

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
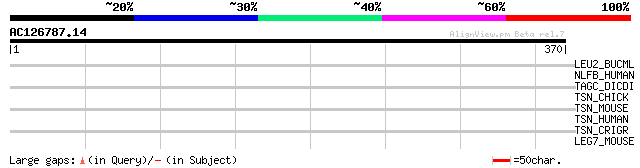
Query= AC126787.14 - phase: 0
(370 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
LEU2_BUCML (Q9EVG2) 3-isopropylmalate dehydratase large subunit ... 38 0.047
NLFB_HUMAN (Q8WX92) Negative elongation factor B (NELF-B) (Cofac... 33 0.88
TAGC_DICDI (Q23868) Prestalk-specific protein tagC precursor (EC... 33 1.5
TSN_CHICK (P79769) Translin 32 3.4
TSN_MOUSE (Q62348) Translin 31 5.7
TSN_HUMAN (Q15631) Translin 31 5.7
TSN_CRIGR (P97891) Translin 31 5.7
LEG7_MOUSE (O54974) Galectin-7 (Gal-7) 30 9.8
>LEU2_BUCML (Q9EVG2) 3-isopropylmalate dehydratase large subunit (EC
4.2.1.33) (Isopropylmalate isomerase) (Alpha-IPM
isomerase) (IPMI) (Fragment)
Length = 442
Score = 37.7 bits (86), Expect = 0.047
Identities = 30/115 (26%), Positives = 56/115 (48%), Gaps = 9/115 (7%)
Query: 58 DFMSSVKSSND---DVVFVEDRGLCVSLLHEIAWEVMPSKVFDPLLRNNSLNYDKFWSWS 114
DF ++KS D D +F D +L +I W P +V + NY+ F S +
Sbjct: 260 DFWQTLKSDKDAFFDKIFTIDIS---NLAPQITWGTNPDQVIS--IDEKIPNYETFSSLT 314
Query: 115 RLN-SVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVES 168
+ N + S C++ LK++ + ++ V I + ARI L L + +L ++++ +
Sbjct: 315 QRNLAKSACKYMGLKKESYLTNISIDKVFIGSXTNARIEDLRLAAKILKNKKISN 369
>NLFB_HUMAN (Q8WX92) Negative elongation factor B (NELF-B) (Cofactor
of BRCA1)
Length = 580
Score = 33.5 bits (75), Expect = 0.88
Identities = 32/113 (28%), Positives = 52/113 (45%), Gaps = 11/113 (9%)
Query: 132 VMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYHIDIDEMGLKL 191
++ L G+W MI DSQ +F + + L + + + SFL DY ++D+
Sbjct: 344 LLALGQGAWDMI--DSQ--VFKEPKMEVELITRFLPMLMSFLV---DDYTFNVDQKLPAE 396
Query: 192 DFIWAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNASDYGGLLGLL 244
+ YPN L + +F L + GL+++LHIT + LL LL
Sbjct: 397 EKAPVSYPNTLPESFTKF----LQEQRMACEVGLYYVLHITKQRNKNALLRLL 445
>TAGC_DICDI (Q23868) Prestalk-specific protein tagC precursor (EC
3.4.21.-)
Length = 1743
Score = 32.7 bits (73), Expect = 1.5
Identities = 23/81 (28%), Positives = 42/81 (51%), Gaps = 9/81 (11%)
Query: 25 WHLSRNKVLFFSGALFITLAVGVHLTPYFPSVSDFMSSVKSSNDDVVFVEDRGLCVSLLH 84
+H S NK+L SG + + L +G+ L + S S +K +N++ F D+ + LLH
Sbjct: 3 FHFSSNKLLLISGLILLVLVIGIKLDIFLSSKS--TEYIKFNNNNNHFKRDKRI---LLH 57
Query: 85 EIAWEVMPSKVFDPLLRNNSL 105
E++ + P ++NN +
Sbjct: 58 N---EIIDTN-NKPSIKNNQI 74
>TSN_CHICK (P79769) Translin
Length = 229
Score = 31.6 bits (70), Expect = 3.4
Identities = 18/59 (30%), Positives = 31/59 (52%), Gaps = 4/59 (6%)
Query: 289 LNTQEKRVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYD 347
L ++ R+ +N V G+Y R +R S + E F+LL++ N ++ DG+ YD
Sbjct: 147 LASELARLAVNSVTAGDYSRPLRISTFINELDSGFRLLNL----KNDSLRKRYDGLKYD 201
>TSN_MOUSE (Q62348) Translin
Length = 228
Score = 30.8 bits (68), Expect = 5.7
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 4/60 (6%)
Query: 288 MLNTQEKRVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYD 347
+L ++ R+ +N V G+Y R + S + E F+LL++ N ++ DG+ YD
Sbjct: 146 ILASELSRLSVNSVTAGDYSRPLHISTFINELDSGFRLLNL----KNDSLRKRYDGLKYD 201
>TSN_HUMAN (Q15631) Translin
Length = 228
Score = 30.8 bits (68), Expect = 5.7
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 4/60 (6%)
Query: 288 MLNTQEKRVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYD 347
+L ++ R+ +N V G+Y R + S + E F+LL++ N ++ DG+ YD
Sbjct: 146 ILASELSRLSVNSVTAGDYSRPLHISTFINELDSGFRLLNL----KNDSLRKRYDGLKYD 201
>TSN_CRIGR (P97891) Translin
Length = 228
Score = 30.8 bits (68), Expect = 5.7
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 4/60 (6%)
Query: 288 MLNTQEKRVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYD 347
+L ++ R+ +N V G+Y R + S + E F+LL++ N ++ DG+ YD
Sbjct: 146 ILASELSRLSVNSVTAGDYSRPLHISTFINELDSGFRLLNL----KNDSLRKRYDGLKYD 201
>LEG7_MOUSE (O54974) Galectin-7 (Gal-7)
Length = 135
Score = 30.0 bits (66), Expect = 9.8
Identities = 21/73 (28%), Positives = 37/73 (49%), Gaps = 2/73 (2%)
Query: 286 NSMLNTQEKRVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMH 345
N L+T E V N QG++ RE R +GI + G PF++L I + + D+ +H
Sbjct: 51 NPRLDTSE--VVFNTKQQGKWGREERGTGIPFQRGQPFEVLLIATEEGFKAVVGDDEYLH 108
Query: 346 YDGAVYEAELHIM 358
+ + A + ++
Sbjct: 109 FHHRLPPARVRLV 121
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,966,170
Number of Sequences: 164201
Number of extensions: 1861097
Number of successful extensions: 3794
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 3794
Number of HSP's gapped (non-prelim): 8
length of query: 370
length of database: 59,974,054
effective HSP length: 112
effective length of query: 258
effective length of database: 41,583,542
effective search space: 10728553836
effective search space used: 10728553836
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 66 (30.0 bits)
Medicago: description of AC126787.14


