

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126781.3 + phase: 0
(77 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
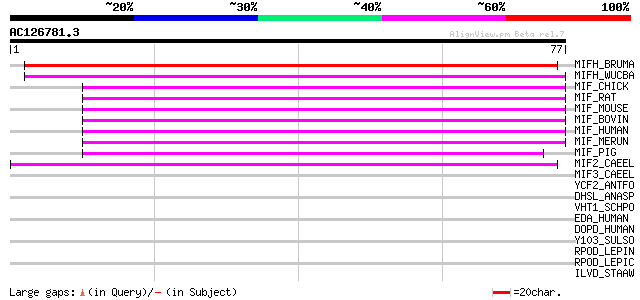
Score E
Sequences producing significant alignments: (bits) Value
MIFH_BRUMA (P91850) Macrophage migration inhibitory factor homol... 62 3e-10
MIFH_WUCBA (O44786) Macrophage migration inhibitory factor homolog 57 9e-09
MIF_CHICK (Q02960) Macrophage migration inhibitory factor (MIF) ... 52 2e-07
MIF_RAT (P30904) Macrophage migration inhibitory factor (MIF) (P... 50 8e-07
MIF_MOUSE (P34884) Macrophage migration inhibitory factor (MIF) ... 50 8e-07
MIF_BOVIN (P80177) Macrophage migration inhibitory factor (MIF) ... 49 2e-06
MIF_HUMAN (P14174) Macrophage migration inhibitory factor (MIF) ... 48 4e-06
MIF_MERUN (O55052) Macrophage migration inhibitory factor (MIF) ... 45 3e-05
MIF_PIG (P80928) Macrophage migration inhibitory factor (MIF) (P... 45 3e-05
MIF2_CAEEL (Q18785) MIF-like protein mif-2 42 4e-04
MIF3_CAEEL (P90835) MIF-like protein mif-3 29 1.9
YCF2_ANTFO (Q859W7) Protein ycf2 28 5.6
DHSL_ANASP (Q8YQL7) Deoxyhypusine synthase-like protein (EC 2.5.... 28 5.6
VHT1_SCHPO (O13880) Vitamin H transporter 1 (H+/biotin symporter... 27 7.3
EDA_HUMAN (Q92838) Ectodysplasin A (Ectodermal dysplasia protein... 27 7.3
DOPD_HUMAN (P30046) D-dopachrome tautomerase (EC 5.3.3.-) (Pheny... 27 7.3
Y103_SULSO (P95958) Hypothetical protein SSO0103 27 9.5
RPOD_LEPIN (P59117) RNA polymerase sigma factor rpoD 27 9.5
RPOD_LEPIC (P61540) RNA polymerase sigma factor rpoD 27 9.5
ILVD_STAAW (P65158) Dihydroxy-acid dehydratase (EC 4.2.1.9) (DAD) 27 9.5
>MIFH_BRUMA (P91850) Macrophage migration inhibitory factor homolog
(BmMIF) (Bm-MIF-1)
Length = 114
Score = 61.6 bits (148), Expect = 3e-10
Identities = 32/74 (43%), Positives = 46/74 (61%)
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
I VNGG A+ FGG+E+P A L SIG + P VN + + +++ L I +R YI+F
Sbjct: 39 IHVNGGQAMVFGGSEDPCAVCVLKSIGCVGPKVNNSHAEKLYKLLADELKIPKNRCYIEF 98
Query: 63 SDVQPSFVGFNGST 76
D++ S + FNGST
Sbjct: 99 VDIEASSMAFNGST 112
>MIFH_WUCBA (O44786) Macrophage migration inhibitory factor homolog
Length = 114
Score = 57.0 bits (136), Expect = 9e-09
Identities = 30/75 (40%), Positives = 44/75 (58%)
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKF 62
I VNGG + FGG+E+P L SIG + P VN + + +++ L I +R YI+
Sbjct: 39 IHVNGGQPMVFGGSEDPCPVCVLKSIGCVGPKVNNSHAEKLYKLLADELKIPKNRCYIES 98
Query: 63 SDVQPSFVGFNGSTF 77
D++ S + FNGSTF
Sbjct: 99 VDIEASSMAFNGSTF 113
>MIF_CHICK (Q02960) Macrophage migration inhibitory factor (MIF)
(Phenylpyruvate tautomerase) (EC 5.3.2.1)
Length = 114
Score = 52.4 bits (124), Expect = 2e-07
Identities = 24/67 (35%), Positives = 39/67 (57%)
Query: 11 IAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFV 70
++FGG+ +P A L SIG + N + + +I +LH+ + R YI + D+ + V
Sbjct: 47 MSFGGSTDPCALCSLYSIGKIGGQQNKTYTKLLCDMIAKHLHVSADRVYINYFDINAANV 106
Query: 71 GFNGSTF 77
G+NGSTF
Sbjct: 107 GWNGSTF 113
>MIF_RAT (P30904) Macrophage migration inhibitory factor (MIF)
(Phenylpyruvate tautomerase) (EC 5.3.2.1)
(Glutathione-binding 13 kDa protein)
Length = 114
Score = 50.4 bits (119), Expect = 8e-07
Identities = 25/67 (37%), Positives = 35/67 (51%)
Query: 11 IAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFV 70
+ F GT +P A L SIG + N S + ++ LHI R YI + D+ + V
Sbjct: 47 MTFSGTSDPCALCSLHSIGKIGGAQNRNYSKLLCGLLSDRLHISPDRVYINYYDMNAANV 106
Query: 71 GFNGSTF 77
G+NGSTF
Sbjct: 107 GWNGSTF 113
>MIF_MOUSE (P34884) Macrophage migration inhibitory factor (MIF)
(Phenylpyruvate tautomerase) (EC 5.3.2.1)
(Glycosylation-inhibiting factor) (GIF) (Delayed early
response protein 6) (DER6)
Length = 114
Score = 50.4 bits (119), Expect = 8e-07
Identities = 25/67 (37%), Positives = 35/67 (51%)
Query: 11 IAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFV 70
+ F GT +P A L SIG + N S + ++ LHI R YI + D+ + V
Sbjct: 47 MTFSGTNDPCALCSLHSIGKIGGAQNRNYSKLLCGLLSDRLHISPDRVYINYYDMNAANV 106
Query: 71 GFNGSTF 77
G+NGSTF
Sbjct: 107 GWNGSTF 113
>MIF_BOVIN (P80177) Macrophage migration inhibitory factor (MIF)
(Phenylpyruvate tautomerase) (EC 5.3.2.1) (p12A)
Length = 114
Score = 49.3 bits (116), Expect = 2e-06
Identities = 25/67 (37%), Positives = 35/67 (51%)
Query: 11 IAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFV 70
+ FGG+ EP A L SIG + N S + ++ L I R YI + D+ + V
Sbjct: 47 MTFGGSSEPCALCSLHSIGKIGGAQNRSYSKLLCGLLTERLRISPDRIYINYYDMNAANV 106
Query: 71 GFNGSTF 77
G+NGSTF
Sbjct: 107 GWNGSTF 113
>MIF_HUMAN (P14174) Macrophage migration inhibitory factor (MIF)
(Phenylpyruvate tautomerase) (EC 5.3.2.1)
(Glycosylation-inhibiting factor) (GIF)
Length = 114
Score = 48.1 bits (113), Expect = 4e-06
Identities = 25/67 (37%), Positives = 35/67 (51%)
Query: 11 IAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFV 70
+AFGG+ EP A L SIG + N S + ++ L I R YI + D+ + V
Sbjct: 47 MAFGGSSEPCALCSLHSIGKIGGAQNRSYSKLLCGLLAERLRISPDRVYINYYDMNAANV 106
Query: 71 GFNGSTF 77
G+N STF
Sbjct: 107 GWNNSTF 113
>MIF_MERUN (O55052) Macrophage migration inhibitory factor (MIF)
(Phenylpyruvate tautomerase) (EC 5.3.2.1)
Length = 114
Score = 45.4 bits (106), Expect = 3e-05
Identities = 23/67 (34%), Positives = 34/67 (50%)
Query: 11 IAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFV 70
+ F G+ +P A L SIG + N S + ++ L I R YI + D+ + V
Sbjct: 47 MTFSGSSDPCALCSLHSIGKIGGAQNRTYSKLLCGLLADRLRISPDRIYINYYDMNAANV 106
Query: 71 GFNGSTF 77
G+NGSTF
Sbjct: 107 GWNGSTF 113
>MIF_PIG (P80928) Macrophage migration inhibitory factor (MIF)
(Phenylpyruvate tautomerase) (EC 5.3.2.1)
(Glycosylation-inhibiting factor) (GIF) (Fragment)
Length = 110
Score = 45.1 bits (105), Expect = 3e-05
Identities = 23/64 (35%), Positives = 33/64 (50%)
Query: 11 IAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFV 70
+AFGG+ EP A L SIG + N S + ++ L I R YI + D+ + V
Sbjct: 47 MAFGGSSEPCALCSLHSIGKIGGAQNRSYSKLLCGLLAERLRISPDRIYINYYDMNAANV 106
Query: 71 GFNG 74
G+NG
Sbjct: 107 GWNG 110
>MIF2_CAEEL (Q18785) MIF-like protein mif-2
Length = 120
Score = 41.6 bits (96), Expect = 4e-04
Identities = 20/76 (26%), Positives = 36/76 (47%)
Query: 1 MCILVNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYI 60
+ + + G + G T +P + SIG + N + ++ I + L + + I
Sbjct: 38 IAVEIAAGARLVHGATHDPVTVISIKSIGAVSAEDNIRNTAAITEFCGKELGLPKDKVVI 97
Query: 61 KFSDVQPSFVGFNGST 76
F D+ P+ VGFNG+T
Sbjct: 98 TFHDLPPATVGFNGTT 113
>MIF3_CAEEL (P90835) MIF-like protein mif-3
Length = 146
Score = 29.3 bits (64), Expect = 1.9
Identities = 14/61 (22%), Positives = 26/61 (41%)
Query: 14 GGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSDVQPSFVGFN 73
G +P A ++ S L P + + + + + L + S I + + P +GFN
Sbjct: 49 GQLTDPLAVLDVTSSTVLTPILTEEYTVALCEFFSQELALDSDAVLINYRSLSPELIGFN 108
Query: 74 G 74
G
Sbjct: 109 G 109
>YCF2_ANTFO (Q859W7) Protein ycf2
Length = 2392
Score = 27.7 bits (60), Expect = 5.6
Identities = 20/70 (28%), Positives = 32/70 (45%), Gaps = 6/70 (8%)
Query: 11 IAFGGTEEPAAYGE-LISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIK-----FSD 64
I G T P LIS LD +N ++ + + ++ +HS RFY K F++
Sbjct: 1770 IIIGSTHLPRKVDPALISPNRLDKLMNIRIIDIFERQEKFSILLHSKRFYFKNKLLHFNE 1829
Query: 65 VQPSFVGFNG 74
+G+NG
Sbjct: 1830 FAYRTMGYNG 1839
>DHSL_ANASP (Q8YQL7) Deoxyhypusine synthase-like protein (EC
2.5.-.-)
Length = 383
Score = 27.7 bits (60), Expect = 5.6
Identities = 18/72 (25%), Positives = 31/72 (43%), Gaps = 2/72 (2%)
Query: 5 VNGGVAIAFGGTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRFYIKFSD 64
VN AIA+ E +I GG + I +++ H F+++F+D
Sbjct: 223 VNETAAIAYNAREGEGKSAAVILGGGSPKNFLLQTQPQIHEVLGLEERGHD--FFVQFTD 280
Query: 65 VQPSFVGFNGST 76
+P G +G+T
Sbjct: 281 ARPDTGGLSGAT 292
>VHT1_SCHPO (O13880) Vitamin H transporter 1 (H+/biotin symporter
vht1)
Length = 568
Score = 27.3 bits (59), Expect = 7.3
Identities = 16/49 (32%), Positives = 27/49 (54%), Gaps = 1/49 (2%)
Query: 2 CILVNGGVAIAFGGTEEPAAYGELISIG-GLDPTVNAKLSSTIAQIIQT 49
C+++ G+A+A + YG L+ IG GL PTV ++ A + +T
Sbjct: 418 CLIIIAGLAVANYAPRAWSRYGGLLMIGFGLGPTVPICMAWCSASMAKT 466
>EDA_HUMAN (Q92838) Ectodysplasin A (Ectodermal dysplasia protein)
(EDA protein)
Length = 391
Score = 27.3 bits (59), Expect = 7.3
Identities = 12/26 (46%), Positives = 14/26 (53%)
Query: 8 GVAIAFGGTEEPAAYGELISIGGLDP 33
G GG+ P G L S+GGLDP
Sbjct: 73 GAESRLGGSGTPGTSGTLSSLGGLDP 98
>DOPD_HUMAN (P30046) D-dopachrome tautomerase (EC 5.3.3.-)
(Phenylpyruvate tautomerase II)
Length = 117
Score = 27.3 bits (59), Expect = 7.3
Identities = 19/74 (25%), Positives = 34/74 (45%), Gaps = 1/74 (1%)
Query: 3 ILVNGGVAIAFGGTEEPAAYGELISIGGLDPTV-NAKLSSTIAQIIQTNLHIHSSRFYIK 61
+ V G+A+A G+ EP A + SIG + N S+ + + L + R I+
Sbjct: 39 VTVRPGLAMALSGSTEPCAQLSISSIGVVGTAEDNRSHSAHFFEFLTKELALGQDRILIR 98
Query: 62 FSDVQPSFVGFNGS 75
F ++ +G G+
Sbjct: 99 FFPLESWQIGKIGT 112
>Y103_SULSO (P95958) Hypothetical protein SSO0103
Length = 272
Score = 26.9 bits (58), Expect = 9.5
Identities = 11/34 (32%), Positives = 21/34 (61%)
Query: 25 LISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSRF 58
++S GGL PT + K + +A+ + NL ++ + F
Sbjct: 75 IVSSGGLGPTWDDKTAEGLAKALGVNLELNKTAF 108
>RPOD_LEPIN (P59117) RNA polymerase sigma factor rpoD
Length = 585
Score = 26.9 bits (58), Expect = 9.5
Identities = 15/43 (34%), Positives = 24/43 (54%)
Query: 15 GTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSR 57
G+EE + G+ I ++ VNA SS +A+ I+ LH +R
Sbjct: 484 GSEEDSELGDFIPDTEVETPVNAAASSILAEQIRQVLHTLPAR 526
>RPOD_LEPIC (P61540) RNA polymerase sigma factor rpoD
Length = 585
Score = 26.9 bits (58), Expect = 9.5
Identities = 15/43 (34%), Positives = 24/43 (54%)
Query: 15 GTEEPAAYGELISIGGLDPTVNAKLSSTIAQIIQTNLHIHSSR 57
G+EE + G+ I ++ VNA SS +A+ I+ LH +R
Sbjct: 484 GSEEDSELGDFIPDTEVETPVNAAASSILAEQIRQVLHTLPAR 526
>ILVD_STAAW (P65158) Dihydroxy-acid dehydratase (EC 4.2.1.9) (DAD)
Length = 562
Score = 26.9 bits (58), Expect = 9.5
Identities = 12/29 (41%), Positives = 19/29 (65%), Gaps = 3/29 (10%)
Query: 7 GGVAIAFGGTEEPAAYGELISIGGLDPTV 35
GG++I FG A G +I +GG+DP++
Sbjct: 375 GGLSILFGNI---APKGAVIKVGGVDPSI 400
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.139 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,650,245
Number of Sequences: 164201
Number of extensions: 266337
Number of successful extensions: 976
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 961
Number of HSP's gapped (non-prelim): 22
length of query: 77
length of database: 59,974,054
effective HSP length: 53
effective length of query: 24
effective length of database: 51,271,401
effective search space: 1230513624
effective search space used: 1230513624
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126781.3


