SNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQL 819
SN+GSS I+ + L K+ K V LK+ RPEFLNR+DE I+F L
Sbjct: 716 SNMGSSYIQSQMEK----LSGSNKEEVIEETKKEVMNMLKKNIRPEFLNRIDETIMFLPL 771
Query: 820 TKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRLLE 879
T+ E+++I + +K V + L +EL +T+ + GY+P +GARP++RAI R L
Sbjct: 772 TETEIRQIVLLQIKGVQKMLAENGVELEMTDAALNFLSQVGYDPEFGARPVKRAIQRYLL 831
Query: 880 DSMAEKMLAREIKEGDSVIVDADSDGNV 907
+ +++K+L++E+ ++IVD++ DG V
Sbjct: 832 NDLSKKLLSQEVDRSKAIIVDSNGDGLV 859
>CLPB_WOLPM (Q73IE4) Chaperone clpB
Length = 853
Score = 709 bits (1831), Expect = 0.0
Identities = 390/872 (44%), Positives = 562/872 (63%), Gaps = 86/872 (9%)
Query: 98 RFTEKAIKVIMLAQEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEV 157
+FTEKA +I AQ +A GH E +L ++ + +G+ ++ + G N++ V
Sbjct: 5 KFTEKAKSLIQSAQMKALGAGHQIFMPEHLLKVMLEDESGLVQDLINACGGNVQSISDAV 64
Query: 158 EKII-------GRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREG 210
+ I G GSG + + +V E S+ A++ +++ E LL GL +
Sbjct: 65 DSAIKKLPVIEGPGSG----GLQLSREIAKVFEDSIGIAKRNKDSFVAVERLLQGLTAQK 120
Query: 211 EGVAARVLENLGADPTNIRTQVIRMVGEGADSVGATVGSGSSNNKMPTLEEYGTNLTKLA 270
+ ++L G P + + V M G++ S +S K+ ++Y ++T+LA
Sbjct: 121 DDTVGKILAENGVTPQKLNSVVAEM------RKGSSADSPNSEEKLNAAKKYTKDVTELA 174
Query: 271 EEGKLDPVVGRQPQIERVTQILGRRTKNNPCLIGEPGVGKTAIAEGLAQRIANGDVPETI 330
+GKLDPV+GR +I R Q+ RRTKNNP LIGEPGVGKTAI EGLA RI DVP +
Sbjct: 175 MQGKLDPVIGRDEEIRRTMQVSLRRTKNNPVLIGEPGVGKTAIVEGLANRIVANDVPLGL 234
Query: 331 EGKKVITLDMGLLVAGTKYRGEFEERLKKLMEEIKQSD-EIILFIDEVHTLIGAGAAEGA 389
KV+ LD+G L+AGTK+RGEFEERLK ++ E+ +++ ++ILFIDE+HTL+GAGA GA
Sbjct: 235 HDAKVLALDLGALIAGTKFRGEFEERLKAVINELSRAEGKVILFIDELHTLVGAGATSGA 294
Query: 390 IDAANILKPALARGELQCIGATTLDEYRKHIEKDPALERRFQPVKVPEPTVPETIQILKG 449
+DA+N+LKPALARGE++CIGATTLDEYR+HIEKDPAL RRFQPV + +PT +TI IL+G
Sbjct: 295 MDASNLLKPALARGEVRCIGATTLDEYRQHIEKDPALARRFQPVFISQPTETDTISILRG 354
Query: 450 LRERYEIHHKLRYTDEALVAAAELSHQYISDRFLPDKAIDLIDEAGSRVRLQHAQLPEEA 509
L+ERYE+HH +R TD A++AAA LS++YI+DRFLPDKAIDLIDEA SRVR++ PE
Sbjct: 355 LKERYEVHHGIRITDGAIIAAATLSNRYITDRFLPDKAIDLIDEAASRVRIEMDSKPEVI 414
Query: 510 RGLEKEVRQIV-------KEKDEAVR------NQEFE----KAGELRDKEMDLKTQISAL 552
LE++V Q+ KE DE + N+E E K +L K K +I+ +
Sbjct: 415 DELERKVIQLKIEAEALKKESDENSKQRLKKINEEIENLNSKFADLNSKWQMEKNKIARI 474
Query: 553 IEKNKEMNKAESE------AGDVGAL-----------------------------VTEVD 577
E ++++ A E +G++G VT D
Sbjct: 475 QETAEKLDNARKELELVQRSGNLGRAGELMYGVIPQLENELKNQEKVTDSFLKKEVTGDD 534
Query: 578 IQHIVASWTGIPVDKVSVDESDRLLKMEDTLHKRIIGQHEAVEAISRAIRRARVGLKNPN 637
I +IV+ WTGIPVD + E ++LL ME+ + +R+IGQ +A+EAIS A+RR+R G+++ N
Sbjct: 535 IANIVSKWTGIPVDNMMHSEKEKLLNMENEIGRRVIGQKDAIEAISNAVRRSRSGVQDTN 594
Query: 638 RPIASFIFSGPTGVGKSELAKALASYYFGSEEAMIRLDMSEFMERHTVSKLIGSPPGYVG 697
+P SF+F GPTGVGK+ELAKALA + F + A++R DMSE+ME+H+VSKLIG+PPGYVG
Sbjct: 595 KPFGSFLFLGPTGVGKTELAKALAEFLFDDQSALLRFDMSEYMEKHSVSKLIGAPPGYVG 654
Query: 698 YTEGGQLTEAVRRRPYTVVLFDEIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLI 757
Y +GG+LTEAVRRRPY V+LFDEIEKA+PD+FN++LQIL++GRLTDS G+ +DF+NT+LI
Sbjct: 655 YEQGGRLTEAVRRRPYQVILFDEIEKANPDIFNLLLQILDEGRLTDSHGKLIDFRNTILI 714
Query: 758 MTSNVGSSVIEKGGRRIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFR 817
+TSN+G+ ++ KG DS N + +V K FRPEFLNRLDE+I+F
Sbjct: 715 LTSNLGAEIMLKG-----------NVDSVRNEVMQIV----KSAFRPEFLNRLDEIIIFH 759
Query: 818 QLTKLEVKEIADIMLKEVFERLKTKEIELSVTERFRERVVDEGYNPSYGARPLRRAIMRL 877
LT+ ++ +I D+ + + L +++ +S+ + +E + GY+P YGARPL+R I
Sbjct: 760 SLTRDDIYKIIDVQFSYLQKTLAKRKLSISLLQEAKELIAQTGYDPEYGARPLKRVIQEC 819
Query: 878 LEDSMAEKMLAREIKEGDSVIVDADSDGNVIV 909
+++++A+ +L+ E+ E D +IV A D ++V
Sbjct: 820 IQNNLAKLVLSGEVVENDELIVYA-LDNEILV 850
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.136 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 104,680,081
Number of Sequences: 164201
Number of extensions: 4623135
Number of successful extensions: 22503
Number of sequences better than 10.0: 748
Number of HSP's better than 10.0 without gapping: 211
Number of HSP's successfully gapped in prelim test: 542
Number of HSP's that attempted gapping in prelim test: 20472
Number of HSP's gapped (non-prelim): 1704
length of query: 926
length of database: 59,974,054
effective HSP length: 120
effective length of query: 806
effective length of database: 40,269,934
effective search space: 32457566804
effective search space used: 32457566804
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 71 (32.0 bits)
Medicago: description of AC126794.6


