

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
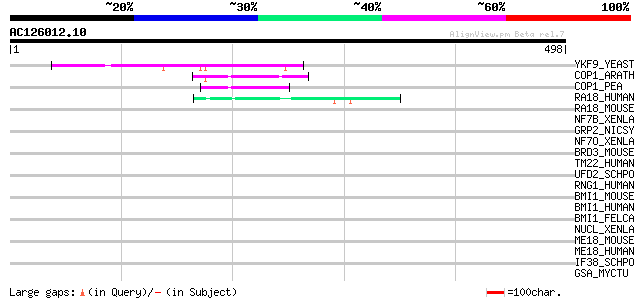
Query= AC126012.10 + phase: 0 /pseudo
(498 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YKF9_YEAST (P35728) Hypothetical 49.6 kDa protein in FBA1-TOA2 i... 87 8e-17
COP1_ARATH (P43254) Ubiquitin ligase protein COP1 (EC 6.3.2.-) (... 54 9e-07
COP1_PEA (P93471) Ubiquitin ligase protein COP1 (EC 6.3.2.-) (Co... 50 1e-05
RA18_HUMAN (Q9NS91) Postreplication repair protein RAD18 (hRAD18... 48 5e-05
RA18_MOUSE (Q9QXK2) Postreplication repair protein RAD18 (mRAD18Sc) 41 0.008
NF7B_XENLA (Q92021) Nuclear factor 7, brain (xnf7) (xnf7-B) 40 0.018
GRP2_NICSY (P27484) Glycine-rich protein 2 40 0.018
NF7O_XENLA (Q91431) Nuclear factor 7, ovary (xnf7-O) 39 0.041
BRD3_MOUSE (Q8K2F0) Bromodomain-containing protein 3 (Bromodomai... 39 0.041
TM22_HUMAN (Q8IYM9) Tripartite motif-containing protein 22 (RING... 38 0.069
UFD2_SCHPO (Q9HE05) Ubiquitin conjugation factor E4 (Ubiquitin f... 37 0.12
RNG1_HUMAN (Q06587) Polycomb complex protein RING1 (RNF1) 37 0.12
BMI1_MOUSE (P25916) Polycomb complex protein BMI-1 37 0.12
BMI1_HUMAN (P35226) Polycomb complex protein BMI-1 37 0.12
BMI1_FELCA (Q9TST0) Polycomb complex protein BMI-1 37 0.12
NUCL_XENLA (P20397) Nucleolin (Protein C23) 37 0.15
ME18_MOUSE (P23798) DNA-binding protein Mel-18 (RING finger prot... 37 0.15
ME18_HUMAN (P35227) DNA-binding protein Mel-18 (RING finger prot... 37 0.15
IF38_SCHPO (O14164) Probable eukaryotic translation initiation f... 37 0.15
GSA_MYCTU (P63506) Glutamate-1-semialdehyde 2,1-aminomutase (EC ... 37 0.15
>YKF9_YEAST (P35728) Hypothetical 49.6 kDa protein in FBA1-TOA2
intergenic region
Length = 441
Score = 87.4 bits (215), Expect = 8e-17
Identities = 65/257 (25%), Positives = 111/257 (42%), Gaps = 35/257 (13%)
Query: 38 SKADEDSKIKALIDTPALDWQQQGSDFGAGRGFRRGAVGGRIGGGRGFGLERKTPPEGYV 97
S E+ +I ++ T W+Q + A + + G PP GY+
Sbjct: 126 SGTTEEERIASMFATQENQWEQTQEEMSAATPVFFKSQTNKNSAQENEG----PPPPGYM 181
Query: 98 CHRCKVSGHFIQHCPTNGDSTFDIKKVRQPTGIPRSMLM---VNPQGSYALQNGSVAVLK 154
C+RC H+I++CPTN D F+ K++R+ TGIP+ L ++P+ + ++
Sbjct: 182 CYRCGGRDHWIKNCPTNSDPNFEGKRIRRTTGIPKKFLKSIEIDPETMTPEEMAQRKIMI 241
Query: 155 PNEAAFDKEMEGLSS---------TRSVG------------DLPPELHCPLCNNVMKDAV 193
+E F ++E S R + DLP +L CPL +++ V
Sbjct: 242 TDEGKFVVQVEDKQSWEDYQRKRENRQIDGDETIWRKGHFKDLPDDLKCPLTGGLLRQPV 301
Query: 194 LTSKCCFKSFCDKCIRNYIM-SKSACV-CLATNILADDLLPNKTLRDAIHRILES----- 246
TSKCC F + + N ++ S C C +IL D L+P++ + L+
Sbjct: 302 KTSKCCNIDFSKEALENALVESDFVCPNCETRDILLDSLVPDQDKEKEVETFLKKQEELH 361
Query: 247 GNSSTENAGSTYQVQDM 263
G+S N T +++ M
Sbjct: 362 GSSKDGNQPETKKMKLM 378
>COP1_ARATH (P43254) Ubiquitin ligase protein COP1 (EC 6.3.2.-)
(Constitutive photomorphogenesis protein 1)
Length = 675
Score = 53.9 bits (128), Expect = 9e-07
Identities = 33/106 (31%), Positives = 53/106 (49%), Gaps = 6/106 (5%)
Query: 165 EGLSSTRSVG--DLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLA 222
+G S +G DL +L CP+C ++KDA LT+ C SFC CI ++ +KS C C +
Sbjct: 33 DGGSGGSEIGAPDLDKDLLCPICMQIIKDAFLTA--CGHSFCYMCIITHLRNKSDCPCCS 90
Query: 223 TNILADDLLPNKTLRDAIHRILESGNSSTENAGSTYQVQDMLSSRC 268
++ + L PN L + + S ++ A Q ++ L C
Sbjct: 91 QHLTNNQLYPNFLLDKLLKK--TSARHVSKTASPLDQFREALQRGC 134
>COP1_PEA (P93471) Ubiquitin ligase protein COP1 (EC 6.3.2.-)
(Constitutive photomorphogenesis protein 1)
Length = 672
Score = 50.4 bits (119), Expect = 1e-05
Identities = 26/80 (32%), Positives = 41/80 (50%), Gaps = 2/80 (2%)
Query: 172 SVGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLATNILADDLL 231
++ DL + CP+C ++KDA LT+ C SFC CI ++ +KS C C + +L
Sbjct: 37 TMSDLDKDFLCPICMQIIKDAFLTA--CGHSFCYMCIITHLRNKSDCPCCGHYLTNSNLF 94
Query: 232 PNKTLRDAIHRILESGNSST 251
PN L + + + S T
Sbjct: 95 PNFLLDKLLKKTSDRQISKT 114
>RA18_HUMAN (Q9NS91) Postreplication repair protein RAD18 (hRAD18)
(hHR18)
Length = 495
Score = 48.1 bits (113), Expect = 5e-05
Identities = 42/201 (20%), Positives = 76/201 (36%), Gaps = 29/201 (14%)
Query: 166 GLSSTRSVGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLATNI 225
GL+ +++ DL L C +C A++ +C ++C CIR ++ K+ C +
Sbjct: 12 GLAVMKTIDDL---LRCGICFEYFNIAMIIPQCSH-NYCSLCIRKFLSYKTQCPTCCVTV 67
Query: 226 LADDLLPNKTLRDAIHRILESGNSSTENAGSTYQVQDMLSSRCPQPKIPSPTSSATSKGE 285
DL N+ L + + + N + +Q L S P S + A
Sbjct: 68 TEPDLKNNRILDELVKSL---------NFARNHLLQFALESPAKSPASSSSKNLAVKVYT 118
Query: 286 PKVSQ--------------VNEGMANIQEIVAE--RKEVSATQQVSEQVKIPRAAVVSEV 329
P S+ + E + E++ + + + S ++ S K V E+
Sbjct: 119 PVASRQSLKQGSRLMDNFLIREMSGSTSELLIKENKSKFSPQKEASPAAKTKETRSVEEI 178
Query: 330 THESMSVKEPEPASQGSAKLV 350
+ K PEP S + K V
Sbjct: 179 APDPSEAKRPEPPSTSTLKQV 199
>RA18_MOUSE (Q9QXK2) Postreplication repair protein RAD18 (mRAD18Sc)
Length = 509
Score = 40.8 bits (94), Expect = 0.008
Identities = 34/135 (25%), Positives = 58/135 (42%), Gaps = 18/135 (13%)
Query: 166 GLSSTRSVGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLATNI 225
GL+ +++ DL L C +C AV+ +C ++C CIR ++ K+ C +
Sbjct: 12 GLAVMKTIDDL---LRCGICFEYFNIAVIIPQCSH-NYCSLCIRKFLSYKTQCPTCCVAV 67
Query: 226 LADDLLPNKTLRDAIHRILESGNSSTENAGSTYQVQDMLSSRCPQPKIPSPTSSATSKGE 285
DL N+ L + + + N T+ +Q L S P I SP SS + K
Sbjct: 68 TEPDLRNNRLLDELV---------KSMNFARTHLLQFALES----PPI-SPVSSTSKKVV 113
Query: 286 PKVSQVNEGMANIQE 300
KV + +++
Sbjct: 114 VKVHNADAAQHPVKQ 128
>NF7B_XENLA (Q92021) Nuclear factor 7, brain (xnf7) (xnf7-B)
Length = 609
Score = 39.7 bits (91), Expect = 0.018
Identities = 22/69 (31%), Positives = 35/69 (49%), Gaps = 4/69 (5%)
Query: 154 KPNEAAFDKE--MEGLSSTRSVGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNY 211
+P +A +++ + SS + GD EL CPLC + KD V+ + C +FC CI
Sbjct: 115 EPKKAKVEEKDASKNASSLGAAGDFAEELTCPLCVELFKDPVMVA--CGHNFCRSCIDKA 172
Query: 212 IMSKSACVC 220
+S+ C
Sbjct: 173 WEGQSSFAC 181
>GRP2_NICSY (P27484) Glycine-rich protein 2
Length = 214
Score = 39.7 bits (91), Expect = 0.018
Identities = 22/55 (40%), Positives = 26/55 (47%), Gaps = 2/55 (3%)
Query: 61 GSDFGAGRGFRRGAVGGRIGGGRGFGLERKTPPEGYVCHRCKVSGHFIQHCPTNG 115
G G G G R G GG GGG G+G G C +C SGHF + C +G
Sbjct: 124 GGGGGYGGGSRYGGGGGGYGGGGGYGGGGSGGGSG--CFKCGESGHFARDCSQSG 176
Score = 34.7 bits (78), Expect = 0.59
Identities = 17/43 (39%), Positives = 21/43 (48%), Gaps = 7/43 (16%)
Query: 73 GAVGGRIGGGRGFGLERKTPPEGYVCHRCKVSGHFIQHCPTNG 115
G GGR GGG G G G C++C GHF + C + G
Sbjct: 178 GGGGGRFGGGGGGG-------GGGGCYKCGEDGHFARECTSGG 213
>NF7O_XENLA (Q91431) Nuclear factor 7, ovary (xnf7-O)
Length = 610
Score = 38.5 bits (88), Expect = 0.041
Identities = 19/53 (35%), Positives = 27/53 (50%), Gaps = 2/53 (3%)
Query: 168 SSTRSVGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVC 220
+S + GD EL CPLC + KD V+ + C +FC CI +S+ C
Sbjct: 132 ASLGAAGDFAEELTCPLCVELFKDPVMVA--CGHNFCRSCIDKVWEGQSSFAC 182
>BRD3_MOUSE (Q8K2F0) Bromodomain-containing protein 3
(Bromodomain-containing FSH-like protein FSRG2)
Length = 726
Score = 38.5 bits (88), Expect = 0.041
Identities = 23/103 (22%), Positives = 45/103 (43%), Gaps = 9/103 (8%)
Query: 271 PKIPSPTSSATSKGEPKVSQVNEGMANIQEIVAERKEVSATQQVSEQVKIPRAAVVSEVT 330
P +P+PT+ SKG E ++ +E + + ++ EQ+K
Sbjct: 422 PALPAPTAPIVSKGAESSRSSEESSSDSGSSDSEEERATRLAELQEQLK---------AV 472
Query: 331 HESMSVKEPEPASQGSAKLVEEEVQQKLVPTDGGKKKKKKKVR 373
HE ++ P ++ K ++E ++K D K+K+K K +
Sbjct: 473 HEQLAALSQAPVNKPKKKKEKKEKEKKKKDKDKDKEKEKHKAK 515
>TM22_HUMAN (Q8IYM9) Tripartite motif-containing protein 22 (RING
finger protein 94) (50 kDa stimulated trans-acting
factor) (Staf-50)
Length = 498
Score = 37.7 bits (86), Expect = 0.069
Identities = 25/85 (29%), Positives = 41/85 (47%), Gaps = 9/85 (10%)
Query: 175 DLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCI-----RNYIMSK--SACVCLATNILA 227
D+ E+ CP+C ++ + + S C SFC CI + I+S+ S+C T
Sbjct: 8 DIEKEVTCPICLELLTEPL--SLDCGHSFCQACITAKIKESVIISRGESSCPVCQTRFQP 65
Query: 228 DDLLPNKTLRDAIHRILESGNSSTE 252
+L PN+ L + + R+ E S E
Sbjct: 66 GNLRPNRHLANIVERVKEVKMSPQE 90
>UFD2_SCHPO (Q9HE05) Ubiquitin conjugation factor E4 (Ubiquitin fusion
degradation protein 2) (UB fusion protein 2)
Length = 1010
Score = 37.0 bits (84), Expect = 0.12
Identities = 23/83 (27%), Positives = 41/83 (48%), Gaps = 1/83 (1%)
Query: 164 MEGLSSTRSVGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLAT 223
++ + +GD+P PL +MKD V+ + S I+ +++S + T
Sbjct: 919 LQEATEEEDMGDIPDYFLDPLMFTIMKDPVVLPRSGI-SIDRSTIKAHLLSDATDPFNRT 977
Query: 224 NILADDLLPNKTLRDAIHRILES 246
+ DD+ PN TLR+ I+ L+S
Sbjct: 978 PLTLDDVTPNDTLREEINTFLKS 1000
>RNG1_HUMAN (Q06587) Polycomb complex protein RING1 (RNF1)
Length = 377
Score = 37.0 bits (84), Expect = 0.12
Identities = 25/85 (29%), Positives = 37/85 (43%), Gaps = 3/85 (3%)
Query: 164 MEGLSSTRSVGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMS--KSACVCL 221
M+G S L EL CP+C +++K+ +T+K C FC CI + S K C
Sbjct: 1 MDGTEIAVSPRSLHSELMCPICLDMLKN-TMTTKECLHRFCSDCIVTALRSGNKECPTCR 59
Query: 222 ATNILADDLLPNKTLRDAIHRILES 246
+ L P+ I +I S
Sbjct: 60 KKLVSKRSLRPDPNFDALISKIYPS 84
>BMI1_MOUSE (P25916) Polycomb complex protein BMI-1
Length = 324
Score = 37.0 bits (84), Expect = 0.12
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 5/76 (6%)
Query: 173 VGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLATNILADDLLP 232
+ +L P L C LC DA +C SFC CI Y+ + C + L
Sbjct: 9 ITELNPHLMCVLCGGYFIDATTIIEC-LHSFCKTCIVRYLETSKYCPICDVQVHKTRPLL 67
Query: 233 N----KTLRDAIHRIL 244
N KTL+D +++++
Sbjct: 68 NIRSDKTLQDIVYKLV 83
>BMI1_HUMAN (P35226) Polycomb complex protein BMI-1
Length = 326
Score = 37.0 bits (84), Expect = 0.12
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 5/76 (6%)
Query: 173 VGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLATNILADDLLP 232
+ +L P L C LC DA +C SFC CI Y+ + C + L
Sbjct: 9 ITELNPHLMCVLCGGYFIDATTIIEC-LHSFCKTCIVRYLETSKYCPICDVQVHKTRPLL 67
Query: 233 N----KTLRDAIHRIL 244
N KTL+D +++++
Sbjct: 68 NIRSDKTLQDIVYKLV 83
>BMI1_FELCA (Q9TST0) Polycomb complex protein BMI-1
Length = 326
Score = 37.0 bits (84), Expect = 0.12
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 5/76 (6%)
Query: 173 VGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLATNILADDLLP 232
+ +L P L C LC DA +C SFC CI Y+ + C + L
Sbjct: 9 ITELNPHLMCVLCGGYFIDATTIIEC-LHSFCKTCIVRYLETSKYCPICDVQVHKTRPLL 67
Query: 233 N----KTLRDAIHRIL 244
N KTL+D +++++
Sbjct: 68 NIRSDKTLQDIVYKLV 83
>NUCL_XENLA (P20397) Nucleolin (Protein C23)
Length = 650
Score = 36.6 bits (83), Expect = 0.15
Identities = 27/60 (45%), Positives = 31/60 (51%), Gaps = 6/60 (10%)
Query: 33 DAPLTSKADEDSKI---KALID--TPALDWQQQG-SDFGAGRGFRRGAVGGRIGGGRGFG 86
DA +A ED +I K +D P D Q+ G FG G GFR G G GGGRGFG
Sbjct: 552 DAKAAKEAMEDGEIDGNKVTLDFAKPKGDSQRGGRGGFGRGGGFRGGRGGRGGGGGRGFG 611
>ME18_MOUSE (P23798) DNA-binding protein Mel-18 (RING finger protein
110) (Zinc finger protein 144)
Length = 342
Score = 36.6 bits (83), Expect = 0.15
Identities = 20/76 (26%), Positives = 34/76 (44%), Gaps = 5/76 (6%)
Query: 173 VGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLATNILAD---- 228
+ +L P L C LC DA +C SFC CI Y+ + C +
Sbjct: 9 ITELNPHLMCALCGGYFIDATTIVEC-LHSFCKTCIVRYLETNKYCPMCDVQVHKTRPLL 67
Query: 229 DLLPNKTLRDAIHRIL 244
+ +KTL+D +++++
Sbjct: 68 SIRSDKTLQDIVYKLV 83
>ME18_HUMAN (P35227) DNA-binding protein Mel-18 (RING finger protein
110) (Zinc finger protein 144)
Length = 344
Score = 36.6 bits (83), Expect = 0.15
Identities = 20/76 (26%), Positives = 34/76 (44%), Gaps = 5/76 (6%)
Query: 173 VGDLPPELHCPLCNNVMKDAVLTSKCCFKSFCDKCIRNYIMSKSACVCLATNILAD---- 228
+ +L P L C LC DA +C SFC CI Y+ + C +
Sbjct: 9 ITELNPHLMCALCGGYFIDATTIVEC-LHSFCKTCIVRYLETNKYCPMCDVQVHKTRPLL 67
Query: 229 DLLPNKTLRDAIHRIL 244
+ +KTL+D +++++
Sbjct: 68 SIRSDKTLQDIVYKLV 83
>IF38_SCHPO (O14164) Probable eukaryotic translation initiation
factor 3 93 kDa subunit (eIF3 p93)
Length = 918
Score = 36.6 bits (83), Expect = 0.15
Identities = 38/135 (28%), Positives = 62/135 (45%), Gaps = 3/135 (2%)
Query: 245 ESGNSSTEN--AGSTYQVQDMLSSRCPQPKIPSPTSSAT-SKGEPKVSQVNEGMANIQEI 301
ES +SS EN S + QD SS + S +SS + S E S+ +E ++
Sbjct: 15 ESVDSSEENRLTSSRLKKQDDSSSEEESSEEESASSSESESSEEESESEESEVEVPKKKA 74
Query: 302 VAERKEVSATQQVSEQVKIPRAAVVSEVTHESMSVKEPEPASQGSAKLVEEEVQQKLVPT 361
VA ++ + + SE+ + + SEV+ ES S E E S+ ++ EE + +
Sbjct: 75 VAASEDSESDSESSEEEEETESEEDSEVSDESESESESESESEEESESEEESDESERSGP 134
Query: 362 DGGKKKKKKKVRMPA 376
KK +K+ PA
Sbjct: 135 SSFLKKPEKEEAKPA 149
>GSA_MYCTU (P63506) Glutamate-1-semialdehyde 2,1-aminomutase (EC
5.4.3.8) (GSA) (Glutamate-1-semialdehyde
aminotransferase) (GSA-AT)
Length = 462
Score = 36.6 bits (83), Expect = 0.15
Identities = 24/86 (27%), Positives = 38/86 (43%), Gaps = 1/86 (1%)
Query: 268 CPQPKIPSPTSSATSKGEPKVSQVNEGMANIQEIVAERKEVSATQQVSEQVKIPRAAVVS 327
C P+ P+ S +S+G P V G A IV ++ A QQ + AAV++
Sbjct: 177 CDDPQRPASPRSQSSRGLPSSPGVT-GAAAADTIVLPYNDIDAVQQTFARFGEQIAAVIT 235
Query: 328 EVTHESMSVKEPEPASQGSAKLVEEE 353
E + +M V P P + + + E
Sbjct: 236 EASPGNMGVVPPGPGFNAALRAITAE 261
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 59,941,081
Number of Sequences: 164201
Number of extensions: 2612956
Number of successful extensions: 9715
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 115
Number of HSP's that attempted gapping in prelim test: 9503
Number of HSP's gapped (non-prelim): 273
length of query: 498
length of database: 59,974,054
effective HSP length: 114
effective length of query: 384
effective length of database: 41,255,140
effective search space: 15841973760
effective search space used: 15841973760
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC126012.10


