

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126018.1 + phase: 0 /pseudo
(596 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
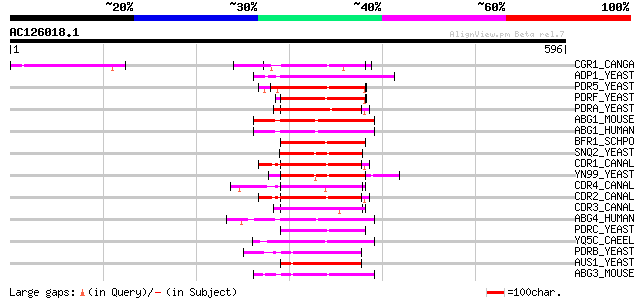
Score E
Sequences producing significant alignments: (bits) Value
CGR1_CANGA (O74208) ATP-binding cassette transporter CGR1 (Pleom... 101 6e-21
ADP1_YEAST (P25371) Probable ATP-dependent permease precursor 96 3e-19
PDR5_YEAST (P33302) Suppressor of toxicity of sporidesmin 95 5e-19
PDRF_YEAST (Q04182) ATP-dependent permease PDR15 92 5e-18
PDRA_YEAST (P51533) ATP-dependent permease PDR10 90 1e-17
ABG1_MOUSE (Q64343) ATP-binding cassette, sub-family G, member 1... 88 7e-17
ABG1_HUMAN (P45844) ATP-binding cassette, sub-family G, member 1... 86 2e-16
BFR1_SCHPO (P41820) Brefeldin A resistance protein 82 5e-15
SNQ2_YEAST (P32568) SNQ2 protein 78 6e-14
CDR1_CANAL (P43071) Multidrug resistance protein CDR1 78 6e-14
YN99_YEAST (P53756) Probable ATP-dependent transporter YNR070W 77 1e-13
CDR4_CANAL (O74676) ABC transporter CDR4 77 2e-13
CDR2_CANAL (P78595) Multidrug resistance protein CDR2 76 3e-13
CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3 75 4e-13
ABG4_HUMAN (Q9H172) ATP-binding cassette, sub-family G, member 4 74 8e-13
PDRC_YEAST (Q02785) ATP-dependent permease PDR12 72 5e-12
YQ5C_CAEEL (Q09466) Putative ABC transporter C16C10.12 in chromo... 69 3e-11
PDRB_YEAST (P40550) ATP-dependent permease PDR11 65 4e-10
AUS1_YEAST (Q08409) ATP-dependent permease AUS1 65 4e-10
ABG3_MOUSE (Q99P81) ATP-binding cassette, sub-family G, member 3 64 9e-10
>CGR1_CANGA (O74208) ATP-binding cassette transporter CGR1
(Pleomorphic drug resistance homolog)
Length = 1542
Score = 101 bits (251), Expect = 6e-21
Identities = 58/142 (40%), Positives = 80/142 (55%), Gaps = 10/142 (7%)
Query: 241 ADSVTESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFR 300
++S+T S G + L + ++ Y V + E++ +L V G +
Sbjct: 862 SESITSGSRGGSPQVGLSKSEAIFHWQNLCYDVPIKTEVRR---------ILNNVDGWVK 912
Query: 301 PGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIH 360
PG LTALMG SGAGKTTL+D LA R T G I GD+ V+G P + +F+R GYC+ D+H
Sbjct: 913 PGTLTALMGASGAGKTTLLDCLAERTTMGVITGDVMVNGRP-RDTSFSRSIGYCQQQDLH 971
Query: 361 SPHVTVHESLLYSAWLRLPSGV 382
TV ESL +SA+LR PS V
Sbjct: 972 LKTATVRESLRFSAYLRQPSSV 993
Score = 46.6 bits (109), Expect = 2e-04
Identities = 37/127 (29%), Positives = 61/127 (47%), Gaps = 11/127 (8%)
Query: 273 VDMPVEM------KEQGVRE-DRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGR 325
+++PV++ K + RE D +LK + G +PG L ++G G+G TTL+ ++
Sbjct: 149 LNLPVKLLNAVWRKARPARESDTFRILKPMDGLLKPGELLVVLGRPGSGCTTLLKSISST 208
Query: 326 KTGGYIDGDIKVS-GYPKKQETFARISGYCEHN---DIHSPHVTVHESLLYSAWLRLPSG 381
G I D +S E G +N DIH PH+TV+++L+ A L+ P
Sbjct: 209 THGFQISKDSVISYNGLTPNEIKKHYRGEVVYNAEADIHLPHLTVYQTLVTVARLKTPQN 268
Query: 382 VDSNTTK 388
T+
Sbjct: 269 RVKGVTR 275
Score = 44.7 bits (104), Expect = 7e-04
Identities = 36/128 (28%), Positives = 58/128 (45%), Gaps = 6/128 (4%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGA-LFFTLVTMMFNGMSEISMTIAKLPVYYKQRD 59
MA I ++F++ + + D + GA +FF ++ F+ + EI P+ K R
Sbjct: 532 MAFILGSMFYKIQ--KGSSADTFYFRGAAMFFAILFNAFSSLLEIFSLYEARPITEKHRT 589
Query: 60 LLFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFR---CVVFHES 116
Y A A S I +IP ++ L+ + Y+++ F + GR F F VF S
Sbjct: 590 YSLYHPSADAFASVISEIPPKIVTAILFNIIFYFLVNFRRDAGRFFFYFLINVIAVFAMS 649
Query: 117 DGFRIVPS 124
FR V S
Sbjct: 650 HLFRCVGS 657
>ADP1_YEAST (P25371) Probable ATP-dependent permease precursor
Length = 1049
Score = 95.5 bits (236), Expect = 3e-19
Identities = 55/153 (35%), Positives = 90/153 (57%), Gaps = 8/153 (5%)
Query: 263 SITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVL 322
+++F++I YSV + GV E +L +SG +PG + A+MG SGAGKTTL+D+L
Sbjct: 383 TLSFENITYSVP---SINSDGVEE---TVLNEISGIVKPGQILAIMGGSGAGKTTLLDIL 436
Query: 323 AGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
A ++ G++ G IKV+G +++F++I G+ + +D P +TV E++L SA LRLP +
Sbjct: 437 AMKRKTGHVSGSIKVNGISMDRKSFSKIIGFVDQDDFLLPTLTVFETVLNSALLRLPKAL 496
Query: 383 DSNTTKVSEHVLYSEWIWLHLLTLL--NEHDRG 413
K + + E + + + NE DRG
Sbjct: 497 SFEAKKARVYKVLEELRIIDIKDRIIGNEFDRG 529
>PDR5_YEAST (P33302) Suppressor of toxicity of sporidesmin
Length = 1511
Score = 95.1 bits (235), Expect = 5e-19
Identities = 51/102 (50%), Positives = 66/102 (64%), Gaps = 1/102 (0%)
Query: 281 EQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGY 340
E ++ + +L V G +PG LTALMG SGAGKTTL+D LA R T G I GDI V+G
Sbjct: 877 EVQIKAETRRILNNVDGWVKPGTLTALMGASGAGKTTLLDCLAERVTMGVITGDILVNGI 936
Query: 341 PKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
P + ++F R GYC+ D+H TV ESL +SA+LR P+ V
Sbjct: 937 P-RDKSFPRSIGYCQQQDLHLKTATVRESLRFSAYLRQPAEV 977
Score = 52.0 bits (123), Expect = 4e-06
Identities = 40/134 (29%), Positives = 66/134 (48%), Gaps = 18/134 (13%)
Query: 268 DIVYS---VDMPVEMKEQGVRE-------DRLVLLKGVSGAFRPGVLTALMGVSGAGKTT 317
D+ Y V++P ++ + G+R+ + +LK + G PG L ++G G+G TT
Sbjct: 142 DVAYQSTVVNIPYKILKSGLRKFQRSKETNTFQILKPMDGCLNPGELLVVLGRPGSGCTT 201
Query: 318 LMDVLAGRKTGGYIDGDIKV--SGYPKK--QETFARISGYCEHNDIHSPHVTVHESLLYS 373
L+ ++ G + D K+ SGY ++ F Y D+H PH+TV E+L+
Sbjct: 202 LLKSISSNTHGFDLGADTKISYSGYSGDDIKKHFRGEVVYNAEADVHLPHLTVFETLVTV 261
Query: 374 AWLRLP----SGVD 383
A L+ P GVD
Sbjct: 262 ARLKTPQNRIKGVD 275
Score = 41.2 bits (95), Expect = 0.008
Identities = 36/127 (28%), Positives = 54/127 (42%), Gaps = 4/127 (3%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
MALI ++FF+ M + D A+FF ++ F+ + EI P+ K R
Sbjct: 533 MALILGSMFFKI-MKKGDTSTFYFRGSAMFFAILFNAFSSLLEIFSLYEARPITEKHRTY 591
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQFR---CVVFHESD 117
Y A A S + +IP L+ + + Y+++ F N G F VF S
Sbjct: 592 SLYHPSADAFASVLSEIPSKLIIAVCFNIIFYFLVDFRRNGGVFFFYLLINIVAVFSMSH 651
Query: 118 GFRIVPS 124
FR V S
Sbjct: 652 LFRCVGS 658
>PDRF_YEAST (Q04182) ATP-dependent permease PDR15
Length = 1529
Score = 91.7 bits (226), Expect = 5e-18
Identities = 50/92 (54%), Positives = 60/92 (64%), Gaps = 1/92 (1%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARI 350
+L V G +PG LTALMG SGAGKTTL+D LA R T G I G+I V G + E+F R
Sbjct: 902 ILNNVDGWVKPGTLTALMGASGAGKTTLLDCLAERVTMGVITGNIFVDG-RLRDESFPRS 960
Query: 351 SGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
GYC+ D+H TV ESL +SA+LR PS V
Sbjct: 961 IGYCQQQDLHLKTATVRESLRFSAYLRQPSSV 992
Score = 47.0 bits (110), Expect = 1e-04
Identities = 32/106 (30%), Positives = 52/106 (48%), Gaps = 8/106 (7%)
Query: 286 EDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI--DGDIKVSGYPKK 343
ED +LK + G PG L ++G G+G TTL+ ++ G I D + +G
Sbjct: 180 EDTFQILKPMDGCLNPGELLVVLGRPGSGCTTLLKSISSNSHGFKIAKDSIVSYNGLSSS 239
Query: 344 --QETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLP----SGVD 383
++ + Y +DIH PH+TV+++L A ++ P GVD
Sbjct: 240 DIRKHYRGEVVYNAESDIHLPHLTVYQTLFTVARMKTPQNRIKGVD 285
Score = 40.0 bits (92), Expect = 0.017
Identities = 28/108 (25%), Positives = 48/108 (43%), Gaps = 1/108 (0%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
MA I ++F++ M +ND A+FF ++ F+ + EI P+ K R
Sbjct: 543 MAFILGSMFYKV-MKKNDTSTFYFRGAAMFFAILFNAFSCLLEIFSLYETRPITEKHRTY 601
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQF 108
Y A A S + ++P L+ + + Y+++ F N G F F
Sbjct: 602 SLYHPSADAFASVLSEMPPKLITAVCFNIIFYFLVDFRRNGGVFFFYF 649
>PDRA_YEAST (P51533) ATP-dependent permease PDR10
Length = 1564
Score = 90.1 bits (222), Expect = 1e-17
Identities = 47/94 (50%), Positives = 61/94 (64%), Gaps = 1/94 (1%)
Query: 284 VREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKK 343
++ + +L V G +PG LTAL+G SGAGKTTL+D LA R T G I GD+ V G P+
Sbjct: 934 IKNGKRRILDNVDGWVKPGTLTALIGASGAGKTTLLDCLAERTTMGLITGDVFVDGRPRD 993
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSAWLR 377
Q +F R GYC+ D+H TV ESL +SA+LR
Sbjct: 994 Q-SFPRSIGYCQQQDLHLKTATVRESLRFSAYLR 1026
Score = 48.5 bits (114), Expect = 5e-05
Identities = 35/104 (33%), Positives = 52/104 (49%), Gaps = 8/104 (7%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGD--IKVSGYPKKQ--ET 346
+LK + G PG L ++G GAG TTL+ ++ G I D I +G+ K+
Sbjct: 195 ILKPMDGCINPGELLVVLGRPGAGCTTLLKSISVNTHGFKISPDTIITYNGFSNKEIKNH 254
Query: 347 FARISGYCEHNDIHSPHVTVHESLLYSAWLRLP----SGVDSNT 386
+ Y +DIH PH+TV ++L A L+ P GVD +T
Sbjct: 255 YRGEVVYNAESDIHIPHLTVFQTLYTVARLKTPRNRIKGVDRDT 298
>ABG1_MOUSE (Q64343) ATP-binding cassette, sub-family G, member 1
(White protein homolog) (ATP-binding cassette
transporter 8)
Length = 666
Score = 87.8 bits (216), Expect = 7e-17
Identities = 52/130 (40%), Positives = 78/130 (60%), Gaps = 7/130 (5%)
Query: 263 SITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVL 322
+I F D+ YSV K++G + LLKG+SG F G L A+MG SGAGK+TLM++L
Sbjct: 76 NIEFKDLSYSVPEGPWWKKKGYK----TLLKGISGKFNSGELVAIMGPSGAGKSTLMNIL 131
Query: 323 AG-RKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSG 381
AG R+TG + G + ++G P+ F ++S Y +D+ PH+TV E+++ SA L+L
Sbjct: 132 AGYRETG--MKGAVLINGMPRDLRCFRKVSCYIMQDDMLLPHLTVQEAMMVSAHLKLQEK 189
Query: 382 VDSNTTKVSE 391
+ V E
Sbjct: 190 DEGRREMVKE 199
>ABG1_HUMAN (P45844) ATP-binding cassette, sub-family G, member 1
(White protein homolog) (ATP-binding cassette
transporter 8)
Length = 678
Score = 86.3 bits (212), Expect = 2e-16
Identities = 51/130 (39%), Positives = 78/130 (59%), Gaps = 7/130 (5%)
Query: 263 SITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVL 322
+I F D+ YSV +++G + LLKG+SG F G L A+MG SGAGK+TLM++L
Sbjct: 76 NIEFRDLSYSVPEGPWWRKKGYK----TLLKGISGKFNSGELVAIMGPSGAGKSTLMNIL 131
Query: 323 AG-RKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSG 381
AG R+TG + G + ++G P+ F ++S Y +D+ PH+TV E+++ SA L+L
Sbjct: 132 AGYRETG--MKGAVLINGLPRDLRCFRKVSCYIMQDDMLLPHLTVQEAMMVSAHLKLQEK 189
Query: 382 VDSNTTKVSE 391
+ V E
Sbjct: 190 DEGRREMVKE 199
>BFR1_SCHPO (P41820) Brefeldin A resistance protein
Length = 1530
Score = 81.6 bits (200), Expect = 5e-15
Identities = 47/92 (51%), Positives = 58/92 (62%), Gaps = 1/92 (1%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARI 350
LL GV G PG LTALMG SGAGKTTL++VLA R G + GD+ V+G TF R
Sbjct: 900 LLNGVQGFVVPGKLTALMGESGAGKTTLLNVLAQRVDTGVVTGDMLVNG-RGLDSTFQRR 958
Query: 351 SGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
+GY + D+H TV E+L +SA LR P+ V
Sbjct: 959 TGYVQQQDVHIGESTVREALRFSAALRQPASV 990
Score = 41.2 bits (95), Expect = 0.008
Identities = 25/107 (23%), Positives = 50/107 (46%), Gaps = 3/107 (2%)
Query: 2 ALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLL 61
+LI ++F+ +++ D G G LFF+++ +SEI+ ++ P+ K R
Sbjct: 552 SLIIGSIFYDMKLNTVDVFSRG---GVLFFSILFCALQSLSEIANMFSQRPIIAKHRASA 608
Query: 62 FYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMFKQF 108
Y A I S I+ +P + +S++ + Y++ G + F
Sbjct: 609 LYHPAADVISSLIVDLPFRFINISVFSIVLYFLTNLKRTAGGFWTYF 655
Score = 32.0 bits (71), Expect = 4.7
Identities = 25/90 (27%), Positives = 41/90 (44%), Gaps = 3/90 (3%)
Query: 302 GVLTALMGVSGAGKTT-LMDVLAGRKTGGYIDGDIKVSGYPKK--QETFARISGYCEHND 358
G L ++G G+G +T L V + ++G G K ++ F Y ND
Sbjct: 187 GELVMVLGQPGSGCSTFLRSVTSDTVHYKRVEGTTHYDGIDKADMKKFFPGDLLYSGEND 246
Query: 359 IHSPHVTVHESLLYSAWLRLPSGVDSNTTK 388
+H P +T E+L ++A R P+ N T+
Sbjct: 247 VHFPSLTTAETLDFAAKCRTPNNRPCNLTR 276
>SNQ2_YEAST (P32568) SNQ2 protein
Length = 1501
Score = 78.2 bits (191), Expect = 6e-14
Identities = 46/90 (51%), Positives = 58/90 (64%), Gaps = 2/90 (2%)
Query: 290 VLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFAR 349
+LL VSG PG +TALMG SGAGKTTL++ LA R G I GD+ V+G P +F R
Sbjct: 870 MLLDNVSGYCIPGTMTALMGESGAGKTTLLNTLAQRNV-GIITGDMLVNGRP-IDASFER 927
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSAWLRLP 379
+GY + DIH +TV ESL +SA +R P
Sbjct: 928 RTGYVQQQDIHIAELTVRESLQFSARMRRP 957
>CDR1_CANAL (P43071) Multidrug resistance protein CDR1
Length = 1501
Score = 78.2 bits (191), Expect = 6e-14
Identities = 49/111 (44%), Positives = 71/111 (63%), Gaps = 5/111 (4%)
Query: 268 DIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKT 327
+I + D+ ++K + +EDR V+L V G +PG +TALMG SGAGKTTL++ L+ R T
Sbjct: 857 EIFFWRDLTYQVKIK--KEDR-VILDHVDGWVKPGQITALMGASGAGKTTLLNCLSERVT 913
Query: 328 GGYI-DGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLR 377
G I DG+ V+G+ +F R GY + D+H P TV E+L +SA+LR
Sbjct: 914 TGIITDGERLVNGH-ALDSSFQRSIGYVQQQDVHLPTSTVREALQFSAYLR 963
Score = 48.5 bits (114), Expect = 5e-05
Identities = 35/104 (33%), Positives = 52/104 (49%), Gaps = 8/104 (7%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI--DGDIKVSGY-PKKQETF 347
+LK + RPG LT ++G GAG +TL+ +A G +I + I G P E
Sbjct: 169 ILKSMDAIMRPGELTVVLGRPGAGCSTLLKTIAVNTYGFHIGKESQITYDGLSPHDIERH 228
Query: 348 ARISG-YCEHNDIHSPHVTVHESLLYSAWLRLP----SGVDSNT 386
R Y D+H PH++V ++L ++A LR P G+D T
Sbjct: 229 YRGDVIYSAETDVHFPHLSVGDTLEFAARLRTPQNRGEGIDRET 272
Score = 42.0 bits (97), Expect = 0.005
Identities = 27/104 (25%), Positives = 45/104 (42%), Gaps = 2/104 (1%)
Query: 4 IAMTLFFRTEMHRNDQDDAGVY--AGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLL 61
+ M L + + Q Y A+FF ++ F+ + EI P+ K +
Sbjct: 523 LVMGLILSSVFYNLSQTTGSFYYRGAAMFFAVLFNAFSSLLEIMSLFEARPIVEKHKKYA 582
Query: 62 FYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF 105
Y A A+ S I ++PV L + F+ Y+++ F N GR F
Sbjct: 583 LYRPSADALASIISELPVKLAMSMSFNFVFYFMVNFRRNPGRFF 626
>YN99_YEAST (P53756) Probable ATP-dependent transporter YNR070W
Length = 1333
Score = 77.4 bits (189), Expect = 1e-13
Identities = 45/92 (48%), Positives = 57/92 (61%), Gaps = 2/92 (2%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARI 350
LL VSG PG LTAL+G SGAGKTTL++ LA R G I GD+ V G P +F R
Sbjct: 747 LLDSVSGYCVPGTLTALIGESGAGKTTLLNTLAQRNV-GTITGDMLVDGLP-MDASFKRR 804
Query: 351 SGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
+GY + D+H +TV ESL +SA +R P +
Sbjct: 805 TGYVQQQDLHVAELTVKESLQFSARMRRPQSI 836
Score = 52.8 bits (125), Expect = 3e-06
Identities = 39/146 (26%), Positives = 72/146 (48%), Gaps = 7/146 (4%)
Query: 279 MKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKT---GGYIDGDI 335
++E+ R ++LK VS + G + ++G GAG T+ + AG + GG G I
Sbjct: 33 IRERKNRNKMKIILKNVSLLAKSGEMVLVLGRPGAGCTSFLKSAAGETSQFAGGVTTGHI 92
Query: 336 KVSGYPKKQ--ETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNTTKVSEHV 393
G P+K+ + + Y D+H PH+TV ++L ++ ++P+ +N TK E++
Sbjct: 93 SYDGIPQKEMMQHYKPDVIYNGEQDVHFPHLTVKQTLDFAISCKMPAKRVNNVTK-EEYI 151
Query: 394 LYSEWIWLHLLTLLNEHD-RGGGRFI 418
+ + + L + D + G FI
Sbjct: 152 TANREFYAKIFGLTHTFDTKVGNDFI 177
>CDR4_CANAL (O74676) ABC transporter CDR4
Length = 1490
Score = 76.6 bits (187), Expect = 2e-13
Identities = 51/148 (34%), Positives = 77/148 (51%), Gaps = 12/148 (8%)
Query: 238 SGRADSVT---ESSHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKG 294
SG + VT ++ + + +L + + D+ Y V + EDR V+L
Sbjct: 817 SGEIEKVTPEFDNEYENNQDKMLQSGGDTFFWRDLTYQVKIK--------SEDR-VILDH 867
Query: 295 VSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARISGYC 354
VSG +PG +TALMG SGAGKTTL++ L+ R T G + I++ +F R GY
Sbjct: 868 VSGWVKPGQVTALMGASGAGKTTLLNALSDRLTTGVVTEGIRLVNGRPLDSSFQRSIGYV 927
Query: 355 EHNDIHSPHVTVHESLLYSAWLRLPSGV 382
+ D+H TV E+L ++A+LR P V
Sbjct: 928 QQQDLHLETSTVREALEFAAYLRQPKSV 955
Score = 47.8 bits (112), Expect = 8e-05
Identities = 29/93 (31%), Positives = 51/93 (54%), Gaps = 4/93 (4%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKV---SGYPKK-QET 346
+LK + G +PG LT ++G GAG +T + +A + G +ID D + S P + ++
Sbjct: 172 ILKPMDGLIKPGELTVVLGRPGAGCSTFLKTIASQTYGYHIDKDSVIRYNSLTPHEIKKH 231
Query: 347 FARISGYCEHNDIHSPHVTVHESLLYSAWLRLP 379
+ YC + H P +TV ++L ++A +R P
Sbjct: 232 YRGEVVYCAETENHFPQLTVGDTLEFAAKMRTP 264
>CDR2_CANAL (P78595) Multidrug resistance protein CDR2
Length = 1499
Score = 75.9 bits (185), Expect = 3e-13
Identities = 48/111 (43%), Positives = 70/111 (62%), Gaps = 5/111 (4%)
Query: 268 DIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKT 327
+I + D+ ++K + +EDR V+L V G +PG +TALMG SGAGKTTL++ L+ R T
Sbjct: 855 EIFFWRDLTYQVKIK--KEDR-VILDHVDGWVKPGQITALMGASGAGKTTLLNCLSERVT 911
Query: 328 GGYI-DGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLR 377
G I DG+ V+G+ +F R GY + D+H TV E+L +SA+LR
Sbjct: 912 TGIITDGERLVNGH-ALDSSFQRSIGYVQQQDVHLETTTVREALQFSAYLR 961
Score = 48.5 bits (114), Expect = 5e-05
Identities = 35/104 (33%), Positives = 52/104 (49%), Gaps = 8/104 (7%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI--DGDIKVSGY-PKKQETF 347
+LK + RPG LT ++G GAG +TL+ +A G +I + I G P E
Sbjct: 167 ILKSMDAIMRPGELTVVLGRPGAGCSTLLKTIAVNTYGFHIGKESQITYDGLSPHDIERH 226
Query: 348 ARISG-YCEHNDIHSPHVTVHESLLYSAWLRLP----SGVDSNT 386
R Y D+H PH++V ++L ++A LR P G+D T
Sbjct: 227 YRGDVIYSAETDVHFPHLSVGDTLEFAARLRTPQNRGEGIDRET 270
Score = 43.1 bits (100), Expect = 0.002
Identities = 29/105 (27%), Positives = 46/105 (43%), Gaps = 3/105 (2%)
Query: 1 MALIAMTLFFRTEMHRNDQDDAGVYAGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDL 60
M LI ++FF R D GALFF+++ F+ + EI P+ K R
Sbjct: 523 MGLILASVFFNL---RKSTDTFYFRGGALFFSVLFNAFSSLLEILSLYEARPIVEKHRKY 579
Query: 61 LFYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF 105
Y A A+ S I ++PV L+ + + Y+++ G F
Sbjct: 580 ALYRPSADALASIISELPVKLLMTMSFNIVYYFMVNLRRTAGNFF 624
>CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3
Length = 1501
Score = 75.5 bits (184), Expect = 4e-13
Identities = 39/99 (39%), Positives = 58/99 (58%)
Query: 284 VREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKK 343
++ + V+L + G +PG +TALMG SGAGKTTL++ L+ R T G I ++ +
Sbjct: 851 IKSEERVILNNIDGWVKPGEVTALMGASGAGKTTLLNALSERLTTGVITSGTRMVNGGEL 910
Query: 344 QETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
+F R GY + D+H TV E+L +SA LR P+ V
Sbjct: 911 DSSFQRSIGYVQQQDLHLETSTVREALKFSARLRQPNSV 949
Score = 48.9 bits (115), Expect = 4e-05
Identities = 28/93 (30%), Positives = 50/93 (53%), Gaps = 4/93 (4%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYI-DGDIKVSGYPKKQETFAR 349
+LK + G +PG +T ++G GAG +T + +A R G ++ DG + + E
Sbjct: 160 ILKPMEGLIKPGEVTVVLGRPGAGCSTFLKTIACRTEGFHVADGSVISYDGITQDEIRNH 219
Query: 350 ISG---YCEHNDIHSPHVTVHESLLYSAWLRLP 379
+ G YC + H P++TV E+L ++A ++ P
Sbjct: 220 LRGEVVYCAETETHFPNLTVGETLEFAALMKTP 252
Score = 35.0 bits (79), Expect = 0.56
Identities = 23/104 (22%), Positives = 44/104 (42%), Gaps = 2/104 (1%)
Query: 4 IAMTLFFRTEMHRNDQDDAGVY--AGALFFTLVTMMFNGMSEISMTIAKLPVYYKQRDLL 61
IAM L + + + + Y +++ L+ ++ + EI + K R+
Sbjct: 514 IAMALILSSVFYNLQPNSSSFYYRTSVMYYALLFNAYSSVLEIYNMYEGRAIVQKHREYA 573
Query: 62 FYPSWAYAIPSWILKIPVSLMEVSLWVFLTYYVIGFDPNVGRMF 105
YP A AI S I P+ ++ L+ + Y+++ F G F
Sbjct: 574 LYPPMADAIGSIISDFPLKVVCSVLFNLILYFMVNFKREPGAFF 617
>ABG4_HUMAN (Q9H172) ATP-binding cassette, sub-family G, member 4
Length = 646
Score = 74.3 bits (181), Expect = 8e-13
Identities = 55/161 (34%), Positives = 87/161 (53%), Gaps = 14/161 (8%)
Query: 234 TTYCSGRADSVTES---SHGKKKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLV 290
TT+ + +TE+ SH K+ V I F ++ YSV +++G +
Sbjct: 34 TTHLKKVENHITEAQRFSHLPKRSAV------DIEFVELSYSVREGPCWRKRGYK----T 83
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARI 350
LLK +SG F L +MG SGAGK+T M++LAG + G + G I V+G P++ TF ++
Sbjct: 84 LLKCLSGKFCRRELIGIMGPSGAGKSTFMNILAGYRESG-MKGQILVNGRPRELRTFRKM 142
Query: 351 SGYCEHNDIHSPHVTVHESLLYSAWLRLPSGVDSNTTKVSE 391
S Y +D+ PH+TV E+++ SA L+L + V+E
Sbjct: 143 SCYIMQDDMLLPHLTVLEAMMVSANLKLSEKQEVKKELVTE 183
>PDRC_YEAST (Q02785) ATP-dependent permease PDR12
Length = 1511
Score = 71.6 bits (174), Expect = 5e-12
Identities = 44/92 (47%), Positives = 55/92 (58%), Gaps = 1/92 (1%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQETFARI 350
LL V G +PG +TALMG SGAGKTTL++VLA R G I GD+ V+ P +F R
Sbjct: 860 LLSDVFGYVKPGKMTALMGESGAGKTTLLNVLAQRINMGVITGDMLVNAKP-LPASFNRS 918
Query: 351 SGYCEHNDIHSPHVTVHESLLYSAWLRLPSGV 382
GY D H ++V ESL ++A LR S V
Sbjct: 919 CGYVAQADNHMAELSVRESLRFAAELRQQSSV 950
Score = 37.0 bits (84), Expect = 0.15
Identities = 27/104 (25%), Positives = 50/104 (47%), Gaps = 8/104 (7%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYID--GDIKVSGYPKKQETFA 348
+++ +G G + ++G GAG +T + L+G +T +D G+ G + E +
Sbjct: 163 IIQNCTGVVESGEMLFVVGRPGAGCSTFLKCLSG-ETSELVDVQGEFSYDGLDQS-EMMS 220
Query: 349 RISGY---CEHNDIHSPHVTVHESLLYSAWLRLPS-GVDSNTTK 388
+ GY C D H P +TV E++ ++ + P +D T K
Sbjct: 221 KYKGYVIYCPELDFHFPKITVKETIDFALKCKTPRVRIDKMTRK 264
>YQ5C_CAEEL (Q09466) Putative ABC transporter C16C10.12 in
chromosome III
Length = 610
Score = 69.3 bits (168), Expect = 3e-11
Identities = 47/132 (35%), Positives = 75/132 (56%), Gaps = 8/132 (6%)
Query: 261 PHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMD 320
P ++T+++I V+ +++ + + LL VSG +PG + ALMG SGAGKTTLM+
Sbjct: 32 PITVTWENI------EVKTRKKLFSKKQKQLLNRVSGIAKPGEMVALMGASGAGKTTLMN 85
Query: 321 VLAGRKTGGY-IDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLP 379
VL R G +G +KV+G K + + ISG+ + +I P +TV E L+ A LR+
Sbjct: 86 VLMCRNMKGLEKNGTVKVNG-TKIGKEISLISGFAQQQEIFIPTLTVDEYLMIQARLRMK 144
Query: 380 SGVDSNTTKVSE 391
+ + +V E
Sbjct: 145 ANKHTRRERVDE 156
>PDRB_YEAST (P40550) ATP-dependent permease PDR11
Length = 1410
Score = 65.5 bits (158), Expect = 4e-10
Identities = 44/127 (34%), Positives = 68/127 (52%), Gaps = 15/127 (11%)
Query: 252 KKGMVLPFEPHSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVS 311
K+ + + + H I++ +I Y++ G ++ L+ SG G LTALMG S
Sbjct: 738 KESVAMETQKHVISWKNINYTI---------GDKK----LINDASGYISSG-LTALMGES 783
Query: 312 GAGKTTLMDVLAGRKTGGYIDGDIKVSGYPKKQ-ETFARISGYCEHNDIHSPHVTVHESL 370
GAGKTTL++VL+ R G + G++ + G P + F R G+ + D+H +TV ESL
Sbjct: 784 GAGKTTLLNVLSQRTESGVVTGELLIDGQPLTNIDAFRRSIGFVQQQDVHLELLTVRESL 843
Query: 371 LYSAWLR 377
S LR
Sbjct: 844 EISCVLR 850
>AUS1_YEAST (Q08409) ATP-dependent permease AUS1
Length = 1394
Score = 65.5 bits (158), Expect = 4e-10
Identities = 38/88 (43%), Positives = 53/88 (60%), Gaps = 2/88 (2%)
Query: 291 LLKGVSGAFRPGVLTALMGVSGAGKTTLMDVLAGRKTGGYIDGDIKVSGYP-KKQETFAR 349
L+ SG G LTALMG SGAGKTTL++VL+ R G + G+I + G+P ++ F R
Sbjct: 765 LINNASGFISSG-LTALMGESGAGKTTLLNVLSQRVETGVVSGEILIDGHPLTDEDAFKR 823
Query: 350 ISGYCEHNDIHSPHVTVHESLLYSAWLR 377
G+ + D+H ++V ESL S LR
Sbjct: 824 SIGFVQQQDLHLDLLSVKESLEISCLLR 851
Score = 34.3 bits (77), Expect = 0.95
Identities = 16/76 (21%), Positives = 34/76 (44%)
Query: 30 FFTLVTMMFNGMSEISMTIAKLPVYYKQRDLLFYPSWAYAIPSWILKIPVSLMEVSLWVF 89
FF+++ F ++++ + + PV KQ L FY +W + + + L V ++
Sbjct: 425 FFSILFFTFLSLADMPIAFQRQPVVKKQSQLHFYTNWVETLSTTVFDYCFKLCLVIVFSI 484
Query: 90 LTYYVIGFDPNVGRMF 105
+ Y++ R F
Sbjct: 485 ILYFLAHLQYKAARFF 500
>ABG3_MOUSE (Q99P81) ATP-binding cassette, sub-family G, member 3
Length = 650
Score = 64.3 bits (155), Expect = 9e-10
Identities = 44/132 (33%), Positives = 69/132 (51%), Gaps = 10/132 (7%)
Query: 262 HSITFDDIVYSVDMPVEMKEQGVREDRLVLLKGVSGAFRPGVLTALMGVSGAGKTTLMDV 321
H+I++ + V S P+ K + L +SG +PG L A+MG ++ L+DV
Sbjct: 40 HNISYQETVQS-GFPLRKKAYVIER-----LSNISGIMKPG-LNAIMGPQDGSRSLLLDV 92
Query: 322 LAGRKTGGYIDGDIKVSGYPKKQETFARISGYCEHNDIHSPHVTVHESLLYSAWLRLPSG 381
LA R+ + GDI ++G P + F SGY ND+ VTV ++L +SA LRLP
Sbjct: 93 LAARRDPRGLSGDILINGKP-RPANFKCTSGYVPQNDVVLGTVTVRDNLEFSAALRLPVT 151
Query: 382 V--DSNTTKVSE 391
+ D +++E
Sbjct: 152 ITRDEKRRRINE 163
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.145 0.488
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 72,508,332
Number of Sequences: 164201
Number of extensions: 3460377
Number of successful extensions: 16602
Number of sequences better than 10.0: 567
Number of HSP's better than 10.0 without gapping: 293
Number of HSP's successfully gapped in prelim test: 290
Number of HSP's that attempted gapping in prelim test: 14618
Number of HSP's gapped (non-prelim): 1382
length of query: 596
length of database: 59,974,054
effective HSP length: 116
effective length of query: 480
effective length of database: 40,926,738
effective search space: 19644834240
effective search space used: 19644834240
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC126018.1


