

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126786.5 - phase: 0
(487 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
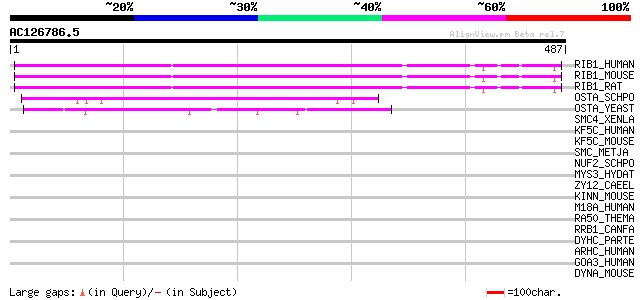
Score E
Sequences producing significant alignments: (bits) Value
RIB1_HUMAN (P04843) Dolichyl-diphosphooligosaccharide--protein g... 316 8e-86
RIB1_MOUSE (Q91YQ5) Dolichyl-diphosphooligosaccharide--protein g... 315 2e-85
RIB1_RAT (P07153) Dolichyl-diphosphooligosaccharide--protein gly... 308 2e-83
OSTA_SCHPO (Q10176) Putative dolichyl-diphosphooligosaccharide--... 122 2e-27
OSTA_YEAST (P41543) Dolichyl-diphosphooligosaccharide--protein g... 116 1e-25
SMC4_XENLA (P50532) Structural maintenance of chromosome 4 (Chro... 42 0.005
KF5C_HUMAN (O60282) Kinesin heavy chain isoform 5C (Kinesin heav... 40 0.010
KF5C_MOUSE (P28738) Kinesin heavy chain isoform 5C (Kinesin heav... 40 0.014
SMC_METJA (Q59037) Chromosome partition protein smc homolog 36 0.26
NUF2_SCHPO (Q10173) Myosin-like protein NUF2 (Nuclear filament-c... 35 0.33
MYS3_HYDAT (P39922) Myosin heavy chain, clone 203 (Fragment) 35 0.33
ZY12_CAEEL (Q23529) Zygote defective protein 12 35 0.44
KINN_MOUSE (P33175) Neuronal kinesin heavy chain (NKHC) (Kinesin... 35 0.44
M18A_HUMAN (Q92614) Myosin XVIIIA (Myosin 18A) (Myosin containin... 35 0.57
RA50_THEMA (Q9X1X1) Probable DNA double-strand break repair rad5... 34 0.97
RRB1_CANFA (Q28298) Ribosome-binding protein 1 (180 kDa ribosome... 33 1.3
DYHC_PARTE (Q27171) Dynein heavy chain, cytosolic (DYHC) 33 1.3
ARHC_HUMAN (Q9NZN5) Rho guanine nucleotide exchange factor 12 (L... 33 1.3
GOA3_HUMAN (Q08378) Golgi autoantigen, golgin subfamily A member... 33 1.7
DYNA_MOUSE (O08788) Dynactin 1 (150 kDa dynein-associated polype... 33 1.7
>RIB1_HUMAN (P04843) Dolichyl-diphosphooligosaccharide--protein
glycosyltransferase 67 kDa subunit precursor (EC
2.4.1.119) (Ribophorin I) (RPN-I)
Length = 607
Score = 316 bits (810), Expect = 8e-86
Identities = 176/488 (36%), Positives = 283/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 133 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPT 192
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP+ ++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 193 RSEDLLDYGPFRDVPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 252
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 253 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 311
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 312 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 371
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 372 IILPEGAKNIEIDSPYEISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 431
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 432 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 487
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ + + + ++ ++ DI + +KS+++ K LT E + LQS + +
Sbjct: 488 KRIGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKALTSE----IALLQSRLKTEGSD 543
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ I+ +QE
Sbjct: 544 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLISGKRQELVTK 599
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 600 IDHILDAL 607
>RIB1_MOUSE (Q91YQ5) Dolichyl-diphosphooligosaccharide--protein
glycosyltransferase 67 kDa subunit precursor (EC
2.4.1.119) (Ribophorin I) (RPN-I)
Length = 608
Score = 315 bits (807), Expect = 2e-85
Identities = 175/488 (35%), Positives = 283/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ VKL +ESYTKL N
Sbjct: 134 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPS 193
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP++++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 194 RSEDVLDYGPFKDIPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 253
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 254 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 312
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+PLFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 313 VEMEIRPRFPLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 372
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 373 IILPEGAKNIQVDSPYDISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 432
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 433 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 488
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ L + + ++ ++ DI + +KS+++ K +T E + LQS + +
Sbjct: 489 KRLGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKAVTSE----IAVLQSRLKTEGSD 544
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ + +QE
Sbjct: 545 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYLENEKLSSGKRQELVTK 600
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 601 IDHILDAL 608
>RIB1_RAT (P07153) Dolichyl-diphosphooligosaccharide--protein
glycosyltransferase 67 kDa subunit precursor (EC
2.4.1.119) (Ribophorin I) (RPN-I)
Length = 605
Score = 308 bits (789), Expect = 2e-83
Identities = 172/488 (35%), Positives = 282/488 (57%), Gaps = 21/488 (4%)
Query: 5 TVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTK 64
TV+TH+L P+P +ITQ++ Q ++F+ + + SPYP K Q++ V+L +ES+TKL N
Sbjct: 131 TVYTHVLHPYPTQITQSEKQFVVFEGNHYFYSPYPTKTQTMRVRLASRNVESHTKLGNPS 190
Query: 65 LQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHYNLVH 124
L YGP++++P +S +H+ENN PF + R IE+SHWGN+ + E+ +L H
Sbjct: 191 RSEDILDYGPFKDIPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENVDLKH 250
Query: 125 GGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGDSKK 184
GA KG FSR DYQ +P G S+ R LP A VYYRDEIGN+STS L
Sbjct: 251 TGAVLKGPFSRYDYQRQP-DSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDS 309
Query: 185 TELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPI-NELVIDTLVVK 243
E+EI PR+ LFGGWKT + +GY LP ++L+ + + L + F + +E VID+L VK
Sbjct: 310 VEMEIRPRFGLFGGWKTHYIVGYNLPSYEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVK 369
Query: 244 VVLPEGSKDISPSVPFPV-KERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYKFN 302
++LPEG+K+I P+ + + E +++LD GRPV+V K N V +H + V+Y FN
Sbjct: 370 IILPEGAKNIQVDSPYDISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHYTFN 429
Query: 303 RLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVS 362
++ ML+EPL++++ F+ LF I+Y+ D SI+K A+ +V QV ++V+
Sbjct: 430 KVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARM----KVACITEQVLTLVN 485
Query: 363 RCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQ 420
+ L + + ++ ++ DI + +KS+++ K +T E + LQS + +
Sbjct: 486 KRLGLYRHFDETVNRYKQSRDISTLNSGKKSLETEHKAVTSE----IAVLQSRLKTEGSD 541
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQE---- 476
+ +V E+ ++ D Q K + S V E+ + G+ ++ +++ + +QE
Sbjct: 542 LCDRVSEM--QKLDAQVKELVLKSAVE--AERLVAGKLKKDTYIENEKLSSGKRQELVTK 597
Query: 477 IDDLMDLI 484
ID ++D +
Sbjct: 598 IDHILDAL 605
>OSTA_SCHPO (Q10176) Putative
dolichyl-diphosphooligosaccharide--protein
glycosyltransferase alpha subunit precursor (EC
2.4.1.119) (Oligosaccharyl transferase alpha subunit)
Length = 450
Score = 122 bits (306), Expect = 2e-27
Identities = 86/330 (26%), Positives = 156/330 (47%), Gaps = 17/330 (5%)
Query: 11 LQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYT--KLENTKLQ-- 66
++P P + Q + Q L++ + SPY +Q + LP A+I++YT ++ +L
Sbjct: 120 VRPLPAAVDQDEEQYLVYLTTLYLDSPYTTDLQRTRLILPSAKIDTYTTYNIDGAELPNR 179
Query: 67 -GSELKYGPYENLP--PFSYLPIVIHFENNQPFAVAKELVREIEISHWGNV-QITEHYNL 122
G+ L Y E + I + +E+ P +L +I++ + + +++E
Sbjct: 180 VGNTLVYESRETITGEDNDLGEIYVRYEHTVPIPRGADLYVQIDMHEYRKMLEVSERVVF 239
Query: 123 VHGGAQSKGEFSRLD-YQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWGD 181
+ A+ K F R Y Y ++A R+ LP + VYY DE+GNI+TS + +
Sbjct: 240 ENHAAKLKSNFDRAKWYMGNFYNPVSTAINRVVYSLPRNSKDVYYTDEVGNITTSHMRVE 299
Query: 182 SKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPINELVIDTLV 241
+T +E+ PRYP+FGGW F + + +P ++F + + +L+I +++ + V
Sbjct: 300 PHQTWIELNPRYPVFGGWNYLFQLDWKMPYEEFRSSKNRREYLDIPLAWSPGDMIYEKAV 359
Query: 242 VKVVLPEGSKDISPSVPFPVKERHETKF-SHLDIAGRPVVVLEKNN---AVPEHNEHFQV 297
V PEG+ +I +P + + TK LD GR V E N VP H
Sbjct: 360 WSYVFPEGATNIEIEMPVEISNSNVTKIHKFLDTLGRQVFTYEALNVTDGVPSDIVHISY 419
Query: 298 YYK----FNRLSMLREPLMLISGFFFLFLA 323
Y F R++++ L+L++ ++ A
Sbjct: 420 EYNSSAFFLRIAIITTLLILLAAAAYMIWA 449
>OSTA_YEAST (P41543) Dolichyl-diphosphooligosaccharide--protein
glycosyltransferase alpha subunit precursor (EC
2.4.1.119) (Oligosaccharyl transferase alpha subunit)
(Oligosaccharyl transferase 64 kDa subunit)
Length = 476
Score = 116 bits (291), Expect = 1e-25
Identities = 93/339 (27%), Positives = 159/339 (46%), Gaps = 22/339 (6%)
Query: 13 PFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKLENTKL----QGS 68
P+PE + ++ Q LL++ + LS Y K S + + + E Y + L G+
Sbjct: 143 PYPEHVGMSEEQHLLWETNRLPLSAYDTKKASFTL-IGSSSFEEYHPPNDESLLGKANGN 201
Query: 69 ELKYGPYENLPPFSYLP-IVIHFENNQPFAVAKELVREIEISHWGN-VQITEHYNLVHGG 126
++GP+E++P FS + I + +N P L R+I +SHW + +Q E+Y L +
Sbjct: 202 SFEFGPWEDIPRFSSNETLAIVYSHNAPLNQVVNLRRDIWLSHWASTIQFEEYYELTNKA 261
Query: 127 AQSKGEFSRLDYQARPYVRGASAFRRLTAK---LPPRAHSVYYRDEIGNISTSSLWGDSK 183
A+ FSRL+ + + +T LP A Y+ D +G +STS ++
Sbjct: 262 AKLSKGFSRLELMKQIQTQNMRQTHFVTVLDMLLPEGATDHYFTDLVGLVSTSH----AE 317
Query: 184 KTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLF---GVDGKRFLNISFGSPINELVIDTL 240
+ I PR+P+FGGW FT+G+ L DFL G D K +I + + V D +
Sbjct: 318 RDHFFIRPRFPIFGGWNYNFTVGWTNKLSDFLHVSSGSDEKFVASIPILNGPPDTVYDNV 377
Query: 241 VVKVVLPEGSK--DISPSVPFPVKERHETKFSHLDI-AGRPVVVLEKNNAVPE-HNEHFQ 296
+ V LPEG++ DI VPF ET+ S+ D+ G + N + + N
Sbjct: 378 ELSVFLPEGAEIFDIDSPVPF-TNVSIETQKSYFDLNKGHVKLTFSYRNLISQVANGQVL 436
Query: 297 VYYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSIS 335
+ Y + + S ++PL + F + V +++++
Sbjct: 437 IKYDYPKSSFFKKPLSIACYIFTALMGVFVLKTLNMNVT 475
>SMC4_XENLA (P50532) Structural maintenance of chromosome 4
(Chromosome-associated protein C) (Chromosome assembly
protein XCAP-C)
Length = 1290
Score = 41.6 bits (96), Expect = 0.005
Identities = 35/140 (25%), Positives = 66/140 (47%), Gaps = 16/140 (11%)
Query: 351 QATIHQVHSIVSRCLTTHDKLEASLRDLSRT--GDIQGCKATRKSV---DSSLKELTKEL 405
Q + +V + C + K + S++ R + T K + D S++ELT++L
Sbjct: 897 QDKLDKVTKEIDECASAITKAQVSIKTADRNLKKSEEAVARTEKEIVANDKSIEELTEDL 956
Query: 406 KS----PVTFLQSSPQAAQILPKVEE----VIAKERDLQEKLMAKHSTVVDCYEKKLGGR 457
K T + +A LP+V+E ++ + + +QEK +H+ + +L
Sbjct: 957 KKLEEKATTVMNECKEAECSLPEVQEQHRSLLQEIKAIQEK---EHALQKEALNIRLNIE 1013
Query: 458 EIENRIASHQQKITALKQEI 477
+I++ IA HQ KI ++EI
Sbjct: 1014 QIDSHIAEHQSKIKYWQKEI 1033
>KF5C_HUMAN (O60282) Kinesin heavy chain isoform 5C (Kinesin heavy
chain neuron-specific 2)
Length = 957
Score = 40.4 bits (93), Expect = 0.010
Identities = 36/156 (23%), Positives = 78/156 (49%), Gaps = 19/156 (12%)
Query: 341 YLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKE 400
Y++K++ EV++ +++ + S + ++ K+ AS R+L+ C+ ++ +K
Sbjct: 590 YISKMK-SEVKSLVNRSKQLESAQMDSNRKMNASERELA------ACQLLISQHEAKIKS 642
Query: 401 LTKELKSPVTFLQSSPQAAQILPKVEEVIAKER--------DLQEKLMAKHSTVVDCYE- 451
LT +++ Q Q + + E +AK R Q+K + + D E
Sbjct: 643 LTDYMQN---MEQKRRQLEESQDSLSEELAKLRAQEKMHEVSFQDKEKEHLTRLQDAEEM 699
Query: 452 KKLGGREIENRIASHQQKITALKQEIDDLMDLIDEI 487
KK +++E+ +HQ++++ L+ EI++ +IDEI
Sbjct: 700 KKALEQQMESHREAHQKQLSRLRDEIEEKQKIIDEI 735
>KF5C_MOUSE (P28738) Kinesin heavy chain isoform 5C (Kinesin heavy
chain neuron-specific 2)
Length = 956
Score = 40.0 bits (92), Expect = 0.014
Identities = 36/156 (23%), Positives = 78/156 (49%), Gaps = 19/156 (12%)
Query: 341 YLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKE 400
Y++K++ EV++ +++ + S + ++ K+ AS R+L+ C+ ++ +K
Sbjct: 589 YISKMK-SEVKSLVNRSKQLESAQMDSNRKMNASERELA------ACQLLISQHEAKIKS 641
Query: 401 LTKELKSPVTFLQSSPQAAQILPKVEEVIAKER--------DLQEKLMAKHSTVVDCYE- 451
LT +++ Q Q + + E +AK R Q+K + + D E
Sbjct: 642 LTDYMQN---MEQKRRQLEESQDSLSEELAKLRAQEKMHEVSFQDKEKEHLTRLQDAEEV 698
Query: 452 KKLGGREIENRIASHQQKITALKQEIDDLMDLIDEI 487
KK +++E+ +HQ++++ L+ EI++ +IDEI
Sbjct: 699 KKALEQQMESHREAHQKQLSRLRDEIEEKQRIIDEI 734
>SMC_METJA (Q59037) Chromosome partition protein smc homolog
Length = 1169
Score = 35.8 bits (81), Expect = 0.26
Identities = 24/106 (22%), Positives = 53/106 (49%), Gaps = 7/106 (6%)
Query: 384 IQGCKATRKSVDSSLKELTKELKSPV----TFLQSSPQAAQILPKVEEVIAKERDLQEKL 439
I K +++ KE+ + K + + ++ Q +I K++ + ++ L+E +
Sbjct: 320 INELKKVEVEIENKKKEIKETQKKIIENRDSIIEKEQQIKEIEEKIKNLNYEKERLKEAI 379
Query: 440 MAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDLMDLID 485
S + E ++ EI + IA +Q ++ LK+E++DL +LI+
Sbjct: 380 AESESIIKHLKESEM---EIADEIAKNQNELYRLKKELNDLDNLIN 422
>NUF2_SCHPO (Q10173) Myosin-like protein NUF2 (Nuclear
filament-containing protein 2) (Nuclear division protein
nuf2)
Length = 441
Score = 35.4 bits (80), Expect = 0.33
Identities = 31/143 (21%), Positives = 67/143 (46%), Gaps = 14/143 (9%)
Query: 350 VQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDI-QGCKATRKSVDSSLKELTKELKSP 408
++ ++ ++ CL DKLE + LS ++ + +K ++ ++L K+L +
Sbjct: 284 IEGDLNACLKVLEECLVELDKLEHATVLLSTNQELCDQIEINKKKLEFRKEQLLKQLSNA 343
Query: 409 VTFL--------QSSPQAAQILPKVEE---VIAKERDLQEKLMAKHSTVVDCYEKKLGG- 456
L Q A Q + + E VI +ER+ + + K + +++ E+K+ G
Sbjct: 344 QEKLEHEQHSRNQKLEAAKQRMDNIREEYKVITQERNKKIQETEKKNAMIEMTEQKIAGM 403
Query: 457 -REIENRIASHQQKITALKQEID 478
E+E++I+S + LK ++
Sbjct: 404 REELESQISSITMEFEKLKSHVE 426
>MYS3_HYDAT (P39922) Myosin heavy chain, clone 203 (Fragment)
Length = 539
Score = 35.4 bits (80), Expect = 0.33
Identities = 42/170 (24%), Positives = 78/170 (45%), Gaps = 35/170 (20%)
Query: 331 DLSISKTSASYLAKIQWEEVQATIHQVHSIVSRCLT---THDKLEASLR----DLSRTGD 383
D +ISK +A K EE++ Q+ + +C T +KLE+S+R DL + D
Sbjct: 189 DETISKMNAE--KKHVDEELKDRTEQLQAAEDKCNNLNKTKNKLESSIREIEQDLKKEKD 246
Query: 384 IQ-GCKATRKSVDSSLKELTKELKSPVTFLQSSP----------------------QAAQ 420
+ + +K V+S LK+ +L T L+ + Q +Q
Sbjct: 247 SKMKLEKEKKKVESDLKDNRDKLSETETRLKETQDLVTKREKSISDLENAKEGLESQISQ 306
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKI 470
+ K++E++AK +L+E+L + + +L +E+E+RI Q ++
Sbjct: 307 LQRKIQELLAKIEELEEELENERKL---RQKSELQRKELESRIEELQDQL 353
>ZY12_CAEEL (Q23529) Zygote defective protein 12
Length = 774
Score = 35.0 bits (79), Expect = 0.44
Identities = 37/143 (25%), Positives = 66/143 (45%), Gaps = 18/143 (12%)
Query: 344 KIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTK 403
KIQ E+ + Q+ R T +++ L+ RT +++ + + + EL
Sbjct: 579 KIQLEKAEVEAQQMREAKMRAETNQAQVDEILK--KRTAELEVNATALQKAKAVIDELEY 636
Query: 404 ELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRI 463
+ PV+ + S + Q +++E K R EKL + +TV +E+ ENR+
Sbjct: 637 NSR-PVS--EDSMTSVQAFKEMKEENEKLRQKVEKLEIELNTVTQGFEQ-------ENRL 686
Query: 464 ---ASHQQKITALKQEIDDLMDL 483
ASHQQ L + ID++M +
Sbjct: 687 LTSASHQQ---VLNRSIDEVMSM 706
>KINN_MOUSE (P33175) Neuronal kinesin heavy chain (NKHC) (Kinesin
heavy chain isoform 5A) (Kinesin heavy chain
neuron-specific 1)
Length = 1027
Score = 35.0 bits (79), Expect = 0.44
Identities = 32/149 (21%), Positives = 71/149 (47%), Gaps = 9/149 (6%)
Query: 341 YLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKE 400
Y++KI+ EV++ + + + + + H K+E + R+LS C+ ++ ++
Sbjct: 590 YISKIK-SEVKSVVKRCRQLENLQVECHRKMEVTGRELS------SCQLLISQHEAKIRS 642
Query: 401 LTKELKSPVTFLQSSPQAAQILP-KVEEVIAKERDLQEKLMAKHSTVVDCYE-KKLGGRE 458
LT+ +++ + ++ L ++ + A E + L K D E KK +
Sbjct: 643 LTEYMQTVELKKRHLEESYDSLSDELARLQAHETVHEVALKDKEPDTQDAEEVKKALELQ 702
Query: 459 IENRIASHQQKITALKQEIDDLMDLIDEI 487
+EN +H +++ L+ EI++ IDE+
Sbjct: 703 MENHREAHHRQLARLRDEINEKQKTIDEL 731
>M18A_HUMAN (Q92614) Myosin XVIIIA (Myosin 18A) (Myosin containing PDZ
domain)
Length = 2054
Score = 34.7 bits (78), Expect = 0.57
Identities = 35/145 (24%), Positives = 70/145 (48%), Gaps = 14/145 (9%)
Query: 331 DLSISKTSAS-YLAKIQWE--EVQATIHQVHSIVSRCLTTHDKLEASL-RDLSRTGDIQG 386
D++ +KT+ L+++Q E E+Q + + ++ + H A RDL++ D+Q
Sbjct: 1722 DIAKAKTALEEQLSRLQREKNEIQNRLEEDQEDMNELMKKHKAAVAQASRDLAQINDLQA 1781
Query: 387 CKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTV 446
L+E + L+S V FL+ S ++ + E AK R+L+ +L + + V
Sbjct: 1782 QLEEANKEKQELQEKLQALQSQVEFLEQSMVDKSLVSRQE---AKIRELETRLEFERTQV 1838
Query: 447 VDCYEKKLGGREIENRIASHQQKIT 471
K+L + +R+ + +K+T
Sbjct: 1839 -----KRL--ESLASRLKENMEKLT 1856
>RA50_THEMA (Q9X1X1) Probable DNA double-strand break repair rad50
ATPase
Length = 852
Score = 33.9 bits (76), Expect = 0.97
Identities = 32/119 (26%), Positives = 56/119 (46%), Gaps = 12/119 (10%)
Query: 369 DKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPKVEEV 428
+KL+ +L T ++ K +KS+ S +++L +++ L+S + + +
Sbjct: 457 EKLDQKRSELENTLNV--LKERKKSLSSLIEDLLMKIEEGKKNLKSIRNQIEKIEEELHR 514
Query: 429 IAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDLMDLIDEI 487
+ DL+EKL K KKL R+IE S QKITA +I + + + EI
Sbjct: 515 LGYSEDLEEKLDEKR--------KKL--RKIEEERHSISQKITAADVQISQIENQLKEI 563
>RRB1_CANFA (Q28298) Ribosome-binding protein 1 (180 kDa ribosome
receptor) (RRp)
Length = 1534
Score = 33.5 bits (75), Expect = 1.3
Identities = 28/107 (26%), Positives = 49/107 (45%), Gaps = 4/107 (3%)
Query: 348 EEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKE-LTKELK 406
E+ QA ++ + ++ +L+A L L TG ++ A + LKE L KE K
Sbjct: 1427 EDEQALRRKLTAEFQEAQSSACRLQAELEKLRSTGPLESSAAEEAT---QLKERLEKEKK 1483
Query: 407 SPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKK 453
++ + ++L +E +AKERD +KL + D K+
Sbjct: 1484 LTSDLGHAATKLQELLKTTQEQLAKERDTVKKLQEQLDKTDDSSSKE 1530
>DYHC_PARTE (Q27171) Dynein heavy chain, cytosolic (DYHC)
Length = 4540
Score = 33.5 bits (75), Expect = 1.3
Identities = 29/114 (25%), Positives = 54/114 (46%), Gaps = 12/114 (10%)
Query: 380 RTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQ-----SSPQAAQILPKVEEVIAKERD 434
R+ +Q ++ ++ S +++ + L + ++ S P + P E I K
Sbjct: 1169 RSKKLQSQESEMNNIQSKIQQDERYLNQQIQEIEEQWKTSKPDSGDCSPNEAEQILKS-- 1226
Query: 435 LQEKLMAKHSTVVDCYEKKLGGREI-ENRIASHQQKITALKQEIDDLMDLIDEI 487
L E+L++ V + YEK +EI + +HQQK+ L + I DL D+ E+
Sbjct: 1227 LNEQLIS----VQEKYEKCSQAKEILKMDPPTHQQKLNVLLESISDLQDVWQEL 1276
>ARHC_HUMAN (Q9NZN5) Rho guanine nucleotide exchange factor 12
(Leukemia-associated RhoGEF)
Length = 1544
Score = 33.5 bits (75), Expect = 1.3
Identities = 34/141 (24%), Positives = 63/141 (44%), Gaps = 7/141 (4%)
Query: 329 HADLSISKTSASYLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCK 388
H +S+ ++ L K + E + +H+ H I + H +++ L D + +
Sbjct: 435 HLKVSVPDEMSADLEKRRPELIPEDLHR-HYIQTMQERVHPEVQRHLEDFRQKRSMGLTL 493
Query: 389 ATRKSVDSSLKELTKEL-KSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVV 447
A +S L +L E K +T + A QI+ K+EEV+ + ++E + V+
Sbjct: 494 A-----ESELTKLDAERDKDRLTLEKERTCAEQIVAKIEEVLMTAQAVEEDKSSTMQYVI 548
Query: 448 DCYEKKLGGREIENRIASHQQ 468
Y K LG + E R H++
Sbjct: 549 LMYMKHLGVKVKEPRNLEHKR 569
>GOA3_HUMAN (Q08378) Golgi autoantigen, golgin subfamily A member 3
(Golgin-160) (Golgi complex-associated protein of 170
kDa) (GCP170)
Length = 1498
Score = 33.1 bits (74), Expect = 1.7
Identities = 24/59 (40%), Positives = 31/59 (51%), Gaps = 1/59 (1%)
Query: 348 EEVQATIHQVHSIVSRCLTTHDKLEASLRDL-SRTGDIQGCKATRKSVDSSLKELTKEL 405
+E A Q+ I + L +EA RD S+ I KATRK +DS LKEL +EL
Sbjct: 829 QETAALKKQMQKIKEQFLQQKVMVEAYRRDATSKDQLISELKATRKRLDSELKELRQEL 887
>DYNA_MOUSE (O08788) Dynactin 1 (150 kDa dynein-associated
polypeptide) (DP-150) (DAP-150) (p150-glued)
Length = 1281
Score = 33.1 bits (74), Expect = 1.7
Identities = 32/120 (26%), Positives = 54/120 (44%), Gaps = 14/120 (11%)
Query: 371 LEASLRDLSRTGDIQGCKATRKSVDSSLKELTK---------ELKSPVTFLQSSPQAAQI 421
L A +RDL ++ + R + LKEL K E KS + Q+ Q
Sbjct: 220 LRAQVRDLEEK--LETLRLKRSEDKAKLKELEKHKIQLEQVQEWKSKMQEQQADLQRR-- 275
Query: 422 LPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREI-ENRIASHQQKITALKQEIDDL 480
L + + + + +E+ M + + D E +E+ E R S QQ++ ALK+ +D+L
Sbjct: 276 LKEARKEAKEALEAKERYMEEMADTADAIEMATLDKEMAEERAESLQQEVEALKERVDEL 335
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 56,175,625
Number of Sequences: 164201
Number of extensions: 2408253
Number of successful extensions: 6995
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 56
Number of HSP's that attempted gapping in prelim test: 6925
Number of HSP's gapped (non-prelim): 112
length of query: 487
length of database: 59,974,054
effective HSP length: 114
effective length of query: 373
effective length of database: 41,255,140
effective search space: 15388167220
effective search space used: 15388167220
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC126786.5


