

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126016.2 - phase: 0
(327 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
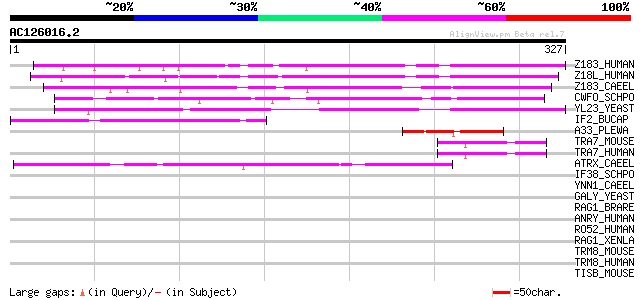
Score E
Sequences producing significant alignments: (bits) Value
Z183_HUMAN (O15541) Zinc finger protein 183 228 2e-59
Z18L_HUMAN (Q8IZP6) Zinc finger protein 183-like 1 201 2e-51
Z183_CAEEL (O17917) Putative zinc finger protein 183 homolog 191 2e-48
CWFO_SCHPO (Q9P6R8) Cell cycle control protein cwf24 157 3e-38
YL23_YEAST (P53769) Hypothetical 29.7 kDa protein in REC102-SFH1... 139 1e-32
IF2_BUCAP (Q8K9H1) Translation initiation factor IF-2 47 9e-05
A33_PLEWA (Q02084) Zinc-binding protein A33 45 2e-04
TRA7_MOUSE (Q922B6) E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.... 45 3e-04
TRA7_HUMAN (Q6Q0C0) E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.... 45 3e-04
ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog (X-li... 44 7e-04
IF38_SCHPO (O14164) Probable eukaryotic translation initiation f... 43 0.001
YNN1_CAEEL (Q03605) Hypothetical RING finger protein T02C1.1 in ... 42 0.002
GALY_YEAST (P19659) Transcription regulatory protein GAL11 42 0.003
RAG1_BRARE (O13033) V(D)J recombination activating protein 1 (RA... 41 0.004
ANRY_HUMAN (Q6UB99) Ankyrin repeat domain protein 11 (Ankyrin re... 41 0.005
RO52_HUMAN (P19474) 52 kDa Ro protein (Sjogren syndrome type A a... 40 0.008
RAG1_XENLA (Q91829) V(D)J recombination activating protein 1 (RA... 40 0.008
TRM8_MOUSE (Q99PJ2) Tripartite motif-containing protein 8 (RING ... 40 0.010
TRM8_HUMAN (Q9BZR9) Tripartite motif-containing protein 8 (RING ... 40 0.010
TISB_MOUSE (P23950) Butyrate response factor 1 (TIS11B protein) 40 0.010
>Z183_HUMAN (O15541) Zinc finger protein 183
Length = 343
Score = 228 bits (581), Expect = 2e-59
Identities = 135/330 (40%), Positives = 191/330 (56%), Gaps = 37/330 (11%)
Query: 15 EQVCSFFRKPVNRKNM---RKR-TIENEDNENDSNNEE--SLMHVQKKNTKADNKLFFST 68
+QVC+F K RK RKR + E E+ S+++E +++ +KK + + +
Sbjct: 12 DQVCTFLFKKPGRKGAAGRRKRPACDPEPGESGSSSDEGCTVVRPEKKRVTHNPMIQKTR 71
Query: 69 GSSKSSA-----SAKPNEESEKPSFH--FESSKEIQV--QHDSKATATLETETDFSRDAR 119
S K A S++ EE+E S ++S++ + D ATA E +T+ RDA+
Sbjct: 72 DSGKQKAAYGDLSSEEEEENEPESLGVVYKSTRSAKPVGPEDMGATAVYELDTEKERDAQ 131
Query: 120 AIRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGS--HGPL 177
AI ER+ K E L+GK ED K+Y+GINNY + + + T + G GP+
Sbjct: 132 AIFERSQK-IQEELRGK----EDDKIYRGINNYQKY---MKPKDTSMGNASSGMVRKGPI 183
Query: 178 RASAHIRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKAR 237
RA H+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GWQ+E+E DE
Sbjct: 184 RAPEHLRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKHGWQIERELDEG---- 239
Query: 238 KMRLATGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHA 297
R ED E S D++ +PF CFICR F +PV TKC+HYFCE CAL+H
Sbjct: 240 --RYGVYEDENYEVGS------DDEEIPFKCFICRQSFQNPVVTKCRHYFCESCALQHFR 291
Query: 298 KNKKCFVCNQPTLGIFNVAHEIRKKMAEDK 327
+C+VC+Q T G+FN A E+ K+ + +
Sbjct: 292 TTPRCYVCDQQTNGVFNPAKELIAKLEKHR 321
>Z18L_HUMAN (Q8IZP6) Zinc finger protein 183-like 1
Length = 322
Score = 201 bits (512), Expect = 2e-51
Identities = 119/321 (37%), Positives = 177/321 (55%), Gaps = 32/321 (9%)
Query: 13 QTEQVCSFFRKPVNRKN---MRKR-TIENEDNENDSNNEESLMHVQKKNTKADNKLFFST 68
Q +QVC+F K RK +RKR + E E+ S+ +E Q + S
Sbjct: 13 QADQVCTFLFKKPGRKGAAGLRKRPACDPEHGESSSSGDEGDTVAQPPRVAPRPRGLHSW 72
Query: 69 GSSKSSASAKPNEESEKPSFHF-----ESSKEIQVQHDSKATATLETETDFSRDARAIRE 123
K++ + EE+ S S+K + + D ATA E +T+ I
Sbjct: 73 --QKAAHGDRRGEEAAPESLDVVYRSTRSAKPVGPE-DMGATADFEQDTEKEHHTPTIL- 128
Query: 124 RALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSH-GPLRASAH 182
+ ++ E+L+G+ E +Y+GI++Y + ++ ++ + +G + GP+RA H
Sbjct: 129 KCSQRVQEALRGR----EHDHIYRGIHSYLRYLKP--KDTSMGNSSSGMARKGPIRAPGH 182
Query: 183 IRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLA 242
+R + R+DYQPDICKDYKETG+CG+GDSCKF+HDR DYK GW++E+E +E R
Sbjct: 183 LRATVRWDYQPDICKDYKETGFCGFGDSCKFLHDRSDYKLGWEIERELEEG------RYC 236
Query: 243 TGEDAEEEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKC 302
ED E S +E+ +PF CFICR F +PV TKC+HYFCE CAL+H +C
Sbjct: 237 ICEDENHEVGS------EEEEIPFRCFICRQAFQNPVVTKCRHYFCESCALEHFRATPRC 290
Query: 303 FVCNQPTLGIFNVAHEIRKKM 323
++C+QPT GIFN A E+ K+
Sbjct: 291 YICDQPTGGIFNPAKELMAKL 311
>Z183_CAEEL (O17917) Putative zinc finger protein 183 homolog
Length = 384
Score = 191 bits (485), Expect = 2e-48
Identities = 117/311 (37%), Positives = 168/311 (53%), Gaps = 34/311 (10%)
Query: 21 FRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNT---KADNKLFFST----GSSKS 73
FRKP R R E+ +E+ + + ++ +++ ++ +L ST SS
Sbjct: 4 FRKPKKRNAPVVRKKESSSDEDQDSEVKDVIQKRRRTNPMVQSTKQLDASTRRADNSSDD 63
Query: 74 SASAKPNEESEKPSFHFESSKEIQVQ--HDSKATATLETETDFSRDARAIRERALKQATE 131
S + N++ + F +S + D ATATLE +TD+S DA+A ER +Q E
Sbjct: 64 SDDSDDNQDIAVATHSFAASGDAGPSGPRDQGATATLEVDTDYSHDAQAQFERVQQQLKE 123
Query: 132 SLKGKSTSSEDVK-LYKGINNYTDHKAGFRREQTIASEKAGGSH--GPLRASAHIRVSAR 188
++ +D K LYKG Y +A + T A G + GP+RA +R + R
Sbjct: 124 GVE------KDGKILYKGSALYGAKEA----KDTAKGNAASGYNRVGPVRAPQFLRQTVR 173
Query: 189 FDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAE 248
+D+ PDICKDYKETG+C +GDSCKF+HDR DYK GW++++E++ G+
Sbjct: 174 WDFAPDICKDYKETGFCTFGDSCKFVHDRSDYKHGWEIDEEYE-----------AGKYGA 222
Query: 249 EEGASLNDEDDDEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQP 308
E+ A+ + D D P CFIC NPFVDP+ TKCKHYFC CALK K+ KC +C Q
Sbjct: 223 EDDANYEIHEGD-DTFPEDCFICGNPFVDPIVTKCKHYFCTGCALKSFQKSSKCPICQQN 281
Query: 309 TLGIFNVAHEI 319
T I N A E+
Sbjct: 282 TENIMNTAKEL 292
>CWFO_SCHPO (Q9P6R8) Cell cycle control protein cwf24
Length = 533
Score = 157 bits (398), Expect = 3e-38
Identities = 93/296 (31%), Positives = 155/296 (51%), Gaps = 35/296 (11%)
Query: 27 RKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADNKLFFSTGSSKSSASAKPNEESEKP 86
R+ ++++ D+++ ++E S N + + +G K+ + ESE+
Sbjct: 34 RQGLKRKKGFKRDDDSGGSSESS-------NEDMRDNIPIVSGRKKTVRLNRLQRESEQ- 85
Query: 87 SFHFESSKEIQVQHDSKATATLET--ETDFSRDARAIRERALKQATESLKGKSTSSEDVK 144
F + K+I V++ S +AT E+ T S RE L + + L +ST +
Sbjct: 86 -FENSALKDINVEYQSNLSATGESVNTTTVSAINEDTREVILGRPSPKLANQSTLPTE-- 142
Query: 145 LYKGINNYT---DHKAGFRREQTIASEKAGGSHGPLRAS--AHIRVSARFDYQPDICKDY 199
L++ N+Y+ + F ++ + GP+ +S + +R++ DYQPD+CKDY
Sbjct: 143 LFQSQNDYSRFLPKRKDFEKKSQV---------GPVLSSNASTVRMNTIIDYQPDVCKDY 193
Query: 200 KETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGEDAEEEGASLNDEDD 259
K TGYCGYGD+CKF+H R DYK+GWQ+++EWD ++ K EEG N++ +
Sbjct: 194 KLTGYCGYGDTCKFLHMREDYKAGWQLDREWDSVQEKYKKGAKL-----EEGMVKNEKKE 248
Query: 260 DEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPTLGIFNV 315
D +PF C IC+ + P++T C H+FCE CA+ + K C C T G+F+V
Sbjct: 249 D---IPFVCLICKKDYRSPIATTCGHHFCEQCAITRYRKTPTCIQCGADTKGLFSV 301
>YL23_YEAST (P53769) Hypothetical 29.7 kDa protein in REC102-SFH1
intergenic region
Length = 259
Score = 139 bits (349), Expect = 1e-32
Identities = 85/306 (27%), Positives = 142/306 (45%), Gaps = 55/306 (17%)
Query: 27 RKNMRKRTIENEDNENDSN----NEESLMHVQKKNTKADNKLFFSTGSSKSSASAKPNEE 82
RK + ++ +E N+ +EE L+ ++ +D +G+S++ + NE
Sbjct: 3 RKRLVNKSSSDEKNQKKRQKINFSEEKLVASDEEKGSSDLMSLAKSGNSRTLQLSHENEG 62
Query: 83 SEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQATESLKGKSTSSED 142
+ + V DS T E +F R A + + + + ++ + S ++
Sbjct: 63 KLQKKGEDLDKYTLTVNDDS----TKEDLLNFERKELAEKAKKRRPSDDNELVLNMSGKN 118
Query: 143 VKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIRVSARFDYQPDICKDYKET 202
+L K IN T+ IR + D+QPD+CKDYK+T
Sbjct: 119 KRLTKQINQPTN----------------------------IRTTVLMDFQPDVCKDYKQT 150
Query: 203 GYCGYGDSCKFMHDRGDYKSGWQMEKEWD-EAEKARKMRLATGEDAEEEGASLNDEDDDE 261
GYCGYGDSCKF+H R D+K+GW++ +EW+ + E ++ + L D
Sbjct: 151 GYCGYGDSCKFLHSRDDFKTGWKLNQEWNADKEDSKAVTL------------------DL 192
Query: 262 DALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPTLGIFNVAHEIRK 321
+ +PF C +C+ + PV T C HYFC C K K KCF+C++ T G VA +++K
Sbjct: 193 EKIPFKCTLCKEDYKSPVVTNCGHYFCGSCFAKDMKKGTKCFICHKETHGSAKVASDLQK 252
Query: 322 KMAEDK 327
+ + K
Sbjct: 253 MLNKRK 258
>IF2_BUCAP (Q8K9H1) Translation initiation factor IF-2
Length = 867
Score = 46.6 bits (109), Expect = 9e-05
Identities = 31/151 (20%), Positives = 70/151 (45%), Gaps = 9/151 (5%)
Query: 1 MEDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKA 60
++++ +K+ E+Q+ + + +N KN+ K T N N+N+ N KKN
Sbjct: 121 LKNTISKAKESQKNIDILEESKANINFKNLNKLTKSNVFNKNEKNKS------LKKNINF 174
Query: 61 DNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARA 120
+N F+S + K++ + + EK +H + + D++ + + +F R+ +
Sbjct: 175 NNHSFYSKKTIKNNTENQKLYKEEKKDYHLTTFIHNRNTEDNRDREIEKNKRNFHRNIKN 234
Query: 121 IRERALKQATESLKGKSTSSEDVKLYKGINN 151
R++ + +K K ++V++ K N
Sbjct: 235 YRQKKNNKQNNQIKSK---KDEVRISKNRKN 262
>A33_PLEWA (Q02084) Zinc-binding protein A33
Length = 624
Score = 45.4 bits (106), Expect = 2e-04
Identities = 22/62 (35%), Positives = 38/62 (60%), Gaps = 6/62 (9%)
Query: 232 EAEKARKMRLATGEDAEEEGASLNDEDDD--EDALPFACFICRNPFVDPVSTKCKHYFCE 289
+A ++ K ++ G D +++ ++DE+DD ED C +CR+ F +PV +C H FC+
Sbjct: 128 KAARSNKRKIEDG-DGDQKKRKVDDEEDDFTED---LTCPLCRSLFKEPVILECGHNFCK 183
Query: 290 HC 291
HC
Sbjct: 184 HC 185
>TRA7_MOUSE (Q922B6) E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.-)
(TNF receptor associated factor 7)
Length = 594
Score = 44.7 bits (104), Expect = 3e-04
Identities = 28/73 (38%), Positives = 41/73 (55%), Gaps = 13/73 (17%)
Query: 253 SLNDEDDDEDALPFA--------CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFV 304
SL +E+++ + L FA C +C + F DPV T C H FC CAL K++KC V
Sbjct: 32 SLPEEEEEPEPLVFAEQPSVKLCCQLCCSVFKDPVITTCGHTFCRRCAL----KSEKCPV 87
Query: 305 CN-QPTLGIFNVA 316
N + T+ + N+A
Sbjct: 88 DNAKLTVVVNNIA 100
>TRA7_HUMAN (Q6Q0C0) E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.-)
(TNF receptor associated factor 7) (Ring finger and WD
repeat domain 1) (RING finger protein 119)
Length = 670
Score = 44.7 bits (104), Expect = 3e-04
Identities = 28/73 (38%), Positives = 41/73 (55%), Gaps = 13/73 (17%)
Query: 253 SLNDEDDDEDALPFA--------CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFV 304
SL +E+++ + L FA C +C + F DPV T C H FC CAL K++KC V
Sbjct: 108 SLPEEEEEPEPLVFAEQPSVKLCCQLCCSVFKDPVITTCGHTFCRRCAL----KSEKCPV 163
Query: 305 CN-QPTLGIFNVA 316
N + T+ + N+A
Sbjct: 164 DNVKLTVVVNNIA 176
>ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog
(X-linked nuclear protein-1)
Length = 1359
Score = 43.5 bits (101), Expect = 7e-04
Identities = 53/262 (20%), Positives = 106/262 (40%), Gaps = 22/262 (8%)
Query: 3 DSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKADN 62
++E K Q+ ++ KP +K K+ + E+D + EES KK+ K
Sbjct: 28 ENERKEKRAQKLKEKREREGKPPPKKRPAKKRKASSSEEDDDDEEESPRKSSKKSRK--- 84
Query: 63 KLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIR 122
S+S + EE K S +S K++ + K+ T + D+ R
Sbjct: 85 -----RAKSESESDESDEEEDRKKS---KSKKKVDQKKKEKSKKKRTTSSSEDEDSDEER 136
Query: 123 ERALKQATESLKGK--STSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRAS 180
E+ K+ ++ K + S SSE+ + + + +K +++ SE++ P + S
Sbjct: 137 EQKSKKKSKKTKKQTSSESSEESEEERKVKKSKKNKEKSVKKRAETSEESDEDEKPSKKS 196
Query: 181 AH-IRVSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKM 239
++ A+ + + + +D KE + K + E E E +K K
Sbjct: 197 KKGLKKKAKSESESE-SEDEKEV-------KKSKKKSKKVVKKESESEDEAPEKKKTEKR 248
Query: 240 RLATGEDAEEEGASLNDEDDDE 261
+ + E + +DE+++E
Sbjct: 249 KRSKTSSEESSESEKSDEEEEE 270
Score = 38.9 bits (89), Expect = 0.018
Identities = 36/148 (24%), Positives = 62/148 (41%), Gaps = 13/148 (8%)
Query: 2 EDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKAD 61
E S+ K T + ++ R+ ++K +K + ++ + EE + KKN +
Sbjct: 116 EKSKKKRTTSSSEDEDSDEEREQKSKKKSKKTKKQTSSESSEESEEERKVKKSKKNKEKS 175
Query: 62 NKLFFSTGSSKSSASAKPNEESE-------KPSFHFESSKEIQVQHDSKATATLETETDF 114
K T S +S KP+++S+ K ES E +V+ K + + +
Sbjct: 176 VKKRAET-SEESDEDEKPSKKSKKGLKKKAKSESESESEDEKEVKKSKKKSKKVVKKESE 234
Query: 115 SRDARAIRERALKQATESLKGKSTSSED 142
S D E K+ TE K TSSE+
Sbjct: 235 SED-----EAPEKKKTEKRKRSKTSSEE 257
>IF38_SCHPO (O14164) Probable eukaryotic translation initiation
factor 3 93 kDa subunit (eIF3 p93)
Length = 918
Score = 42.7 bits (99), Expect = 0.001
Identities = 50/218 (22%), Positives = 98/218 (44%), Gaps = 25/218 (11%)
Query: 3 DSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIE--------NEDNENDSNNEESLMHVQ 54
DS+A+S ++ + ++ S K + + + + E +E +E +S +EES + V
Sbjct: 11 DSDAESVDSSEENRLTSSRLKKQDDSSSEEESSEEESASSSESESSEEESESEESEVEVP 70
Query: 55 KKNTKADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDF 114
KK A ++ S S+SS + E E ES E + + +S+ + E E+D
Sbjct: 71 KKKAVAASE--DSESDSESSEEEEETESEEDSEVSDESESESESESESEEESESEEESDE 128
Query: 115 SRDA-------RAIRERALKQATESLKGKST--SSEDVKLYKGINNYTDHKAGFRREQTI 165
S + + +E A + L+G+S+ SS++ + + + + D R E+ I
Sbjct: 129 SERSGPSSFLKKPEKEEAKPAGLKFLRGESSEESSDEEEGRRVVKSAKDK----RYEEFI 184
Query: 166 ASEKAGGSHGPLRASAHIRVSARFDYQPDICKDYKETG 203
+ + + ++ I VS FD+ + + KE G
Sbjct: 185 SCMET--IKNAMSSNNWIVVSNEFDHLNKVSQKCKEAG 220
Score = 32.7 bits (73), Expect = 1.3
Identities = 29/135 (21%), Positives = 62/135 (45%), Gaps = 9/135 (6%)
Query: 2 EDSEAKSTENQQTEQVCSFFRKPVNRKNMRKRTIENEDNENDSNNEESLMHVQKKNTKAD 61
E+ A S+E++ +E+ V +K +ED+E+DS + E +++ T+++
Sbjct: 44 EEESASSSESESSEEESESEESEVEVPK-KKAVAASEDSESDSESSE-----EEEETESE 97
Query: 62 NKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAI 121
S S +S + ++ ESE+ S E S E + S E E + +
Sbjct: 98 ED---SEVSDESESESESESESEEESESEEESDESERSGPSSFLKKPEKEEAKPAGLKFL 154
Query: 122 RERALKQATESLKGK 136
R + +++++ +G+
Sbjct: 155 RGESSEESSDEEEGR 169
>YNN1_CAEEL (Q03605) Hypothetical RING finger protein T02C1.1 in
chromosome III
Length = 160
Score = 42.0 bits (97), Expect = 0.002
Identities = 19/50 (38%), Positives = 26/50 (52%), Gaps = 3/50 (6%)
Query: 260 DEDALPFACFICRNPFVDPVSTKCKHYFCEHCALKHHAKNKKCFVCNQPT 309
DED F C +C + FV+P +C H +C C H N+KC +C T
Sbjct: 3 DED---FCCAVCLDFFVEPCIIECGHSYCRFCIESHLNINEKCPLCRAHT 49
>GALY_YEAST (P19659) Transcription regulatory protein GAL11
Length = 1081
Score = 41.6 bits (96), Expect = 0.003
Identities = 38/188 (20%), Positives = 83/188 (43%), Gaps = 18/188 (9%)
Query: 2 EDSEAKSTENQQTEQVCSFFRKPVNRKNMRK-RTIENEDNENDSNNEES----------- 49
E ++ ST + +Q SF RK +N+K +RK + + + N N++NN S
Sbjct: 320 EKTQLGSTMTEPVKQ--SFIRKYINQKALRKIQALRDVKNNNNANNNGSNLQRAQNVPMN 377
Query: 50 -LMHVQKKNTKADNKLFFSTGSSKSSASAKPNEESEKPSFHFESSKEIQVQHDSKATATL 108
+ Q++NT ++ + S + ++ S + N S+ + + Q Q ++A A
Sbjct: 378 IIQQQQQQNTNNNDTIATSATPNAAAFSQQQNASSKLYQMQQQQQAQAQAQAQAQAQAQA 437
Query: 109 ETETDFSRDARA-IRERALKQATESLKGKSTSSEDVKLYKGINNYTDHKAGFRREQTIAS 167
+ + ++ A+A + +A QA + ++ + + + H+ + +Q A
Sbjct: 438 QAQAQAAQAAQAQAQAQAQAQAQAQAQAQAQAQAQAQAQAQAQAHAQHQPSQQPQQ--AQ 495
Query: 168 EKAGGSHG 175
++ HG
Sbjct: 496 QQPNPLHG 503
>RAG1_BRARE (O13033) V(D)J recombination activating protein 1
(RAG-1)
Length = 1057
Score = 41.2 bits (95), Expect = 0.004
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 1/41 (2%)
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKH-HAKNKKCFVCNQ 307
C +C + DPV + C+H FC C +++ HA C CNQ
Sbjct: 296 CQVCDHLLSDPVQSPCRHLFCRLCIIRYTHALGPNCPTCNQ 336
>ANRY_HUMAN (Q6UB99) Ankyrin repeat domain protein 11 (Ankyrin
repeat-containing cofactor-1)
Length = 2664
Score = 40.8 bits (94), Expect = 0.005
Identities = 61/271 (22%), Positives = 102/271 (37%), Gaps = 55/271 (20%)
Query: 40 NENDSNNE-ESLMHVQKKNTKADNKLFFSTGSSKSSASAKPNEESEKPSF---------H 89
N SN + E+ + V T + L T +S +S + +EE + PSF +
Sbjct: 260 NPQQSNRKGETPLKVANSPTMVNLLLGKGTYTSSEESSTESSEEEDAPSFAPSSSVDGNN 319
Query: 90 FESSKEIQVQHDS------KATATLETETDFSRDARAIR----------ERALKQATESL 133
+S E ++H + KATA ++ E +F D R ++ ++ T+S
Sbjct: 320 TDSEFEKGLKHKAKNPEPQKATAPVKDEYEFDEDDEQDRVPPVDDKHLLKKDYRKETKSN 379
Query: 134 KGKSTSSEDVKLYKGINNYTDHKAGFRREQTIASEKAG----GSHGPLRASAHIRVSARF 189
S +VK Y N KA R + E+ G+ LR SAH +
Sbjct: 380 SFISIPKMEVKSYTKNNTIAPKKASHRILSDTSDEEDASVTVGTGEKLRLSAHTILPGSK 439
Query: 190 DYQPDICKDYKETGYCGYGDSCKFMHDRGDYKSGWQMEKEWDEAEKARKMRLATGED--- 246
+P K KE +++K+ + K R++R D
Sbjct: 440 TREPSNAKQQKEKN---------------------KVKKKRKKETKGREVRFGKRSDKFC 478
Query: 247 -AEEEGASLNDEDDDEDALPFACFICRNPFV 276
+E E S +DD D+L + + +P V
Sbjct: 479 SSESESESSESGEDDRDSLGSSGCLKGSPLV 509
Score = 30.0 bits (66), Expect = 8.3
Identities = 22/68 (32%), Positives = 32/68 (46%), Gaps = 1/68 (1%)
Query: 81 EESEKPSFHFESSKEIQVQHDSKATATLETETDFSRDARAIRERALKQATE-SLKGKSTS 139
++ EK S KE + + +T D SR R ++R+ K E SLK KS
Sbjct: 696 KKEEKLSKMKLEEKEWLFKDEKSLKRIKDTNKDISRSFREEKDRSNKAEKERSLKEKSPK 755
Query: 140 SEDVKLYK 147
E ++LYK
Sbjct: 756 EEKLRLYK 763
>RO52_HUMAN (P19474) 52 kDa Ro protein (Sjogren syndrome type A
antigen) (SS-A) (Ro(SS-A)) (52 kDa ribonucleoprotein
autoantigen Ro/SS-A) (Tripartite motif-containing
protein 21)
Length = 475
Score = 40.0 bits (92), Expect = 0.008
Identities = 18/41 (43%), Positives = 23/41 (55%), Gaps = 1/41 (2%)
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALK-HHAKNKKCFVCNQ 307
C IC +PFV+PVS +C H FC+ C + C VC Q
Sbjct: 16 CPICLDPFVEPVSIECGHSFCQECISQVGKGGGSVCPVCRQ 56
>RAG1_XENLA (Q91829) V(D)J recombination activating protein 1
(RAG-1)
Length = 1045
Score = 40.0 bits (92), Expect = 0.008
Identities = 14/29 (48%), Positives = 18/29 (61%)
Query: 267 ACFICRNPFVDPVSTKCKHYFCEHCALKH 295
+C +C + DPV T CKH FC C LK+
Sbjct: 293 SCLVCEHILSDPVQTSCKHLFCRICILKY 321
>TRM8_MOUSE (Q99PJ2) Tripartite motif-containing protein 8 (RING
finger protein 27) (Glioblastoma-expressed RING finger
protein)
Length = 551
Score = 39.7 bits (91), Expect = 0.010
Identities = 19/43 (44%), Positives = 24/43 (55%), Gaps = 3/43 (6%)
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNK---KCFVCNQ 307
C IC + FV+PV CKH FC C + AK+ +C CNQ
Sbjct: 15 CPICLHVFVEPVQLPCKHNFCRGCIGEAWAKDSGLVRCPECNQ 57
>TRM8_HUMAN (Q9BZR9) Tripartite motif-containing protein 8 (RING
finger protein 27) (Glioblastoma-expressed RING finger
protein)
Length = 551
Score = 39.7 bits (91), Expect = 0.010
Identities = 19/43 (44%), Positives = 24/43 (55%), Gaps = 3/43 (6%)
Query: 268 CFICRNPFVDPVSTKCKHYFCEHCALKHHAKNK---KCFVCNQ 307
C IC + FV+PV CKH FC C + AK+ +C CNQ
Sbjct: 15 CPICLHVFVEPVQLPCKHNFCRGCIGEAWAKDSGLVRCPECNQ 57
>TISB_MOUSE (P23950) Butyrate response factor 1 (TIS11B protein)
Length = 338
Score = 39.7 bits (91), Expect = 0.010
Identities = 28/98 (28%), Positives = 46/98 (46%), Gaps = 20/98 (20%)
Query: 128 QATESLKGK---STSSEDVKLYKGINNYTDHKAGFRREQTIASEKAGGSHGPLRASAHIR 184
Q SLKG+ S SS D + + D E+ + ++K GS G + +S
Sbjct: 66 QLLSSLKGEPAPSLSSRD-------SRFRDRSFSEGGERLLPTQKQPGS-GQVNSSR--- 114
Query: 185 VSARFDYQPDICKDYKETGYCGYGDSCKFMHDRGDYKS 222
Y+ ++C+ ++E G C YGD C+F H + +S
Sbjct: 115 ------YKTELCRPFEENGACKYGDKCQFAHGIHELRS 146
Score = 32.3 bits (72), Expect = 1.7
Identities = 9/26 (34%), Positives = 17/26 (64%)
Query: 191 YQPDICKDYKETGYCGYGDSCKFMHD 216
Y+ ++C+ + G+C YG C F+H+
Sbjct: 153 YKTELCRTFHTIGFCPYGPRCHFIHN 178
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.127 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,309,927
Number of Sequences: 164201
Number of extensions: 1620898
Number of successful extensions: 7792
Number of sequences better than 10.0: 347
Number of HSP's better than 10.0 without gapping: 126
Number of HSP's successfully gapped in prelim test: 225
Number of HSP's that attempted gapping in prelim test: 7204
Number of HSP's gapped (non-prelim): 692
length of query: 327
length of database: 59,974,054
effective HSP length: 110
effective length of query: 217
effective length of database: 41,911,944
effective search space: 9094891848
effective search space used: 9094891848
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC126016.2


