

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
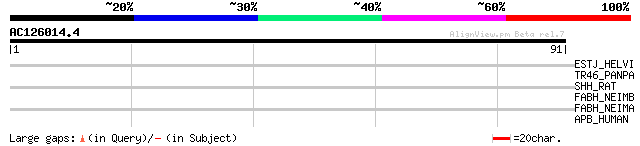
Query= AC126014.4 + phase: 0
(91 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ESTJ_HELVI (P12992) Juvenile hormone esterase precursor (EC 3.1.... 29 2.4
TR46_PANPA (Q646E1) Taste receptor type 2 member 46 (T2R46) 27 6.9
SHH_RAT (Q63673) Sonic hedgehog protein precursor (SHH) 27 9.1
FABH_NEIMB (Q9JXR6) 3-oxoacyl-[acyl-carrier-protein] synthase II... 27 9.1
FABH_NEIMA (Q9JW56) 3-oxoacyl-[acyl-carrier-protein] synthase II... 27 9.1
APB_HUMAN (P04114) Apolipoprotein B-100 precursor (Apo B-100) [C... 27 9.1
>ESTJ_HELVI (P12992) Juvenile hormone esterase precursor (EC
3.1.1.59) (JH esterase)
Length = 564
Score = 28.9 bits (63), Expect = 2.4
Identities = 17/46 (36%), Positives = 23/46 (49%), Gaps = 5/46 (10%)
Query: 23 LQVIWHACVWLIWKETNSR-----IFNHKTKEVGLLLDSVKLTSFL 63
L + HAC L W+ETNSR + + + V D +K SFL
Sbjct: 8 LAFLLHACTALAWQETNSRSVVAHLDSGIIRGVPRSADGIKFASFL 53
>TR46_PANPA (Q646E1) Taste receptor type 2 member 46 (T2R46)
Length = 309
Score = 27.3 bits (59), Expect = 6.9
Identities = 12/34 (35%), Positives = 18/34 (52%)
Query: 7 HRFGNLAGAPRSTHTFLQVIWHACVWLIWKETNS 40
H F + G + TFL V+WH W+ +E +S
Sbjct: 275 HPFILIWGNKKLKQTFLSVLWHVRYWVKGEEPSS 308
>SHH_RAT (Q63673) Sonic hedgehog protein precursor (SHH)
Length = 437
Score = 26.9 bits (58), Expect = 9.1
Identities = 10/26 (38%), Positives = 15/26 (57%)
Query: 13 AGAPRSTHTFLQVIWHACVWLIWKET 38
AG P H + Q+++H WL+ ET
Sbjct: 401 AGPPAGIHWYSQLLYHIGTWLLDSET 426
>FABH_NEIMB (Q9JXR6) 3-oxoacyl-[acyl-carrier-protein] synthase III
(EC 2.3.1.41) (Beta-ketoacyl-ACP synthase III) (KAS III)
Length = 320
Score = 26.9 bits (58), Expect = 9.1
Identities = 11/29 (37%), Positives = 16/29 (54%)
Query: 32 WLIWKETNSRIFNHKTKEVGLLLDSVKLT 60
W++ + N RI K +GL +D V LT
Sbjct: 243 WIVPHQANRRIIESTAKHLGLSMDKVVLT 271
>FABH_NEIMA (Q9JW56) 3-oxoacyl-[acyl-carrier-protein] synthase III
(EC 2.3.1.41) (Beta-ketoacyl-ACP synthase III) (KAS III)
Length = 320
Score = 26.9 bits (58), Expect = 9.1
Identities = 11/29 (37%), Positives = 16/29 (54%)
Query: 32 WLIWKETNSRIFNHKTKEVGLLLDSVKLT 60
W++ + N RI K +GL +D V LT
Sbjct: 243 WIVPHQANRRIIESTAKHLGLSMDKVVLT 271
>APB_HUMAN (P04114) Apolipoprotein B-100 precursor (Apo B-100)
[Contains: Apolipoprotein B-48 (Apo B-48)]
Length = 4563
Score = 26.9 bits (58), Expect = 9.1
Identities = 12/23 (52%), Positives = 15/23 (65%)
Query: 39 NSRIFNHKTKEVGLLLDSVKLTS 61
NSR F TK++GL +S K TS
Sbjct: 295 NSRFFGEGTKKMGLAFESTKSTS 317
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.137 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,874,068
Number of Sequences: 164201
Number of extensions: 342896
Number of successful extensions: 906
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 899
Number of HSP's gapped (non-prelim): 7
length of query: 91
length of database: 59,974,054
effective HSP length: 67
effective length of query: 24
effective length of database: 48,972,587
effective search space: 1175342088
effective search space used: 1175342088
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC126014.4


