

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126015.12 - phase: 0
(1620 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
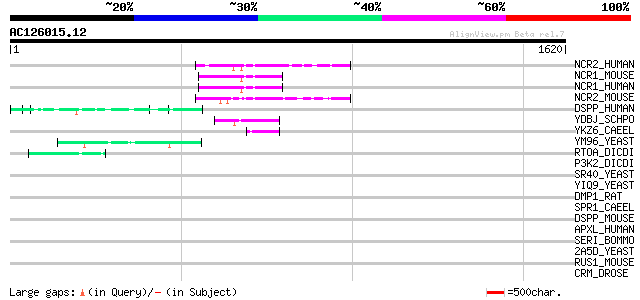
Sequences producing significant alignments: (bits) Value
NCR2_HUMAN (Q9Y618) Nuclear receptor corepressor 2 (N-CoR2) (Sil... 102 9e-21
NCR1_MOUSE (Q60974) Nuclear receptor corepressor 1 (N-CoR1) (N-C... 102 9e-21
NCR1_HUMAN (O75376) Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) 102 9e-21
NCR2_MOUSE (Q9WU42) Nuclear receptor corepressor 2 (N-CoR2) (Sil... 100 2e-20
DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contai... 57 3e-07
YDBJ_SCHPO (Q10369) Hypothetical protein C22E12.19 in chromosome I 54 5e-06
YKZ6_CAEEL (P34333) Hypothetical protein C14B9.6 in chromosome III 50 4e-05
YM96_YEAST (Q04893) Hypothetical 113.1 kDa protein in PRE5-FET4 ... 47 3e-04
RTOA_DICDI (P54681) Protein rtoA (Ratio-A) 47 6e-04
P3K2_DICDI (P54674) Phosphatidylinositol 3-kinase 2 (EC 2.7.1.13... 43 0.008
SR40_YEAST (P32583) Suppressor protein SRP40 42 0.011
YIQ9_YEAST (P40442) Hypothetical 99.7 kDa protein in SDL1 5'regi... 42 0.018
DMP1_RAT (P98193) Dentin matrix acidic phosphoprotein 1 precurso... 42 0.018
SPR1_CAEEL (Q18919) Suppressor of presenilin 1 41 0.023
DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin... 41 0.031
APXL_HUMAN (Q13796) Apical-like protein (APXL protein) 40 0.052
SERI_BOMMO (P07856) Sericin precursor (Silk gum protein) 40 0.068
2A5D_YEAST (P38903) Serine/threonine protein phosphatase 2A, 56 ... 40 0.068
RUS1_MOUSE (Q8BG26) RUN and SH3 domain containing protein 1 39 0.089
CRM_DROSE (Q8MX88) Cramped protein 39 0.089
>NCR2_HUMAN (Q9Y618) Nuclear receptor corepressor 2 (N-CoR2)
(Silencing mediator of retinoic acid and thyroid hormone
receptor) (SMRT) (SMRTe) (Thyroid-,
retinoic-acid-receptor-associated corepressor) (T3
receptor-associating factor) (TRAC) (CTG repeat pr
Length = 2517
Score = 102 bits (254), Expect = 9e-21
Identities = 104/480 (21%), Positives = 201/480 (41%), Gaps = 68/480 (14%)
Query: 541 SSYGDTYKSIIASNKESANRAHKLFTKLVPKECKKHGNMGVSNDSFSHTSILQKFAEKKQ 600
S + + I N++ A AH++ L P+ V ++ S +++ E +
Sbjct: 221 SKHRSLVQIIYDENRKKAEAAHRILEGLGPQ---------VELPLYNQPSDTRQYHENIK 271
Query: 601 FERFKERVIALKFKALHHL---WKEDM--RLLSIRKCRPKSHKKNELNVRTTCSSN---- 651
+ + + L FK +H WK+ R + + K ++ E N R +
Sbjct: 272 INQAMRKKLILYFKRRNHARKQWKQKFCQRYDQLMEALEKKVERIENNPRRRAKESKVRE 331
Query: 652 --------MKNRSSIRSRFTFPAGNHLSLVP---------TTEIINFTSKLLSESQAQLQ 694
++ + ++ R G S + +EII+ S+ E+ +
Sbjct: 332 YYEKQFPEIRKQRELQERMQSRVGQRGSGLSMSAARSEHEVSEIIDGLSE--QENLEKQM 389
Query: 695 RNTLKMPALILDEKEKMVTKFISSNGLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFG 754
R +P ++ D ++ + KFI+ NGL+ DP+ + K+R ++N W+ +EKE F EKF
Sbjct: 390 RQLAVIPPMLYDADQQRI-KFINMNGLMADPMKVYKDRQVMNMWSEQEKETFREKFMQHP 448
Query: 755 KDFRKIASFLDHKTTADCIEFYYKNHKSECFEKLKRKDIGKLGKSYAAKTNLMASGKKWN 814
K+F IASFL+ KT A+C+ +YY K+E ++ L R+ + GKS + ++
Sbjct: 449 KNFGLIASFLERKTVAECVLYYYLTKKNENYKSLVRRSYRRRGKSQQQQQQQQQQQQQQQ 508
Query: 815 HEVNVSSLDILSAASVMADVIAGNKRMRGRRYLLGYGNVKASRGEDSIIERSNSFDTLGD 874
+ + ++ D K N K ED + E+++ DT G+
Sbjct: 509 QQ------PMPRSSQEEKDEKEKEKEAEKEEEKPEVENDK----EDLLKEKTD--DTSGE 556
Query: 875 ERETAAAADVLAGICGSFSSEAMSSCITSSIDPVDGNKETKFLKANPLFKQPLTPDISQN 934
+ + A A T++ + T+ + ++ +TP Q+
Sbjct: 557 DNDEKEAV-------------ASKGRKTANSQGRRKGRITRSMANEANSEEAITP--QQS 601
Query: 935 ADDETCSDESCGEATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRK 994
A+ + E++ WT++E + + G+++ I+R VG+K CK F+ +K
Sbjct: 602 AE---LASMELNESSRWTEEEMETAKKGLLEHGRNWSAIARMVGSKTVSQCKNFYFNYKK 658
Score = 34.3 bits (77), Expect = 2.9
Identities = 17/66 (25%), Positives = 32/66 (47%)
Query: 738 WTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSECFEKLKRKDIGKLG 797
WT EE E + G+++ IA + KT + C FY+ K + +++ ++ K+
Sbjct: 615 WTEEEMETAKKGLLEHGRNWSAIARMVGSKTVSQCKNFYFNYKKRQNLDEILQQHKLKME 674
Query: 798 KSYAAK 803
K A+
Sbjct: 675 KERNAR 680
>NCR1_MOUSE (Q60974) Nuclear receptor corepressor 1 (N-CoR1) (N-CoR)
(Retinoid X receptor interacting protein 13) (RIP13)
Length = 2453
Score = 102 bits (254), Expect = 9e-21
Identities = 71/262 (27%), Positives = 128/262 (48%), Gaps = 19/262 (7%)
Query: 550 IIASNKESANRAHKLFTKLVPKE----CKKHGNMGVSNDSFSHTSILQK-----FAEKKQ 600
I N++ A AHK+F L PK + + V +++ +++K F +
Sbjct: 239 IYDENRKKAEEAHKIFEGLGPKVELPLYNQPSDTKVYHENIKTNQVMRKKLILFFKRRNH 298
Query: 601 FERFKERVIALKFKALHHLWKEDM-RLLSIRKCRPKSHKKNELNVRTTCSSNMKNRSSIR 659
+ +E+ I ++ L W++ + R+ + + + K K E + + R
Sbjct: 299 ARKQREQKICQRYDQLMEAWEKKVDRIENNPRRKAKESKTREYYEKQFPEIRKQREQQER 358
Query: 660 SRFTFPAGNHLSLV------PTTEIINFTSKLLSESQAQLQRNTLKMPALILDEKEKMVT 713
+ G LS +EII+ S+ E+ + R +P ++ D +++ V
Sbjct: 359 FQRVGQRGAGLSATIARSEHEISEIIDGLSE--QENNEKQMRQLSVIPPMMFDAEQRRV- 415
Query: 714 KFISSNGLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCI 773
KFI+ NGL+EDP+ + K+R +N WT EKE+F +KF K+F IAS+L+ K+ DC+
Sbjct: 416 KFINMNGLMEDPMKVYKDRQFMNVWTDHEKEIFKDKFIQHPKNFGLIASYLERKSVPDCV 475
Query: 774 EFYYKNHKSECFEKLKRKDIGK 795
+YY K+E ++ L R++ GK
Sbjct: 476 LYYYLTKKNENYKALVRRNYGK 497
Score = 38.1 bits (87), Expect = 0.20
Identities = 34/131 (25%), Positives = 58/131 (43%), Gaps = 6/131 (4%)
Query: 947 EATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFF--SKTRKCLGLNLANPV 1004
E + WT++E + + G+++ I++ VGTK++ CK F+ K R L L
Sbjct: 623 ETSRWTEEEMEVAKKGLVEHGRNWAAIAKMVGTKSEAQCKNFYFNYKRRHNLDNLLQQHK 682
Query: 1005 PGINGSPLND-DANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTFHDESNPL 1063
+ P + D + ES A V A D + S + + + A N+ ES P
Sbjct: 683 QKASRKPREERDVSQCES---VASTVSAQEDEDIEASNEEENPEDSEGAENSSDTESAPS 739
Query: 1064 EATSLSAKLNE 1074
+ +AK +E
Sbjct: 740 PSPVEAAKSSE 750
Score = 32.7 bits (73), Expect = 8.3
Identities = 18/57 (31%), Positives = 28/57 (48%), Gaps = 1/57 (1%)
Query: 725 PLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
P I E + WT EE E+ + G+++ IA + K+ A C FY+ N+K
Sbjct: 614 PEPISTEPVETSRWTEEEMEVAKKGLVEHGRNWAAIAKMVGTKSEAQCKNFYF-NYK 669
>NCR1_HUMAN (O75376) Nuclear receptor corepressor 1 (N-CoR1) (N-CoR)
Length = 2440
Score = 102 bits (254), Expect = 9e-21
Identities = 71/262 (27%), Positives = 128/262 (48%), Gaps = 19/262 (7%)
Query: 550 IIASNKESANRAHKLFTKLVPKE----CKKHGNMGVSNDSFSHTSILQK-----FAEKKQ 600
I N++ A AHK+F L PK + + V +++ +++K F +
Sbjct: 239 IYDENRKKAEEAHKIFEGLGPKVELPLYNQPSDTKVYHENIKTNQVMRKKLILFFKRRNH 298
Query: 601 FERFKERVIALKFKALHHLWKEDM-RLLSIRKCRPKSHKKNELNVRTTCSSNMKNRSSIR 659
+ +E+ I ++ L W++ + R+ + + + K K E + + R
Sbjct: 299 ARKQREQKICQRYDQLMEAWEKKVDRIENNPRRKAKESKTREYYEKQFPEIRKQREQQER 358
Query: 660 SRFTFPAGNHLSLV------PTTEIINFTSKLLSESQAQLQRNTLKMPALILDEKEKMVT 713
+ G LS +EII+ S+ E+ + R +P ++ D +++ V
Sbjct: 359 FQRVGQRGAGLSATIARSEHEISEIIDGLSE--QENNEKQMRQLSVIPPMMFDAEQRRV- 415
Query: 714 KFISSNGLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCI 773
KFI+ NGL+EDP+ + K+R +N WT EKE+F +KF K+F IAS+L+ K+ DC+
Sbjct: 416 KFINMNGLMEDPMKVYKDRQFMNVWTDHEKEIFKDKFIQHPKNFGLIASYLERKSVPDCV 475
Query: 774 EFYYKNHKSECFEKLKRKDIGK 795
+YY K+E ++ L R++ GK
Sbjct: 476 LYYYLTKKNENYKALVRRNYGK 497
Score = 36.6 bits (83), Expect = 0.58
Identities = 13/43 (30%), Positives = 26/43 (60%)
Query: 947 EATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFF 989
E + WT++E + + G+++ I++ VGTK++ CK F+
Sbjct: 624 ETSRWTEEEMEVAKKGLVEHGRNWAAIAKMVGTKSEAQCKNFY 666
Score = 34.3 bits (77), Expect = 2.9
Identities = 29/110 (26%), Positives = 49/110 (44%), Gaps = 14/110 (12%)
Query: 725 PLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSEC 784
P I E + WT EE E+ + G+++ IA + K+ A C FY+ N+K
Sbjct: 615 PEPISTEPVETSRWTEEEMEVAKKGLVEHGRNWAAIAKMVGTKSEAQCKNFYF-NYK--- 670
Query: 785 FEKLKRKDIGKLGKSYAAKTNLMASGKKWNHEVNVSSLD-ILSAASVMAD 833
+R ++ L + + KT+ +K E +VS + + S S D
Sbjct: 671 ----RRHNLDNLLQQHKQKTS-----RKPREERDVSQCESVASTVSAQED 711
>NCR2_MOUSE (Q9WU42) Nuclear receptor corepressor 2 (N-CoR2)
(Silencing mediator of retinoic acid and thyroid hormone
receptor) (SMRT) (SMRTe) (Thyroid-,
retinoic-acid-receptor-associated corepressor) (T3
receptor-associating factor) (TRAC)
Length = 2472
Score = 100 bits (250), Expect = 2e-20
Identities = 111/481 (23%), Positives = 196/481 (40%), Gaps = 74/481 (15%)
Query: 541 SSYGDTYKSIIASNKESANRAHKLFTKLVPKECKKHGNMGVSNDSFSHTSILQKFAEKKQ 600
S + + I N++ A AH++ L P+ N + + + KK
Sbjct: 221 SKHRSLVQIIYDENRKKAEAAHRILEGLGPQVELPLYNQPSDTRQYHENIKINQAMRKKL 280
Query: 601 FERFKERVIALK---------FKALHHLWKEDMRLLSIRKCR---------------PKS 636
FK R A K + L W++ + + R P+
Sbjct: 281 ILYFKRRNHARKQWEQRFCQRYDQLMEAWEKKVERIENNPRRRAKESKVREYYEKQFPEI 340
Query: 637 HKKNELNVRTTCSSNMKNRSSIRSRFTFPAGNHLSLVPTTEIINFTSKLLSESQAQLQRN 696
K+ EL R S + R S S + + +S EII+ S+ E+ + R
Sbjct: 341 RKQRELQERM--QSRVGQRGSGLSMSAARSEHEVS-----EIIDGLSE--QENLEKQMRQ 391
Query: 697 TLKMPALILDEKEKMVTKFISSNGLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKD 756
+P ++ D ++ + KFI+ NGL++DP+ + K+R + N W+ +E++ F EKF K+
Sbjct: 392 LAVIPPMLYDADQQRI-KFINMNGLMDDPMKVYKDRQVTNMWSEQERDTFREKFMQHPKN 450
Query: 757 FRKIASFLDHKTTADCIEFYYKNHKSECFEKLKRKDIGKLGKSYAAKTNLMASGKKWNHE 816
F IASFL+ KT A+C+ +YY K+E ++ L R+ + GKS + ++ +
Sbjct: 451 FGLIASFLERKTVAECVLYYYLTKKNENYKSLVRRSYRRRGKS---QQQQQQQQQQQQQQ 507
Query: 817 VNVSSLDILSAASVMADVIAGNKRMRGRRYLLGYGNVKASRGEDSIIERSNSFDTLG--- 873
+ SS + + ++ + + E + + + DT G
Sbjct: 508 MARSSQEEKEEKEKEKEADKEEEK-------------QDAENEKEELSKEKTDDTSGEDN 554
Query: 874 DERETAAAADVLAGICGSFSSEAMSSCITSSIDPVDGNKETKFLKANPLFKQPLTPDISQ 933
DE+E A+ G + S IT S+ ++ET TP S
Sbjct: 555 DEKEAVAS----KGRKTANSQGRRKGRITRSMANEANHEET------------ATPQQS- 597
Query: 934 NADDETCSDESCGEATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTR 993
E S E E++ WT++E + + G+++ I+R VG+K CK F+ +
Sbjct: 598 ---SELASME-MNESSRWTEEEMETAKKGLLEHGRNWSAIARMVGSKTVSQCKNFYFNYK 653
Query: 994 K 994
K
Sbjct: 654 K 654
Score = 34.3 bits (77), Expect = 2.9
Identities = 17/66 (25%), Positives = 32/66 (47%)
Query: 738 WTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSECFEKLKRKDIGKLG 797
WT EE E + G+++ IA + KT + C FY+ K + +++ ++ K+
Sbjct: 611 WTEEEMETAKKGLLEHGRNWSAIARMVGSKTVSQCKNFYFNYKKRQNLDEILQQHKLKME 670
Query: 798 KSYAAK 803
K A+
Sbjct: 671 KERNAR 676
>DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contains:
Dentin phosphoprotein (Dentin phosphophoryn) (DPP);
Dentin sialoprotein (DSP)]
Length = 1253
Score = 57.4 bits (137), Expect = 3e-07
Identities = 120/561 (21%), Positives = 211/561 (37%), Gaps = 91/561 (16%)
Query: 3 SEEPGHGYGVSRSGDKMLEEDGRPLVSRGDGKYGRSSR-DNRGGPFGQRDWRGHSWEASN 61
S+ GY DK ++ D +G +S DN G + N
Sbjct: 443 SDSNSDGYDSYDFDDKSMQGDDPNSSDESNGNDDANSESDNNSSSRGDASYNSDE-SKDN 501
Query: 62 GSPNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGTGPR 121
G+ + S+ +D DDS + S +SD + +N D + K+ +G G
Sbjct: 502 GNGSDSKGAED--------DDSDSTSDTNNSD--SNGNGNNGNDDNDKSD--SGKGKSDS 549
Query: 122 CDRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVTAVE 181
D ++S SS S S S S SS S S + +DS + + + ++
Sbjct: 550 SDSDSSDSS-----------NSSDSSDSSDSDSSDSNSSSDSDSSDSDSSDSSDSDSSDS 598
Query: 182 SNSGEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNM 241
SNS +++ SS S+ + S + + K + D D S D S S++ +
Sbjct: 599 SNSSDSSDSSDSSDSSDSSDS--------SDSKSDSSKSESDSSDSDSKSDSSDSNSSDS 650
Query: 242 EPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSNLTD 301
S S + S + SD + + SS + SSS +D SN +D
Sbjct: 651 SDNSDSSDS--SNSSNSSDSSDSSDSSDSSSSSDSSS--------------SSDSSNSSD 694
Query: 302 SPAPGFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDDPSSDDSGLVRSNAINKLLIW 361
S ++ + + D D DS +S SS SD +S DS
Sbjct: 695 SSDSSDSSNSSESSDSSDSSDSDSSDSSDSSNSNSSDSDSSNSSDSS------------D 742
Query: 362 KADISKVLEMTESEIDLLENELKSLKSESVDRSECPVASGSQQADSSSKFYEERVEVSQK 421
+D S ++S D ++ S S+S D S+ +S S ++ SS
Sbjct: 743 SSDSSDSSNSSDSS-DSSDSSNSSDSSDSSDSSDSSDSSNSSDSNDSSN----------- 790
Query: 422 VIRPVPLKIISSDEPNTVKMPQSTNLCSIHENDKEEDIDSPGSATSKFVEPLPVNAVSSS 481
SSD ++ S+N ++ D DS S+ S N+ SS
Sbjct: 791 ----------SSDSSDSSNSSDSSNSSDSSDSSDSSDSDSSNSSDSS-------NSSDSS 833
Query: 482 YTRGYDNLSRDMNAVQSTMMKCFVRCNRKNTSVSACNNVNTPTEVKDSLGDVTFGANLCS 541
+ + S ++ S+ R + N+S S+ ++ ++ + D + +N S
Sbjct: 834 DSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSSDSSDSSDSSNSSDSSDSSD-SSDSNESS 892
Query: 542 SYGDTYKSIIASNKESANRAH 562
+ D+ S +S+ +S++ ++
Sbjct: 893 NSSDSSDSSNSSDSDSSDSSN 913
Score = 50.1 bits (118), Expect = 5e-05
Identities = 79/360 (21%), Positives = 135/360 (36%), Gaps = 18/360 (5%)
Query: 60 SNGSPNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGTG 119
S+ S + S ++ +S DS S S N+ + + D + N +
Sbjct: 610 SSDSSDSSDSSDSKSDSSKSESDSSDSDSKSDSSDSNSSDSSDNSDSSDSSNSSNSSDSS 669
Query: 120 PRCDRENSLSSIDWKPLKWTRSGSLSSRGS---GFSHSSSSRSMAGTDSYEGKPNLKHKN 176
D +S SS D + + S SS S S SS S + +DS + + +
Sbjct: 670 DSSDSSDSSSSSDSSSSSDSSNSSDSSDSSDSSNSSESSDSSDSSDSDSSDSSDSSNSNS 729
Query: 177 VTAVESNSGEATACVTS--------SMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPG 228
+ SNS +++ S S S D+++ + + D D
Sbjct: 730 SDSDSSNSSDSSDSSDSSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSNDSS 789
Query: 229 SNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGK 288
++ D S SS + SS S + D S + S +S ++ SS + +SS D
Sbjct: 790 NSSDSSDSSNSSDSSNSSDSSDSSDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSS--DS 847
Query: 289 VGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDDPSSDDSG 348
++D+D SN +DS + + D D + S SD +S DS
Sbjct: 848 SDSSDSDSSNRSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSNESSNSSDSSDSSNSSDSD 907
Query: 349 LVRSNAINKLLIWKADISKVLEMTESEIDLLENELKSLKSESVDRSECPVASGSQQADSS 408
S+ + +D S + +ES + +N S S S D S+ +S S + +S
Sbjct: 908 SSDSSNSSD----SSDSSNSSDSSESS-NSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNS 962
Score = 47.4 bits (111), Expect = 3e-04
Identities = 96/428 (22%), Positives = 156/428 (36%), Gaps = 37/428 (8%)
Query: 38 SSRDNRGGPFGQRDWRGHSWEASNGSPNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNT 97
S NR D S ++SN S + +NE + DS S+ SD ++
Sbjct: 853 SDSSNRSDSSNSSD-SSDSSDSSNSSDSSDSSDSSDSNESSNSSDSSDSSNSSDSDSSDS 911
Query: 98 WEQHNLKDQHAKTGGVNGLGTGPRCDRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSS 157
+ D + + + +S +S D + + S SS S+SS S
Sbjct: 912 SNSSDSSDSSNSSDSSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDS 971
Query: 158 RSMAGTDSYEGKPNLKHKNVTAVESNSGEATACVTSSMPSEDATSRKKPRLNWGEGLAKY 217
+DS + N + ++ S+S +++ SS S+ + S +
Sbjct: 972 SDSNSSDSSDSS-NSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSN-------SSD 1023
Query: 218 EKKKVDVPDPGSNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSS 277
D D + D S SS + SS S + D S +G SD +S ++ SS + S
Sbjct: 1024 SSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDS-SGSSD-SSDSSDSSDSSDS 1081
Query: 278 SPGVDDKLLGKVGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKLD-VDSLNSLGSSIVEL 336
S D + +D S+ +DS + + D D DS NS SS
Sbjct: 1082 SDS-SDSSDSSDSSESSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSSD 1140
Query: 337 VQSDDPSSDDSGLVRSNAINKLLIWKADISKVLEMTESEIDLLENELKSLKSESVDRSEC 396
SSD S S+ +D S + ++S D ++ S +ES D S+
Sbjct: 1141 SSDSSDSSDSSDSSDSSD-------SSDSSDSSDSSDSS-DSSDSSDSSDSNESSDSSDS 1192
Query: 397 PVASGSQQADSSSKFYEERVEVSQKVIRPVPLKIISSDEPNTVKMPQSTNLCSIHENDKE 456
+S S + SS + S S+DE ++ + N N +
Sbjct: 1193 SDSSDSSNSSDSSDSSDSSDSTSD-----------SNDESDSQSKSGNGN-----NNGSD 1236
Query: 457 EDIDSPGS 464
D DS GS
Sbjct: 1237 SDSDSEGS 1244
Score = 43.9 bits (102), Expect = 0.004
Identities = 70/308 (22%), Positives = 109/308 (34%), Gaps = 18/308 (5%)
Query: 3 SEEPGHGYGVSRSGDKMLEEDGRPLVSRGDGKYGRSSRDNRGGPFGQRDWRGHSWEASNG 62
S + + S SGD D S SS + S +S+
Sbjct: 950 SSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDS 1009
Query: 63 SPNL----SRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGT 118
S + S D +N S D S + S SD N+ + + D + + G+
Sbjct: 1010 SDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSS---DSSGS 1066
Query: 119 GPRCDRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVT 178
D +S S D + S SS S S SS S + + + + +
Sbjct: 1067 SDSSDSSDSSDSSDSSDSSDSSDSSDSSESSDSSDSSDSSDSSDSSDSSDSSDSSDSSDS 1126
Query: 179 AVESNSGEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSA 238
+ SNS +++ SS S+ + S + D D + D S SS
Sbjct: 1127 SDSSNSSDSSDSSDSSDSSDSSDSSDSS----DSSDSSDSSDSSDSSDSSDSSDSSDSSD 1182
Query: 239 GNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSN 298
N SS S + D S S +S ++ SS + S S D K GN +N+ S+
Sbjct: 1183 SNESSDSSDSSDSSDSSN-----SSDSSDSSDSSDSTSDSNDESDSQ-SKSGNGNNNGSD 1236
Query: 299 LTDSPAPG 306
+DS + G
Sbjct: 1237 -SDSDSEG 1243
Score = 38.5 bits (88), Expect = 0.15
Identities = 83/420 (19%), Positives = 155/420 (36%), Gaps = 23/420 (5%)
Query: 1008 NGSPLNDDANGGESDTDDACVVEAGSVVDADKSGNKTDEDLPSDALNTFHDESNPLEATS 1067
N S +D ++ SD+ D+ ++ + D+ S + +D SD+ ++ D S +S
Sbjct: 575 NSSSDSDSSDSDSSDSSDSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSKSDSSKSESDSS 634
Query: 1068 LSAKLNESREISGTEVCLENVDVASVACAINVESKLGSDVSGVGLCTTDKSGSVNGVGLG 1127
S ++S + S + +N D + + + N S S ++D S S +
Sbjct: 635 DSDSKSDSSD-SNSSDSSDNSDSSDSSNSSNSSDSSDSSDSSDSSSSSDSSSSSDSSNSS 693
Query: 1128 GTVRESISASEIIKPRECGSVALDRTVSEGSSGGLCLGSEVERQRVSAPHCVVDKDVEHV 1187
+ S S++ S D + S SS S+ S+ D +
Sbjct: 694 DSSDSSDSSNSSESSDSSDSSDSDSSDSSDSSNSNSSDSDSSNSSDSSDSS-DSSDSSNS 752
Query: 1188 ADAGVVVELKNC-----VLESSTAANVSFSPVVNSCSGLSFGSENKHVSFGKPHTSALSM 1242
+D+ + N +SS +++ S S N S S S++ + S + +
Sbjct: 753 SDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSNDSSNSSDSSDSSNSSDSSNSSDSSDS 812
Query: 1243 SMSDLQATANSLLLKAAAAQCEKTVSQDRL-SSTCDIQGGRDMRCHSSGSNGDHQLPLSG 1301
S S ++NS ++ + + S D SS D S SN S
Sbjct: 813 SDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSSDSSDSSD 872
Query: 1302 SHVETVSVLQGYSMQVPIKKEVDGDVNCSSSAAEFPLLPQKVKQTDGHFKPSFHSSNSEK 1361
S + S S D N SS++++ +D S +SS+S
Sbjct: 873 SSNSSDSSDSSDS----------SDSNESSNSSD---SSDSSNSSDSDSSDSSNSSDSSD 919
Query: 1362 TSRNGDVKLFGKILTNPSSTQNPNLTAKRSEENGSHHPKLNNK--SSNLNFTGHQNSDEN 1419
+S + D N +S+ + N + + S+ +N SSN + + NS ++
Sbjct: 920 SSNSSDSSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDS 979
Score = 36.6 bits (83), Expect = 0.58
Identities = 97/503 (19%), Positives = 178/503 (35%), Gaps = 25/503 (4%)
Query: 930 DISQNADDETCSDESCGEATEWTDDETAAFLQAVSSFGKDFEKI--SRCVGTKAQEHCKR 987
D S DD SDES G ++ + + + +S+ D K + A++
Sbjct: 457 DKSMQGDDPNSSDESNGNDDANSESDNNSSSRGDASYNSDESKDNGNGSDSKGAEDDDSD 516
Query: 988 FFSKTRKCLGLNLANPVPGINGSPLND--DANGGESDTDDACVVEAGSVVDADKSGNKTD 1045
S T N + G NG+ ND D+ G+SD+ D+ ++ + D+ S +
Sbjct: 517 STSDTN-----NSDSNGNGNNGNDDNDKSDSGKGKSDSSDSDSSDSSNSSDSSDSSDSDS 571
Query: 1046 EDLPSDALNTFHDESNPLEATSLSAKLNESREISGTEVCLENVDVASVACAINVESKLGS 1105
D S + + D + + S S+ + S + S + ++ D + + + + SK S
Sbjct: 572 SDSNSSSDSDSSDSDSSDSSDSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSKSDSSKSES 631
Query: 1106 DVS-----GVGLCTTDKSGSVNGVGLGGTVRESISASEIIKPRECGSVALDRTVSEGSSG 1160
D S + S N + + S S S + D + S SS
Sbjct: 632 DSSDSDSKSDSSDSNSSDSSDNSDSSDSSNSSNSSDSSDSSDSSDSSSSSDSSSSSDSSN 691
Query: 1161 GLCLGSEVERQRVSAPHCVVDKDVEHVADAGVVVELKNCVLESSTAANVSFSPVVNSCSG 1220
+ S D +D+ + +SS +++ S S + S
Sbjct: 692 SSDSSDSSDSSNSSESSDSSDSSDSDSSDSSDSSNSNSSDSDSSNSSDSSDSSDSSDSSN 751
Query: 1221 LSFGSENKHVSFGKPHTSALSMSMSDLQATANSLLLKAAAAQCEKTVSQDRLSSTCDIQG 1280
S S++ S S+ S SD ++NS ++ + + S S++ D
Sbjct: 752 SSDSSDSSDSS--NSSDSSDSSDSSDSSDSSNSSDSNDSSNSSDSSDS----SNSSDSSN 805
Query: 1281 GRDMRCHSSGSNGDHQLPLSGSHVETVSVLQGYSMQVPIKKEVDGDVNCSSSAAEFPLLP 1340
D S S+ D S+ S S D + SS+ ++
Sbjct: 806 SSDSSDSSDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSS 865
Query: 1341 QKVKQTD----GHFKPSFHSSNSEKTSRNGDVKLFGKILTNPSSTQNPNLTAKRSEENGS 1396
+D S SS+S ++S + D ++ S+ + N + N S
Sbjct: 866 DSSDSSDSSNSSDSSDSSDSSDSNESSNSSD-SSDSSNSSDSDSSDSSNSSDSSDSSNSS 924
Query: 1397 HHPKLNNKSSNLNFTGHQNSDEN 1419
+ +N S N N + NS ++
Sbjct: 925 DSSESSNSSDNSNSSDSSNSSDS 947
Score = 35.4 bits (80), Expect = 1.3
Identities = 61/307 (19%), Positives = 115/307 (36%), Gaps = 40/307 (13%)
Query: 864 ERSNSFDTLGDERETAAAADVLAGICGSFSSEAMSSCITSSIDPVDGNKETKFLKANPLF 923
+ SNS D+ D +++ ++D S SS++ S S D D + ++ +N
Sbjct: 597 DSSNSSDS-SDSSDSSDSSD------SSDSSDSKSDSSKSESDSSDSDSKSDSSDSN--- 646
Query: 924 KQPLTPDISQNADDETCSDESCGEATEWTDDETAAFLQAVSSFGKDFEKISRCVGTKAQE 983
+ D S N+D S+ S + + D + + + SS D S +
Sbjct: 647 ----SSDSSDNSDSSDSSNSSNSSDSSDSSDSSDSSSSSDSSSSSDSSNSSDSSDSSDSS 702
Query: 984 HCKRFFSKTRKCLGLNLANPVPGINGSPLNDDANGGESDTDDACVVEAGSVVDADKSGNK 1043
+ + + S +D +N SD+D + ++ D+ S N
Sbjct: 703 NSSESSDSSDSS----------DSDSSDSSDSSNSNSSDSDSSNSSDSSDSSDSSDSSNS 752
Query: 1044 TDEDLPSDALNT-----------FHDESNPLEATSLSAKLNESREISGTEVCLENVDVAS 1092
+D SD+ N+ D SN ++ S+ ++S + S + + D +
Sbjct: 753 SDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSND-SSNSSDSSDSSNSSDSSNSSDSSD 811
Query: 1093 VACAINVESKLGSDVSGVGLCTTDKSGSVNGVGLGGTVRESISASEIIKPRECGSVALDR 1152
+ + + +S SD S ++D S S N + S S+ R S + D
Sbjct: 812 SSDSSDSDSSNSSDSSN----SSDSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSSDS 867
Query: 1153 TVSEGSS 1159
+ S SS
Sbjct: 868 SDSSDSS 874
>YDBJ_SCHPO (Q10369) Hypothetical protein C22E12.19 in chromosome I
Length = 661
Score = 53.5 bits (127), Expect = 5e-06
Identities = 52/205 (25%), Positives = 88/205 (42%), Gaps = 16/205 (7%)
Query: 598 KKQFERFKERVIALKFKALHHLWKEDMRLLSIRKCRPKSHKKNELNV-RTTCSSNMKN-- 654
+++F+R K+ ++ + K H L K+ RL S K ++N V R T KN
Sbjct: 99 REKFQRDKQTIVGVMRKRRHVLNKKIKRLQSHWKQVLGRWEENIARVDRLTEIDKTKNAK 158
Query: 655 -------RSSIRSRFTFPAGNHLSLVPTTEIINFTSKLLSE----SQAQLQRNTLKMPAL 703
RS+ + F AG+ + E + +KL + S +P +
Sbjct: 159 KSEPFIKRSTRKVMSNFTAGDIVR--SEEEFLEILAKLEQQEKEASNVSEASRIATIPPM 216
Query: 704 ILDEKEKMVTKFISSNGLVEDPLAIEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASF 763
IL E+E F + LV D +SM + W E+ +F+++F GK F KIA
Sbjct: 217 ILSEEEVKSQYFNDQSRLVTDCPKFYHFQSMPDIWNEEQHSIFVQQFILHGKKFGKIAEA 276
Query: 764 LDHKTTADCIEFYYKNHKSECFEKL 788
+ K + +C+ YY ++ + L
Sbjct: 277 VPGKNSKECVLHYYLTKRTTDYRAL 301
Score = 32.7 bits (73), Expect = 8.3
Identities = 34/182 (18%), Positives = 73/182 (39%), Gaps = 12/182 (6%)
Query: 951 WTDDETAAFLQAVSSFGKDFEKISRCVGTKAQEHCKRFFSKTRKCLGLNL----ANPVPG 1006
W +++ + F+Q GK F KI+ V K + C + T++ A G
Sbjct: 251 WNEEQHSIFVQQFILHGKKFGKIAEAVPGKNSKECVLHYYLTKRTTDYRALVASATKTKG 310
Query: 1007 INGSPLNDDANGGESDTDDACV---VEAGSVVDADKSGNK--TDEDLPSDALNTFHDESN 1061
L GG+ + + + +EA + +++ N + + +D +NT+ + +
Sbjct: 311 RRRKKLLPSQRGGKKKSKGSALMVDIEAADINKTEENINNQFQEASVTADNMNTWDNTPS 370
Query: 1062 PLEATSLSAKLNESREISGTEVCLENVDVASVACAINVESKLGSDVSGVGLCTTDKSGSV 1121
S + +N + ++++ + A I K + + + TDKS +V
Sbjct: 371 VENVESANENVNNHNADEQMDEKIKSLVEGNSAYEI---EKGAQEPDPMSIDMTDKSETV 427
Query: 1122 NG 1123
+G
Sbjct: 428 SG 429
>YKZ6_CAEEL (P34333) Hypothetical protein C14B9.6 in chromosome III
Length = 1780
Score = 50.4 bits (119), Expect = 4e-05
Identities = 29/99 (29%), Positives = 52/99 (52%), Gaps = 4/99 (4%)
Query: 692 QLQRNTLKMPALILDEKEKMVTKFISSNGLVEDPLAIEKERSMIN---PWTSEEKELFLE 748
+++ K+P L L E E+ + +F+ G + + E +S+++ W+ EE+ LF
Sbjct: 170 KIRLGVAKIPRL-LTESERKMDEFVERPGSILKDMKKEHRQSVLDRLEEWSPEERSLFKS 228
Query: 749 KFAAFGKDFRKIASFLDHKTTADCIEFYYKNHKSECFEK 787
+ A K F + F KT +D + FYY N K+E ++K
Sbjct: 229 RQADHVKIFHGLTEFFVDKTASDLVLFYYMNKKTEDYKK 267
>YM96_YEAST (Q04893) Hypothetical 113.1 kDa protein in PRE5-FET4
intergenic region
Length = 1140
Score = 47.4 bits (111), Expect = 3e-04
Identities = 89/448 (19%), Positives = 160/448 (34%), Gaps = 46/448 (10%)
Query: 141 SGSLSSRGSGFSHSS----SSRSMAGTDSYEGKPNLKHKNVTA-----VESNSGEATACV 191
SGS S+ + SHSS SS + T + + VT V S+S ++ V
Sbjct: 4 SGSKSTTATTTSHSSTTTTSSTTSTTTPTTTSTTSTTSTKVTTSPEIIVSSSSTLVSSVV 63
Query: 192 TSSMPSEDATSRKKPRLNWGEGLAK------YEKKKVDVPDPGSNKDGSVSSAGNMEPCS 245
S +S + E L Y + + + + + S
Sbjct: 64 PEFTSSSSLSSDTIASILSSESLVSIFSSLSYTSSDISSTSVNDVESSTSGPSNSYSALS 123
Query: 246 SISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSNLTDSPAP 305
S + L + + S A + + S+ G + LGK + V T S +
Sbjct: 124 STNAQLSSSTTETDSISSSAIQTSSPQTSSSNGGGSSSEPLGK-----SSVLETTASSSD 178
Query: 306 GFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDDPSSDDSGLVRSNAINKLLIWKADI 365
F D ++S GS++ + + D S + V S++ + D+
Sbjct: 179 TTAVTSSTFTTLTDVSSSPKISSSGSAVTSVGTTSDASKE----VFSSSTS-------DV 227
Query: 366 SKVLEMTESEIDLLENELKSLKSESVDRSECPVASGSQQADSSSKFYEERVEVSQKVIRP 425
S +L T S +E S + + PV+S + A SSS E S V
Sbjct: 228 SSLLSSTSSPASSTISETLPFSSTILSITSSPVSSEAPSATSSSVSSEASSSTSSSVSSE 287
Query: 426 VPL---KIISSDEPNTVKMPQSTNLCSIHENDKEEDIDSPGS---------ATSKFVEPL 473
PL ++SS+ P++ S+ S + +I S S ATS V
Sbjct: 288 APLATSSVVSSEAPSSTSSVVSSEAPSSTSSSVSSEISSTTSSSVSSEAPLATSSVVSSE 347
Query: 474 PVNAVSSSYTRGYDNLSRDMNAVQSTMMKCFVRCNRKNTSVSACNNVNTPTEVKDSLGD- 532
++ SSS + + + + ++ + V + +S S+ + P+ S+
Sbjct: 348 APSSTSSSVSSEISSTTSSSVSSEAPLATSSVVSSEAPSSTSSSVSSEAPSSTSSSVSSE 407
Query: 533 --VTFGANLCSSYGDTYKSIIASNKESA 558
+ +++ S T S+++S SA
Sbjct: 408 APSSTSSSVSSEISSTKSSVMSSEVSSA 435
>RTOA_DICDI (P54681) Protein rtoA (Ratio-A)
Length = 400
Score = 46.6 bits (109), Expect = 6e-04
Identities = 54/225 (24%), Positives = 88/225 (39%), Gaps = 7/225 (3%)
Query: 56 SWEASNGSPNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNG 115
S +++GS + S + + S S + S S N+ Q + ++ + G G
Sbjct: 89 SSSSNSGSQSTSNSGSEASGSSNSGSQSTSNSGSEASGSSNSGSQSSTDSSNSGSQGSTG 148
Query: 116 LGTGPRCDRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHK 175
+S +S + SGS S GS S S SS S G N +
Sbjct: 149 SSNSGSQSSTDSSNSGSQSSTDSSNSGSQGSTGSSNSGSESSGSSNSGSESSGSSNSGSE 208
Query: 176 NVTAVESNSGEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSV 235
+ + SNSG ++ +S+ SE ++ N G + GS+ GS
Sbjct: 209 SSSG-SSNSGSESSSGSSNSGSESSSGSS----NSGSESSSGSSNSGSESSSGSSNSGSE 263
Query: 236 SSAGNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPG 280
SS+G+ S S + + +G S+ S + SS + SSS G
Sbjct: 264 SSSGSSNSGSESSSGSSNSGSESSGSSNSGSES--SSDSGSSSDG 306
Score = 38.5 bits (88), Expect = 0.15
Identities = 53/283 (18%), Positives = 103/283 (35%), Gaps = 14/283 (4%)
Query: 59 ASNGSPNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGT 118
+S+GS + + +++ S S + S S N+ Q + + ++ G + G+
Sbjct: 74 SSSGSNSTASSEGSVSSSSNSGSQSTSNSGSEASGSSNSGSQ-STSNSGSEASGSSNSGS 132
Query: 119 GPRCDRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVT 178
D NS S +GS +S + SS+S S + TDS +
Sbjct: 133 QSSTDSSNSGSQ--------GSTGSSNSGSQSSTDSSNSGSQSSTDSSNSGSQGSTGSSN 184
Query: 179 AVESNSGEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSA 238
+ +SG + + SS S + N G + GS+ GS SS+
Sbjct: 185 SGSESSGSSNSGSESSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSS 244
Query: 239 GNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSN 298
G+ S S + + + S + + S + S S G ++D+ S+
Sbjct: 245 GSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSGSSNSGSESSSDSGSSS 304
Query: 299 LTDSPAPGFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDD 341
+ F + L+++ +D D + G + ++
Sbjct: 305 DGKTTCISFHD-----TLSINTVDDDEIECTGKGETRCISDNN 342
>P3K2_DICDI (P54674) Phosphatidylinositol 3-kinase 2 (EC 2.7.1.137)
(PI3-kinase) (PtdIns-3-kinase) (PI3K)
Length = 1858
Score = 42.7 bits (99), Expect = 0.008
Identities = 52/218 (23%), Positives = 97/218 (43%), Gaps = 12/218 (5%)
Query: 135 PLKWTRSGSLSSRGSGFSHSSS-SRSMAGTDSYEGKPNLKHKNVTAVESNSGEATACVTS 193
P K + S L S S SS+ + S++ + + V V+ +S +T ++
Sbjct: 344 PKKVSSSKLLIPPSSNVSSSSNITLSLSSSSPSSSSSSSTSTVVPIVQLSSSNSTNSPST 403
Query: 194 SMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNMEPCSSI---SPN 250
S+P+ S+ P ++ + + ++++ + SN + + +S+ + SS+ SP
Sbjct: 404 SLPTTPRLSQ--PTTSYTQLIPSQQQQQPTESNSSSNTNTTTTSSSSSSSSSSLTISSPQ 461
Query: 251 LVDKSPKVTGFSDCASPATPSSVACSSSPG-VDDKLLGKVGNADNDVSNLTDSPAPGFQN 309
+ S +++ F ++ T SS SSPG V +K+ +GN N + N
Sbjct: 462 PSNNSIRISAFGRSSTQFTISSNGIPSSPGQVSNKVYNNIGNLSNSSGERVKNKQYSMLN 521
Query: 310 HLQKFYLNLDKLDVDSL-NSLGS--SIVELVQSDDPSS 344
+K LD+ D+ S S+GS SI + S P S
Sbjct: 522 ISKKTI--LDESDISSSPRSIGSPNSIRASISSQLPPS 557
>SR40_YEAST (P32583) Suppressor protein SRP40
Length = 406
Score = 42.4 bits (98), Expect = 0.011
Identities = 66/297 (22%), Positives = 118/297 (39%), Gaps = 28/297 (9%)
Query: 64 PNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGTGPRCD 123
P LS + +++ E++S S + SS S ++ + + + + + + D
Sbjct: 12 PKLSVKEKEI--EEKSSSSSSSSSSSSSSSSSSSSSSSSSGESSSSSSSSSSSSSSDSSD 69
Query: 124 RENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVTAVESN 183
+S SS + S S S S S SSSS S + + S + ++ ES
Sbjct: 70 SSDSESSSSSSSSSSSSSSSSDSESSSESDSSSSGSSSSSSSSSDE--------SSSESE 121
Query: 184 SGEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNMEP 243
S + T +EDA KK + E + E ++ GS S + +
Sbjct: 122 SEDETKKRARESDNEDAKETKKAKTE-PESSSSSESSSSGSSSSSESESGSESDSDSSSS 180
Query: 244 CSSISPNLVD-----KSPKVTGFSDCASPATPSSV--------ACSSSPGVDDKLLGKVG 290
SS S + D +S + SD +S + SS + SSS D
Sbjct: 181 SSSSSDSESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSSSSSSDSDSDSDS 240
Query: 291 NADNDVSNLTDSPAPGFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDDPSSDDS 347
++D+D S +DS + + + + D D DS + GSS +++ + ++D+S
Sbjct: 241 SSDSDSSGSSDSSSSSDSSSDES--TSSDSSDSDSDSDSGSS--SELETKEATADES 293
Score = 37.7 bits (86), Expect = 0.26
Identities = 65/302 (21%), Positives = 115/302 (37%), Gaps = 27/302 (8%)
Query: 229 SNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGK 288
S+ S SS+ + E SS S + S + SD S ++ SS + SSS D + +
Sbjct: 38 SSSSSSSSSSSSGESSSSSSSSSSSSSSDSSDSSDSESSSSSSSSSSSSSSSSDSESSSE 97
Query: 289 VGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSD-DPSSDDS 347
++ + S+ + S + + + + D+ + S E ++ +P S S
Sbjct: 98 SDSSSSGSSSSSSSSSDESSSESE----SEDETKKRARESDNEDAKETKKAKTEPESSSS 153
Query: 348 GLVRSNAINKLLIWKADISKVLEMTESEIDLLENELKSLKSESVDRSECPVASGSQQADS 407
S+ + S+ +ES+ D + S SES S+ +S S +DS
Sbjct: 154 SESSSSG-------SSSSSESESGSESDSDSSSSSSSSSDSESDSESDSQSSSSSSSSDS 206
Query: 408 SSKFYEERVEVSQKVIRPVPLKIISSDEPNTVKMPQSTNLCSIHENDKEEDIDSPGSATS 467
SS + SS + ++ S++ S ++D D DS GS+ S
Sbjct: 207 SSDSDSSSSD--------------SSSDSDSSSSSSSSSSDSDSDSDSSSDSDSSGSSDS 252
Query: 468 KFVEPLPVNAVSSSYTRGYDNLSRDMNAVQSTMMKCFVRCNRKNTSVSACNNVNTPTEVK 527
+ +SS + D+ S D + K K A +N +TP+
Sbjct: 253 SSSSDSSSDESTSSDSSDSDSDS-DSGSSSELETKEATADESKAEETPASSNESTPSASS 311
Query: 528 DS 529
S
Sbjct: 312 SS 313
Score = 33.5 bits (75), Expect = 4.9
Identities = 47/226 (20%), Positives = 88/226 (38%), Gaps = 10/226 (4%)
Query: 1203 SSTAANVSFSPVVNSCSGLSFGSENKHVSFGKPHTSALSMSMSDLQATANSLLLKAAAAQ 1262
SS ++ S S +S S S S + S +S+ S S SD ++++ S + ++
Sbjct: 48 SSGESSSSSSSSSSSSSSDSSDSSDSESSSSSSSSSSSSSSSSDSESSSESDSSSSGSSS 107
Query: 1263 CEKTVSQDRLSSTCDIQGGRDMRCHSSGSNGDHQL------PLSGSHVETVSVLQGYSMQ 1316
+ S D SS + + R S + + P S S E+ S S +
Sbjct: 108 SSSS-SSDESSSESESEDETKKRARESDNEDAKETKKAKTEPESSSSSESSSSGSSSSSE 166
Query: 1317 VPIKKEVDGDVNCSSSAA---EFPLLPQKVKQTDGHFKPSFHSSNSEKTSRNGDVKLFGK 1373
E D D + SSS++ E + S S+S + + D
Sbjct: 167 SESGSESDSDSSSSSSSSSDSESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSS 226
Query: 1374 ILTNPSSTQNPNLTAKRSEENGSHHPKLNNKSSNLNFTGHQNSDEN 1419
++ S + + + ++ S+ +GS ++ SS+ T +SD +
Sbjct: 227 SSSSSSDSDSDSDSSSDSDSSGSSDSSSSSDSSSDESTSSDSSDSD 272
>YIQ9_YEAST (P40442) Hypothetical 99.7 kDa protein in SDL1 5'region
precursor
Length = 995
Score = 41.6 bits (96), Expect = 0.018
Identities = 70/369 (18%), Positives = 141/369 (37%), Gaps = 32/369 (8%)
Query: 214 LAKYEKKKVDVPDPGSNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGF--SDCASPATPS 271
L +Y + S+ SS+G++ SIS ++ + S T S S ++ S
Sbjct: 22 LGQYYSNSTSISSNSSSTSVVSSSSGSV----SISSSIAETSSSATDILSSITQSASSTS 77
Query: 272 SVACSSSPGVDDKLLGKVGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKLDVDSLNSLGS 331
V+ S P + V + + VS+++ S + + DV S S +
Sbjct: 78 GVSSSVGPSSSSVVSSSVSQSSSSVSDVSSSVSQS----------SSSASDVSSSVSQSA 127
Query: 332 SIVELVQSDDPSSDDSGLVRSNAINKLLIWKADISKVLEMTESEIDLLENELKSLKSESV 391
S V S S S S+++++ +D+S + + S + + + S +
Sbjct: 128 SSTSDVSSSVSQSSSSASDVSSSVSQSSSSASDVSSSVSQSASSASDVSSSVSQSASSTS 187
Query: 392 DRSECPVASGSQQADSSSKFYEERVEVSQKVIRPVPLKIISSDEPNTVKMPQSTNLCSIH 451
D S S S +D SS + S + + SS + + QS + S
Sbjct: 188 DVSSSVSQSSSSASDVSSSVSQSSSSASD--VSSSVSQSASSTSDVSSSVSQSASSTSGV 245
Query: 452 ENDKEEDIDSPGSATSKFVEPLPVNAVSSSYTRGYDNLSRDMNAVQSTMMKCFVRCNRKN 511
+ + + S ++S F + + SS+ T S ++++ S+ N
Sbjct: 246 SSSGSQSVSSASGSSSSFPQ-----STSSASTASGSATSNSLSSITSSASS--ASATASN 298
Query: 512 TSVSACNNVNTPTEVKDSLGDVTFGANLCSSYG-----DTYKSIIASNKESANRAHKLFT 566
+ S+ + PT GD+T + ++ G +++ +K S + K++
Sbjct: 299 SLSSSDGTIYLPTTTIS--GDLTLTGKVIATEGVVVAAGAKLTLLDGDKYSFSADLKVYG 356
Query: 567 KLVPKECKK 575
L+ K+ K+
Sbjct: 357 DLLVKKSKE 365
>DMP1_RAT (P98193) Dentin matrix acidic phosphoprotein 1 precursor
(Dentin matrix protein-1) (DMP-1) (AG1)
Length = 489
Score = 41.6 bits (96), Expect = 0.018
Identities = 61/262 (23%), Positives = 104/262 (39%), Gaps = 29/262 (11%)
Query: 6 PGHGYGVSRSGDKMLEEDGRPLVSRGDGKYGRSSRDNRGGPFGQRDWRGHS-WEASNGSP 64
P G S D EED GD +G DN GP +R W G S ++ S
Sbjct: 69 PAGGLSKSAGMDADKEEDED---DSGDDTFG--DEDNGPGP-EERQWGGPSRLDSDEDSA 122
Query: 65 NLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNT------WEQHNLKDQHAKTGGVNG--- 115
+ ++ +D +++ S D+P+ S HSD ++ Q + +++ GG G
Sbjct: 123 DTTQSSEDSTSQENSAQDTPSDSKDHHSDEADSRPEAGDSTQDSESEEYRVGGGSEGESS 182
Query: 116 LGTGPRCDRENSLSS------IDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYE-- 167
G G D E S D + + +G S G +S++ + E
Sbjct: 183 HGDGSEFDDEGMQSDDPGSTRSDRGHTRMSSAGIRSEESKGDHEPTSTQDSDDSQDVEFS 242
Query: 168 GKPNLKHKNVTAVESNSGE---ATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDV 224
+ + + V+ E + GE + + T S+ +ED S+++ R E A+ + ++ D
Sbjct: 243 SRKSFRRSRVSE-EDDRGELADSNSRETQSVSTEDFRSKEESRSETQEDTAETQSQE-DS 300
Query: 225 PDPGSNKDGSVSSAGNMEPCSS 246
P+ S AG SS
Sbjct: 301 PEGQDPSSESSEEAGEPSQESS 322
>SPR1_CAEEL (Q18919) Suppressor of presenilin 1
Length = 558
Score = 41.2 bits (95), Expect = 0.023
Identities = 17/54 (31%), Positives = 31/54 (56%)
Query: 728 IEKERSMINPWTSEEKELFLEKFAAFGKDFRKIASFLDHKTTADCIEFYYKNHK 781
+ + + + WT +E LF + FGK+F +I S L H++ ++FYY++ K
Sbjct: 188 VARRNELKDVWTDQEITLFENCYQIFGKNFSQIRSALCHRSLQSIVQFYYESKK 241
>DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin
matrix protein-3) (DMP-3) [Contains: Dentin
phosphoprotein (Dentin phosphophoryn) (DPP); Dentin
sialoprotein (DSP)]
Length = 934
Score = 40.8 bits (94), Expect = 0.031
Identities = 84/407 (20%), Positives = 139/407 (33%), Gaps = 46/407 (11%)
Query: 3 SEEPGHGYGVSRSGDKMLEEDGRPLVSRGDGKYGRSSRDNRGGPFGQRDWRGHSWEASNG 62
S++ S S D D D SS D+ D G S + +
Sbjct: 552 SDDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNSSSDSS-------DSSGSSDSSDSS 604
Query: 63 SPNLSRRPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGTGPRC 122
S D ++ S D S + S SD ++ + + D + + +
Sbjct: 605 DTCDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSSSSDSSDSSSCSDSSDSSDSS 664
Query: 123 DRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVTAVES 182
D +S S D + S + S SSSS S +DS + + + +S
Sbjct: 665 DSSDSSDSSDSSSSDSSSSSNSSDSSDSSDSSSSSDSSDSSDSSDSS-----DSSGSSDS 719
Query: 183 NSGEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNME 242
+ A++ +SS S D++S D D + D S SS +
Sbjct: 720 SDSSASSDSSSSSDSSDSSS---------------SSDSSDSSDSSDSSDSSESSDSSNS 764
Query: 243 PCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSNLTDS 302
SS S + D S S +S ++ SS + +SS D + +D SN +DS
Sbjct: 765 SDSSDSSDSSDSSD-----SSDSSDSSDSSDSSNSSDSSD----SSDSSDSSDSSNSSDS 815
Query: 303 PAPGFQNHLQKFYLNLDKLDVDSLNSLGSSIVELVQSDDPSSDDSGLVRSNAINKLLIWK 362
+ + D D + S SD S DS ++ +
Sbjct: 816 SDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDS 875
Query: 363 ADISKVLEMTESEIDLLENELKSLKSESVDRSECPVASGSQQADSSS 409
+D S + ++S+ K S D S+ SG+ +DS+S
Sbjct: 876 SDSSNSSDSSDSD----------SKDSSSDSSDGDSKSGNGNSDSNS 912
Score = 40.0 bits (92), Expect = 0.052
Identities = 63/294 (21%), Positives = 102/294 (34%), Gaps = 16/294 (5%)
Query: 13 SRSGDKMLEEDGRPLVSRGDGKYGRSSRDNRGGPFGQRDWRGHSWEASNGSPNL----SR 68
S S D D S D S D+ S +SN S + S
Sbjct: 637 SDSSDSSSSSDSSDSSSCSDSSDSSDSSDSSDSSDSSDSSSSDSSSSSNSSDSSDSSDSS 696
Query: 69 RPQDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLKDQHAKTGGVNGLGTGPRCDRENSL 128
D ++ S D S + S SD + + + D + + + D +S
Sbjct: 697 SSSDSSDSSDSSDSSDSSGSSDSSDSSASSDSSSSSDSSDSSSSSDSSDSSDSSDSSDSS 756
Query: 129 SSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEGKPNLKHKNVTAVESNSGEAT 188
S D + S SS S S SS S + + + + + ++ SNS +++
Sbjct: 757 ESSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSS 816
Query: 189 ACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKDGSVSSAGNMEPCSSIS 248
SS S+ + S + D D + D S SS + SS S
Sbjct: 817 DSSDSSDSSDSSDSSD----------SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDS 866
Query: 249 PNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNADNDVSNLTDS 302
+ D S + S +S ++ S SSS D GN+D++ + +DS
Sbjct: 867 SDSSDSSD--SSDSSNSSDSSDSDSKDSSSDSSDGDSKSGNGNSDSNSDSNSDS 918
>APXL_HUMAN (Q13796) Apical-like protein (APXL protein)
Length = 1616
Score = 40.0 bits (92), Expect = 0.052
Identities = 50/200 (25%), Positives = 76/200 (38%), Gaps = 26/200 (13%)
Query: 52 WRGHSWEASNGSPNLSRRPQDMNNEQRSVDDSPTYSSHPHS-----DFVNTWEQHNLK-- 104
WR HSW A+ S + P+ + S P++S H+ D ++WEQ NL+
Sbjct: 112 WRPHSWHATKFSDS---HPELAASPFTSTSGCPSWSGRHHASSSSHDLSSSWEQTNLQRT 168
Query: 105 -DQHAKTGGVNGLG-TGPRCDRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRS--- 159
D + G V+ L R S SSID GS S R S + S+S S
Sbjct: 169 LDHFSSLGSVDSLDHPSSRLSVAKSNSSID-------HLGSHSKRDSAYGSFSTSSSTPD 221
Query: 160 --MAGTDSYEGKPNLKHKNVTAVESNSGEATACVTSSMPSEDATSRKKPRL--NWGEGLA 215
++ D+ + L + G SE+ S PR+ + G+G
Sbjct: 222 HTLSKADTSSAENILYTVGLWEAPRQGGRQAQAAGDPQGSEEKLSCFPPRVPGDSGKGPR 281
Query: 216 KYEKKKVDVPDPGSNKDGSV 235
+ + PG + G V
Sbjct: 282 PEYNAEPKLAAPGRSNFGPV 301
>SERI_BOMMO (P07856) Sericin precursor (Silk gum protein)
Length = 389
Score = 39.7 bits (91), Expect = 0.068
Identities = 62/306 (20%), Positives = 116/306 (37%), Gaps = 38/306 (12%)
Query: 11 GVSRSGDKMLEEDGRPLVSRGDGKYGRSSRDNRGGPFGQRDWRGHSWEASNGSPNLSRRP 70
GVS SG +D + + G K S+ + Q S E+++ S + S
Sbjct: 78 GVSTSGRSQNYKDSKQAIISGGTKSSNSNVQSDEKSASQSSSSRSSQESASYSSSSSSST 137
Query: 71 QDMNNEQRSVDDSPTYSSHPHSDFVNTWEQHNLK------DQHAKTGGVNGLGTGPRCDR 124
++ ++ S SS+ S+ + + +++ G V+ G+ D
Sbjct: 138 EESSSSSSRAASSTDASSNTDSNSNSAGSSTSGGRRTYGYSSNSRDGSVSSTGSSSNTDS 197
Query: 125 ENSLS-------SIDWKPLKWTRSGSLSSRGSGF-----SHSSSSRSMAGTDSYEGKPNL 172
+S + S + +R GS+S+ GS S+S SR G+ S+E
Sbjct: 198 NSSNAGSSTSGGSSTYGYSSNSRDGSVSTTGSSSNTDSNSNSVGSRRSGGSSSHEDSSKS 257
Query: 173 KHKNVTAVESNSGEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKVDVPDPGSNKD 232
+ +NV+ S+S T S + +S R +G +++D
Sbjct: 258 RDENVSTTGSSSN------TDSNSNSVGSSTSGGRRTYGYS--------------SNSRD 297
Query: 233 GSVSSAGNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVDDKLLGKVGNA 292
GSVSS G+ S S ++ + + +S + SV+ + S D G++
Sbjct: 298 GSVSSTGSSSNTDSNSNSVGSSTSGGSSTYGYSSNSRDGSVSSTGSSSNTDSNSNSAGSS 357
Query: 293 DNDVSN 298
+ S+
Sbjct: 358 TSGGSS 363
Score = 39.3 bits (90), Expect = 0.089
Identities = 59/235 (25%), Positives = 95/235 (40%), Gaps = 31/235 (13%)
Query: 1225 SENKHVSFGKPHTSALSMSMSDLQATANSLLLKAAAAQCEKTVSQDRLSSTCDIQGGRDM 1284
S N +V + S S S S ++ + S ++++ E + S R +S+ D D
Sbjct: 102 SSNSNVQSDEKSASQSSSSRSSQESASYSS--SSSSSTEESSSSSSRAASSTDASSNTDS 159
Query: 1285 RCHSSGSNGDHQLPLSGSHVETVSVLQGYSMQVPIKKEVDGDVNCSSSAAEFPLLPQKV- 1343
+S+GS+ SG GYS DG V+ + S++
Sbjct: 160 NSNSAGSS------TSGGRRT-----YGYS-----SNSRDGSVSSTGSSSNTDSNSSNAG 203
Query: 1344 KQTDGHFKPSFHSSNSEKTSRNGDVKLFGKILTNPSSTQNPNLTAKRSEENGSHHPKLNN 1403
T G +SSNS R+G V G +N S N ++ ++RS + SH +
Sbjct: 204 SSTSGGSSTYGYSSNS----RDGSVSTTGSS-SNTDSNSN-SVGSRRSGGSSSHEDSSKS 257
Query: 1404 KSSNLNFTG-HQNSDENLNFLKFGLENVPVMSYGYWEGNAIQSRQSGLSSLPDSS 1457
+ N++ TG N+D N N + +YGY + SR +SS SS
Sbjct: 258 RDENVSTTGSSSNTDSNSNSVGSSTSG-GRRTYGY----SSNSRDGSVSSTGSSS 307
Score = 35.0 bits (79), Expect = 1.7
Identities = 48/189 (25%), Positives = 73/189 (38%), Gaps = 14/189 (7%)
Query: 104 KDQHAKTGGVNGLGTGPRCDR-ENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAG 162
+D+ T G G+ T R ++S +I K + S S S S SSSSRS
Sbjct: 67 QDKSKYTSGPEGVSTSGRSQNYKDSKQAIISGGTKSSNSNVQSDEKSA-SQSSSSRSSQE 125
Query: 163 TDSYEGKPNLKHKNVTAVESNSGEATACVTSSMPSEDATSRKKPRLNWGEGLAKYEKKKV 222
+ SY + + T S+S A T + + D+ S G G Y
Sbjct: 126 SASYSSSSS----SSTEESSSSSSRAASSTDASSNTDSNSNSAGSSTSG-GRRTYGYSS- 179
Query: 223 DVPDPGSNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGFSDCASPATPSSVACSSSPGVD 282
+++DGSVSS G+ S S N + + +S + SV+ + S
Sbjct: 180 ------NSRDGSVSSTGSSSNTDSNSSNAGSSTSGGSSTYGYSSNSRDGSVSTTGSSSNT 233
Query: 283 DKLLGKVGN 291
D VG+
Sbjct: 234 DSNSNSVGS 242
>2A5D_YEAST (P38903) Serine/threonine protein phosphatase 2A, 56 kDa
regulatory subunit, delta isoform (PP2A, B subunit, B'
delta isoform) (RTS1 protein) (SCS1 protein)
Length = 757
Score = 39.7 bits (91), Expect = 0.068
Identities = 52/216 (24%), Positives = 87/216 (40%), Gaps = 34/216 (15%)
Query: 109 KTGGVNGLGTGPRCDRENSLSSIDWKPLKWTRSGSLSSRGSGFSHSSSSRSMAGTDSYEG 168
KT G + + + D+E + S SS G +HSS+S + + + + +G
Sbjct: 12 KTTGSSSSSSSKKKDKEKEKEKSSTTSSTSKKPASASSSSHGTTHSSASSTGSKSTTEKG 71
Query: 169 KPN--------------LKHKNVTAVESNSGEATACVTSSMPSEDATSRKKPRLNWGEGL 214
K + K K T S+S + +SS+ ++S KK G+
Sbjct: 72 KQSGSVPSQGKHHSSSTSKTKTATTPSSSSSSSR---SSSVSRSGSSSTKKTSSRKGQEQ 128
Query: 215 AKYEKKKVDVPDPGSNKDGSVSSAGNMEPCSSISPNLVDKSPKVTGFSDCASP------- 267
+K ++ P + S SSA M P ++ V K K T D A P
Sbjct: 129 SKQSQQ----PSQSQKQGSSSSSAAIMNPTPVLT---VTKDDKSTSGEDHAHPTLLGAVS 181
Query: 268 ATPSS-VACSSSPGVDDKLLGKVGNADNDVSNLTDS 302
A PSS ++ +S V + + GN++N+ N+ S
Sbjct: 182 AVPSSPISNASGTAVSSDV--ENGNSNNNNMNINTS 215
>RUS1_MOUSE (Q8BG26) RUN and SH3 domain containing protein 1
Length = 893
Score = 39.3 bits (90), Expect = 0.089
Identities = 63/238 (26%), Positives = 99/238 (41%), Gaps = 42/238 (17%)
Query: 218 EKKKVDVPDPGSNKDGSVSSAGNMEPCSS-IS------PNLVDKSPKVTGFS-----DCA 265
E++ V DPG + S+SS ++ P S +S P D +P+ + A
Sbjct: 90 EEEAVSPSDPGCSS--SLSSCSDLSPDESPVSVYSRDLPGNEDANPQPSTLELGSPLAPA 147
Query: 266 SPAT--PSSVACS--SSPGVDDKLLGKVGNADNDVSNLTDSPAPGFQNHLQKFYLNLDKL 321
P+T P S CS S G+ + + N ++ D P+PG L++
Sbjct: 148 GPSTCSPDSFCCSPDSCSGISSPPGPDLDSNCNALTTCQDLPSPG-----------LEEE 196
Query: 322 DVDSLNSLGSSIVELVQSDDPSSDDSGLVRSNAINKLLIWKADISKV------LEMTESE 375
+ L +S EL +++D D S IN IWK D K +E ++S
Sbjct: 197 EDSGEQDLATS--ELSETEDGRIDAGKAEPSWKINP--IWKIDTEKTEAGWKTIEDSDSG 252
Query: 376 IDLLENELKSLKSESVDRSECPVASGSQQADSSSKFYEERVEVSQKVIRPVPLKIISS 433
EN SLK+ES + C + + D+ R +VSQ+ PVP + I+S
Sbjct: 253 RKTDENTNSSLKTESGKLASCLNTNSGSKIDAGKTDGGWRGDVSQE---PVPHRTITS 307
>CRM_DROSE (Q8MX88) Cramped protein
Length = 975
Score = 39.3 bits (90), Expect = 0.089
Identities = 22/65 (33%), Positives = 32/65 (48%), Gaps = 9/65 (13%)
Query: 925 QPLTPDISQN--------ADDETCSDESCGEA-TEWTDDETAAFLQAVSSFGKDFEKISR 975
+P+TP S+ A +T S G T WT+ E F A++ FGKDFE ++
Sbjct: 79 RPMTPPPSEREPNKKEEKAAQKTPSQLKTGSGKTTWTNVERNCFFDALNEFGKDFEAVAN 138
Query: 976 CVGTK 980
C+ K
Sbjct: 139 CINAK 143
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.310 0.128 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 193,204,415
Number of Sequences: 164201
Number of extensions: 8552487
Number of successful extensions: 19412
Number of sequences better than 10.0: 100
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 85
Number of HSP's that attempted gapping in prelim test: 19138
Number of HSP's gapped (non-prelim): 282
length of query: 1620
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1496
effective length of database: 39,613,130
effective search space: 59261242480
effective search space used: 59261242480
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 73 (32.7 bits)
Medicago: description of AC126015.12


