

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126786.7 - phase: 0 /pseudo
(521 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
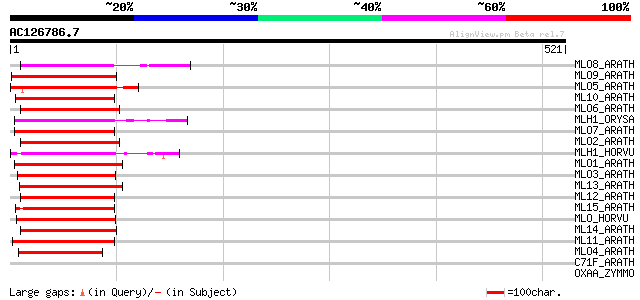
Score E
Sequences producing significant alignments: (bits) Value
MLO8_ARATH (O22757) MLO-like protein 8 (AtMlo8) 148 4e-35
MLO9_ARATH (Q94KB4) MLO-like protein 9 (AtMlo9) 140 1e-32
MLO5_ARATH (O22815) MLO-like protein 5 (AtMlo5) 135 3e-31
ML10_ARATH (Q9FKY5) MLO-like protein 10 (AtMlo10) 119 1e-26
MLO6_ARATH (Q94KB7) MLO-like protein 6 (AtMlo6) 111 4e-24
MLH1_ORYSA (O49914) MLO protein homolog 1 110 7e-24
MLO7_ARATH (O22752) MLO-like protein 7 (AtMlo7) 110 1e-23
MLO2_ARATH (Q9SXB6) MLO-like protein 2 (AtMlo2) 106 2e-22
MLH1_HORVU (O49873) MLO protein homolog 1 103 1e-21
MLO1_ARATH (O49621) MLO-like protein 1 (AtMlo1) (MLO protein hom... 103 1e-21
MLO3_ARATH (Q94KB9) MLO-like protein 3 (AtMlo3) 101 5e-21
ML13_ARATH (Q94KB2) MLO-like protein 13 (AtMlo13) (AtMlo20) 94 7e-19
ML12_ARATH (O80961) MLO-like protein 12 (AtMlo12) (AtMlo18) 94 7e-19
ML15_ARATH (O80580) MLO-like protein 15 (AtMlo15) 93 1e-18
MLO_HORVU (P93766) MLO protein 89 2e-17
ML14_ARATH (Q94KB1) MLO-like protein 14 (AtMlo14) 84 1e-15
ML11_ARATH (Q9FI00) MLO-like protein 11 (AtMlo11) 82 3e-15
MLO4_ARATH (O23693) MLO-like protein 4 (AtMlo4) 70 1e-11
C71F_ARATH (P58046) Cytochrome P450 71A15 (EC 1.14.-.-) 37 0.12
OXAA_ZYMMO (Q9RNL5) Inner membrane protein oxaA 34 1.1
>MLO8_ARATH (O22757) MLO-like protein 8 (AtMlo8)
Length = 593
Score = 148 bits (373), Expect = 4e-35
Identities = 81/159 (50%), Positives = 96/159 (59%), Gaps = 25/159 (15%)
Query: 11 RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGF 70
+QL+ TPTWAVAAVC +++S+LLEKL+HK V +R K ALL+ALEKIKAELMVLGF
Sbjct: 39 KQLNQTPTWAVAAVCTFFIVVSVLLEKLLHKVGKVLWDRHKTALLDALEKIKAELMVLGF 98
Query: 71 ISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTADHPGEPALEPQGTEHEPTPNIPAG 130
ISLLLTFGQ YI +CIP + TMLPC P PN+
Sbjct: 99 ISLLLTFGQTYILDICIPSHVARTMLPC------------------------PAPNLKK- 133
Query: 131 EGESKGEHHRRLLSYERRFLGGGGGGPGCKPWTGYISYI 169
E + GE HRRLLS+E RFL GG P GY+ I
Sbjct: 134 EDDDNGESHRRLLSFEHRFLSGGEASPTKCTKEGYVELI 172
>MLO9_ARATH (Q94KB4) MLO-like protein 9 (AtMlo9)
Length = 460
Score = 140 bits (352), Expect = 1e-32
Identities = 67/100 (67%), Positives = 80/100 (80%), Gaps = 1/100 (1%)
Query: 2 GGGGGGGDS-RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEK 60
GGGGGGG+ RQLD TPTWAV+ VC +I++ISI+LE +IHK VFE +KK AL EALEK
Sbjct: 4 GGGGGGGEGPRQLDQTPTWAVSTVCGVIILISIILELIIHKVGEVFERKKKKALFEALEK 63
Query: 61 IKAELMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPCQP 100
IK ELMVLGFISLLLTFGQNYI+ +C+P +Y + M C P
Sbjct: 64 IKNELMVLGFISLLLTFGQNYIASICVPSRYGHAMSFCGP 103
>MLO5_ARATH (O22815) MLO-like protein 5 (AtMlo5)
Length = 501
Score = 135 bits (340), Expect = 3e-31
Identities = 70/125 (56%), Positives = 87/125 (69%), Gaps = 9/125 (7%)
Query: 1 MGGGGGGGDS----RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLE 56
M GGGGG S R+LD TPTWAV+ VC +I++ISI+LE +IHK VF ER+K AL E
Sbjct: 1 MAGGGGGSTSGEGPRELDQTPTWAVSTVCGVIILISIVLELMIHKIGEVFTERRKKALYE 60
Query: 57 ALEKIKAELMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTADHPGEPALEP 116
AL+KIK ELMVLGFISLLLTFGQNYI+ +C+ +Y + M C P D P + +P
Sbjct: 61 ALQKIKNELMVLGFISLLLTFGQNYIASLCVASRYGHAMSFCGPY-----DGPSGESKKP 115
Query: 117 QGTEH 121
+ TEH
Sbjct: 116 KTTEH 120
>ML10_ARATH (Q9FKY5) MLO-like protein 10 (AtMlo10)
Length = 569
Score = 119 bits (299), Expect = 1e-26
Identities = 56/93 (60%), Positives = 72/93 (77%)
Query: 6 GGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAEL 65
GG + L TPTWAVA VC +++S+LLEK +H+ A+ E+ K++LLEALEKIKAEL
Sbjct: 29 GGAKEKGLSQTPTWAVALVCTFFILVSVLLEKALHRVATWLWEKHKNSLLEALEKIKAEL 88
Query: 66 MVLGFISLLLTFGQNYISKVCIPVKYSNTMLPC 98
M+LGFISLLLTFG+ YI K+CIP K + +MLPC
Sbjct: 89 MILGFISLLLTFGEQYILKICIPEKAAASMLPC 121
>MLO6_ARATH (Q94KB7) MLO-like protein 6 (AtMlo6)
Length = 583
Score = 111 bits (278), Expect = 4e-24
Identities = 53/93 (56%), Positives = 70/93 (74%)
Query: 11 RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGF 70
+ L+ T TWAVA VC ++++ISI++EKLIHK S F+++ K AL EALEK+KAELM++GF
Sbjct: 8 KTLEETSTWAVAVVCFVLLLISIVIEKLIHKIGSWFKKKNKKALYEALEKVKAELMLMGF 67
Query: 71 ISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAE 103
ISLLLT GQ YIS +CIP + +M PC E
Sbjct: 68 ISLLLTIGQGYISNICIPKNIAASMHPCSASEE 100
>MLH1_ORYSA (O49914) MLO protein homolog 1
Length = 540
Score = 110 bits (276), Expect = 7e-24
Identities = 66/163 (40%), Positives = 89/163 (54%), Gaps = 34/163 (20%)
Query: 5 GGGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAE 64
GG SR+L TPTWAVA VCA++V++S +E +H + F R+K A+ +AL+KIKAE
Sbjct: 3 GGRSGSRELPETPTWAVAVVCAVLVLVSAAMEHGLHNLSHWFRRRQKKAMGDALDKIKAE 62
Query: 65 LMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTADHPGEPALEPQGTEHEPT 124
LM+LGFISLLLT Q ISK+CIP +N +LPC+ G+ A+E +
Sbjct: 63 LMLLGFISLLLTVAQAPISKICIPKSAANILLPCK---------AGQDAIEEE------- 106
Query: 125 PNIPAGEGESKGEHHRRLLSYERRFLGGGGGGPGCKPWTGYIS 167
A G RR L G GGG C + G ++
Sbjct: 107 ----AASG--------------RRSLAGAGGGDYCSKFDGKVA 131
>MLO7_ARATH (O22752) MLO-like protein 7 (AtMlo7)
Length = 542
Score = 110 bits (274), Expect = 1e-23
Identities = 53/94 (56%), Positives = 71/94 (75%)
Query: 5 GGGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAE 64
G ++L TPTWAVA VC +++IS LLEK + + A+ ++ K++LLEALEKIKAE
Sbjct: 25 GAPSGGKELSQTPTWAVAVVCTFLILISHLLEKGLQRLANWLWKKHKNSLLEALEKIKAE 84
Query: 65 LMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPC 98
LM+LGFISLLLTFG+ YI K+C+P K + +MLPC
Sbjct: 85 LMILGFISLLLTFGEPYILKICVPRKAALSMLPC 118
>MLO2_ARATH (Q9SXB6) MLO-like protein 2 (AtMlo2)
Length = 573
Score = 106 bits (264), Expect = 2e-22
Identities = 53/93 (56%), Positives = 67/93 (71%)
Query: 11 RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGF 70
R L+ T TWAVA VC +++ ISI+LE IHK + F+++ K AL EALEK+KAELM+LGF
Sbjct: 8 RTLEETSTWAVAVVCFVLLFISIVLEHSIHKIGTWFKKKHKQALFEALEKVKAELMLLGF 67
Query: 71 ISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAE 103
ISLLLT GQ IS +CI K ++TM PC E
Sbjct: 68 ISLLLTIGQTPISNICISQKVASTMHPCSAAEE 100
>MLH1_HORVU (O49873) MLO protein homolog 1
Length = 544
Score = 103 bits (257), Expect = 1e-21
Identities = 66/165 (40%), Positives = 90/165 (54%), Gaps = 35/165 (21%)
Query: 1 MGGGGGGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEK 60
M G GG R+L TPTWAVA VCA+++++S+ +E +HK F + +K AL EALEK
Sbjct: 1 MAGPAGG---RELSDTPTWAVAVVCAVMILVSVAMEHALHKLGHWFHKWRKKALGEALEK 57
Query: 61 IKAELMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTADHPGEPALEPQGTE 120
+KAELM++GFISLLL Q+ +S++CI + MLPC+P + G
Sbjct: 58 MKAELMLVGFISLLLIVTQDPVSRICISKEAGEKMLPCKP-------YDG---------- 100
Query: 121 HEPTPNIPAGEGESKGEHHRRLL------SYERRFLGGGGGGPGC 159
AG G+ K ++HRRLL RRFL G C
Sbjct: 101 --------AGGGKGK-DNHRRLLWLQGESETHRRFLAAPAGVDVC 136
>MLO1_ARATH (O49621) MLO-like protein 1 (AtMlo1) (MLO protein
homolog 1) (AtMLO-H1)
Length = 526
Score = 103 bits (256), Expect = 1e-21
Identities = 51/102 (50%), Positives = 68/102 (66%)
Query: 5 GGGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAE 64
G GG+ L+ TPTW VA VC +IV IS+ +E+L+H F +V +++K+ L EAL+K+K E
Sbjct: 2 GHGGEGMSLEFTPTWVVAGVCTVIVAISLAVERLLHYFGTVLKKKKQKPLYEALQKVKEE 61
Query: 65 LMVLGFISLLLTFGQNYISKVCIPVKYSNTMLPCQPLAERTA 106
LM+LGFISLLLT Q ISK C+ MLPC + R A
Sbjct: 62 LMLLGFISLLLTVFQGLISKFCVKENVLMHMLPCSLDSRREA 103
>MLO3_ARATH (Q94KB9) MLO-like protein 3 (AtMlo3)
Length = 508
Score = 101 bits (251), Expect = 5e-21
Identities = 46/92 (50%), Positives = 68/92 (73%)
Query: 8 GDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMV 67
G R L TPTWA+A VC + +SI LE+LI+ ++ ++ +K +LLEA+EK+K+ LMV
Sbjct: 14 GAVRSLQETPTWALATVCFFFIAVSICLERLINLLSTRLKKNRKTSLLEAVEKLKSVLMV 73
Query: 68 LGFISLLLTFGQNYISKVCIPVKYSNTMLPCQ 99
LGF+SL+L + +SK+CIP+KY+N MLPC+
Sbjct: 74 LGFMSLMLNVTEGEVSKICIPIKYANRMLPCR 105
>ML13_ARATH (Q94KB2) MLO-like protein 13 (AtMlo13) (AtMlo20)
Length = 478
Score = 94.4 bits (233), Expect = 7e-19
Identities = 47/98 (47%), Positives = 64/98 (64%), Gaps = 1/98 (1%)
Query: 10 SRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLG 69
S L+ TPTW VA +C IIV++S+L E+ +H + R++ AL EAL+K+K ELM+LG
Sbjct: 6 SGSLEYTPTWVVAFICFIIVLLSLLAERGLHHLGKCLKRRQQDALFEALQKLKEELMLLG 65
Query: 70 FISLLLTFGQNYISKVCIPVKYSNTMLPC-QPLAERTA 106
FISL+LT Q I +C+P N M PC +PL E A
Sbjct: 66 FISLMLTVSQAAIRHICVPPALVNNMFPCKKPLEEHHA 103
>ML12_ARATH (O80961) MLO-like protein 12 (AtMlo12) (AtMlo18)
Length = 576
Score = 94.4 bits (233), Expect = 7e-19
Identities = 46/88 (52%), Positives = 62/88 (70%)
Query: 11 RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGF 70
R L+ TPTWAVA VC +++ ISI++E +H F+++ K AL EALEK+KAELM+LGF
Sbjct: 6 RSLEETPTWAVAVVCFVLLFISIMIEYFLHFIGHWFKKKHKKALSEALEKVKAELMLLGF 65
Query: 71 ISLLLTFGQNYISKVCIPVKYSNTMLPC 98
ISLLL Q +S++CIP + T PC
Sbjct: 66 ISLLLVVLQTPVSEICIPRNIAATWHPC 93
>ML15_ARATH (O80580) MLO-like protein 15 (AtMlo15)
Length = 496
Score = 93.2 bits (230), Expect = 1e-18
Identities = 47/93 (50%), Positives = 64/93 (68%), Gaps = 2/93 (2%)
Query: 6 GGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAEL 65
GGG + L+ TPTW VA VC++IV IS +E+LIH+ F+ + L AL+KIK EL
Sbjct: 3 GGGTT--LEYTPTWVVALVCSVIVSISFAVERLIHRAGKHFKNNDQKQLFGALQKIKEEL 60
Query: 66 MVLGFISLLLTFGQNYISKVCIPVKYSNTMLPC 98
M++GFISLLL+ GQ+ I+K+CI + S LPC
Sbjct: 61 MLVGFISLLLSVGQSKIAKICISKELSEKFLPC 93
>MLO_HORVU (P93766) MLO protein
Length = 533
Score = 89.4 bits (220), Expect = 2e-17
Identities = 47/94 (50%), Positives = 66/94 (70%), Gaps = 1/94 (1%)
Query: 7 GGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELM 66
G +R+L TP+WAVA V A +V++S+L+E +HK F+ R K AL EALEK+KAELM
Sbjct: 6 GVPARELPETPSWAVAVVFAAMVLVSVLMEHGLHKLGHWFQHRHKKALWEALEKMKAELM 65
Query: 67 VLGFISLLLTFGQN-YISKVCIPVKYSNTMLPCQ 99
++GFISLLL Q+ I+K+CI ++ M PC+
Sbjct: 66 LVGFISLLLIVTQDPIIAKICISEDAADVMWPCK 99
>ML14_ARATH (Q94KB1) MLO-like protein 14 (AtMlo14)
Length = 554
Score = 83.6 bits (205), Expect = 1e-15
Identities = 40/91 (43%), Positives = 61/91 (66%), Gaps = 1/91 (1%)
Query: 11 RQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGF 70
R L +TPTW+VA V I V +S+++E+ IH+ ++ ++ K+ L ALEK+K ELM+LGF
Sbjct: 10 RTLGLTPTWSVATVLTIFVFVSLIVERSIHRLSNWLQKTKRKPLFAALEKMKEELMLLGF 69
Query: 71 ISLLLTFGQNYISKVCIPVKYSN-TMLPCQP 100
ISLLLT + I+ +C+ + N +PC P
Sbjct: 70 ISLLLTATSSTIANICVSSSFHNDRFVPCTP 100
>ML11_ARATH (Q9FI00) MLO-like protein 11 (AtMlo11)
Length = 573
Score = 82.4 bits (202), Expect = 3e-15
Identities = 38/97 (39%), Positives = 64/97 (65%), Gaps = 1/97 (1%)
Query: 3 GGGGGGDSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIK 62
G + R L ++PTW+VA V + V++S+++E+ I++ ++ + K+ + ALEK+K
Sbjct: 8 GNEADSNERSLALSPTWSVAIVLTVFVVVSLIVERSIYRLSTWLRKTKRKPMFAALEKMK 67
Query: 63 AELMVLGFISLLLTFGQNYISKVCIPVK-YSNTMLPC 98
ELM+LGFISLLLT + I+ +C+P Y++ LPC
Sbjct: 68 EELMLLGFISLLLTATSSTIANICVPSSFYNDRFLPC 104
>MLO4_ARATH (O23693) MLO-like protein 4 (AtMlo4)
Length = 573
Score = 70.5 bits (171), Expect = 1e-11
Identities = 32/79 (40%), Positives = 55/79 (69%)
Query: 9 DSRQLDVTPTWAVAAVCAIIVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVL 68
+ R L TPT++VA+V ++V + L+E+ I++F ++ ++ AL +LEK+K ELM+L
Sbjct: 7 EGRSLAETPTYSVASVVTVLVFVCFLVERAIYRFGKWLKKTRRKALFTSLEKMKEELMLL 66
Query: 69 GFISLLLTFGQNYISKVCI 87
G ISLLL+ +IS++C+
Sbjct: 67 GLISLLLSQSARWISEICV 85
>C71F_ARATH (P58046) Cytochrome P450 71A15 (EC 1.14.-.-)
Length = 496
Score = 37.0 bits (84), Expect = 0.12
Identities = 19/66 (28%), Positives = 36/66 (53%), Gaps = 1/66 (1%)
Query: 28 IVIISILLEKLIHKFASVFEERKKHALLEALEKIKAELMVLGFISLLLTFGQNYISKVCI 87
+ ++ +L +K++ FA V EE + ++E +EK ++ L LL+T + S+V
Sbjct: 130 VCVVHLLNKKMVQSFAKVREEERS-VMMEKVEKASSDSSPLNLSKLLITLTSDVASRVSF 188
Query: 88 PVKYSN 93
K+SN
Sbjct: 189 GKKHSN 194
>OXAA_ZYMMO (Q9RNL5) Inner membrane protein oxaA
Length = 579
Score = 33.9 bits (76), Expect = 1.1
Identities = 19/65 (29%), Positives = 35/65 (53%), Gaps = 1/65 (1%)
Query: 65 LMVLGFISLLLTFGQNYISKVCIPVKYSNTML-PCQPLAERTADHPGEPALEPQGTEHEP 123
L+++ +S L+ FG ++++K + + T QP AE TA+ G+ L+P +
Sbjct: 10 LVIVTILSALILFGWSFVTKHWLSTPPAPTQQGKNQPKAELTAEESGDKPLKPVDQVLQE 69
Query: 124 TPNIP 128
TP +P
Sbjct: 70 TPRLP 74
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.354 0.158 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,954,428
Number of Sequences: 164201
Number of extensions: 2119677
Number of successful extensions: 16307
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 16268
Number of HSP's gapped (non-prelim): 38
length of query: 521
length of database: 59,974,054
effective HSP length: 115
effective length of query: 406
effective length of database: 41,090,939
effective search space: 16682921234
effective search space used: 16682921234
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC126786.7


