

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.6 + phase: 0
(530 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
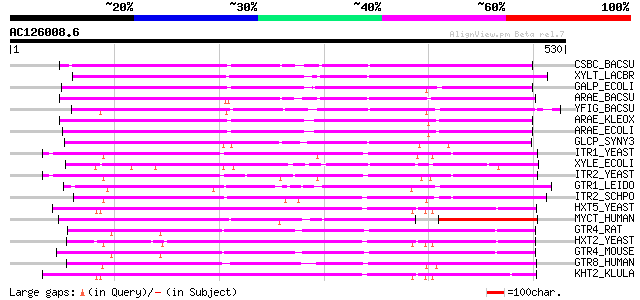
Score E
Sequences producing significant alignments: (bits) Value
CSBC_BACSU (P46333) Probable metabolite transport protein csbC 265 3e-70
XYLT_LACBR (O52733) D-xylose-proton symporter (D-xylose transpor... 257 6e-68
GALP_ECOLI (P37021) Galactose-proton symporter (Galactose transp... 250 7e-66
ARAE_BACSU (P96710) Arabinose-proton symporter (Arabinose transp... 248 2e-65
YFIG_BACSU (P54723) Hypothetical metabolite transport protein yfiG 245 2e-64
ARAE_KLEOX (P45598) Arabinose-proton symporter (Arabinose transp... 233 9e-61
ARAE_ECOLI (P09830) Arabinose-proton symporter (Arabinose transp... 233 9e-61
GLCP_SYNY3 (P15729) Glucose transport protein 227 7e-59
ITR1_YEAST (P30605) Myo-inositol transporter 1 218 3e-56
XYLE_ECOLI (P09098) D-xylose-proton symporter (D-xylose transpor... 217 7e-56
ITR2_YEAST (P30606) Myo-inositol transporter 2 216 1e-55
GTR1_LEIDO (Q01440) Membrane transporter D1 214 3e-55
ITR2_SCHPO (P87110) Myo-inositol transporter 2 214 4e-55
HXT5_YEAST (P38695) Probable glucose transporter HXT5 212 2e-54
MYCT_HUMAN (Q96QE2) Proton myo-inositol cotransporter (H(+)-myo-... 206 1e-52
GTR4_RAT (P19357) Solute carrier family 2, facilitated glucose t... 206 2e-52
HXT2_YEAST (P23585) High-affinity glucose transporter HXT2 205 3e-52
GTR4_MOUSE (P14142) Solute carrier family 2, facilitated glucose... 204 6e-52
GTR8_HUMAN (Q9NY64) Solute carrier family 2, facilitated glucose... 201 4e-51
KHT2_KLULA (P53387) Hexose transporter 2 201 5e-51
>CSBC_BACSU (P46333) Probable metabolite transport protein csbC
Length = 461
Score = 265 bits (676), Expect = 3e-70
Identities = 149/452 (32%), Positives = 250/452 (54%), Gaps = 15/452 (3%)
Query: 48 RNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLL 107
+ TRKY+I F +L +L GYD GV+SGA++FI D+ +T + ++ +L + ++
Sbjct: 2 KKDTRKYMIY--FFGALGGLLYGYDTGVISGALLFINNDIPLTTLTEGLVVSMLLLGAIF 59
Query: 108 GSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISP 167
GS G SD GR+ + + +++F +G + + + +L+ R++ G+ +G + P
Sbjct: 60 GSALSGTCSDRWGRRKVVFVLSIIFIIGALACAFSQTIGMLIASRVILGLAVGGSTALVP 119
Query: 168 IYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFI 227
+Y++E++P RG+L T + I GI+L Y+ NY F+ +WR M+ + +P+V +
Sbjct: 120 VYLSEMAPTKIRGTLGTMNNLMIVTGILLAYIVNYLFTPFE---AWRWMVGLAAVPAVLL 176
Query: 228 GFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKP 287
+ +PESPRWLV + EEAR ++ T+ D K++E LAE++Q G+ E K
Sbjct: 177 LIGIAFMPESPRWLVKRGSEEEARRIMNITH-DPKDIEMELAEMKQ-------GEAEKKE 228
Query: 288 VWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAV 347
+L +R ML+ G+G+ FQQ GI+ +YY+P I AG+ + L T+ +
Sbjct: 229 TTLGVLKAK-WIRPMLLIGVGLAIFQQAVGINTVIYYAPTIFTKAGLGTSASALG-TMGI 286
Query: 348 GITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLVIALGILFVC 407
GI + + A++LID+VGRK LLI ++G+T L + L + ++F+
Sbjct: 287 GILNVIMCITAMILIDRVGRKKLLIWGSVGITLSLAALSGVLLTLGLSASTAWMTVVFLG 346
Query: 408 GNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGG 467
+ F+ GPV WVL E+FP + R A+ + + +V++ F + A+
Sbjct: 347 VYIVFYQATWGPVVWVLMPELFPSKARGAATGFTTLVLSAANLIVSLVFPLMLSAMGIAW 406
Query: 468 TFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
F +FS I L+ F F +VPETKGKSLE+IE
Sbjct: 407 VFMVFSVICLLSFFFAFYMVPETKGKSLEEIE 438
>XYLT_LACBR (O52733) D-xylose-proton symporter (D-xylose
transporter)
Length = 457
Score = 257 bits (656), Expect = 6e-68
Identities = 139/454 (30%), Positives = 248/454 (54%), Gaps = 13/454 (2%)
Query: 61 FASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGGGRTSDIIG 120
F +L +L GYD GV+SGA++FI++ + + Q +++ + + ++LG+ G +SD G
Sbjct: 12 FGALGGLLFGYDTGVISGAILFIQKQMNLGSWQQGWVVSAVLLGAILGAAIIGPSSDRFG 71
Query: 121 RKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEISPNLTRG 180
R+ + L+A++F +G + +P + L+I R++ G+ +G + P Y+AE++P+ RG
Sbjct: 72 RRKLLLLSAIIFFVGALGSAFSPEFWTLIISRIILGMAVGAASALIPTYLAELAPSDKRG 131
Query: 181 SLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFIIPESPRW 240
++++ ++ + GI+L Y++NY+FSG + WR ML +P+ + I+PESPR+
Sbjct: 132 TVSSLFQLMVMTGILLAYITNYSFSGF--YTGWRWMLGFAAIPAALLFLGGLILPESPRF 189
Query: 241 LVMQNRIEEARSVLLKTNE-DEKEVEERLAEIQQAAGFANSGKYEDKPVWRELLSPPPAL 299
LV ++EAR VL N+ D+ V + + +IQ++A + G W EL +
Sbjct: 190 LVKSGHLDEARHVLDTMNKHDQVAVNKEINDIQESAKIVSGG-------WSELFG--KMV 240
Query: 300 RRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAI 359
R LI G+G+ FQQ+ G + +YY+P I G + LL A + +GI + +A+
Sbjct: 241 RPSLIIGIGLAIFQQVMGCNTVLYYAPTIFTDVGFGVSAALL-AHIGIGIFNVIVTAIAV 299
Query: 360 VLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLVIALGILFVCGNVAFFSVGLGP 419
++DK+ RK ++ +GM LF M + + + ++ + +AFFS GP
Sbjct: 300 AIMDKIDRKKIVNIGAVGMGISLFVMSIGMKFSGGSQTAAIISVIALTVYIAFFSATWGP 359
Query: 420 VCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALA 479
V WV+ E+FPL +R ++ +V N + +V+++F S+ D G F + + +
Sbjct: 360 VMWVMIGEVFPLNIRGLGNSFASVINWTANMIVSLTFPSLLDFFGTGSLFIGYGILCFAS 419
Query: 480 IVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEM 513
I FV V ET+ +SLE IE + G E+
Sbjct: 420 IWFVQKKVFETRNRSLEDIEATLRAKTGEDAAEL 453
>GALP_ECOLI (P37021) Galactose-proton symporter (Galactose
transporter)
Length = 464
Score = 250 bits (638), Expect = 7e-66
Identities = 142/454 (31%), Positives = 249/454 (54%), Gaps = 19/454 (4%)
Query: 50 STRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGS 109
S + A+L +L G D+GV++GA+ FI ++ +IT E+++ + + +G+
Sbjct: 10 SNKAMTFFVCFLAALAGLLFGLDIGVIAGALPFIADEFQITSHTQEWVVSSMMFGAAVGA 69
Query: 110 LGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIY 169
+G G S +GRK ++ + A++F G + AP+ +VL++ R+L G+ +G +P+Y
Sbjct: 70 VGSGWLSFKLGRKKSLMIGAILFVAGSLFSAAAPNVEVLILSRVLLGLAVGVASYTAPLY 129
Query: 170 IAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGF 229
++EI+P RGS+ + ++ I +GI+ Y+S+ AFS +WR ML V I+P++ +
Sbjct: 130 LSEIAPEKIRGSMISMYQLMITIGILGAYLSDTAFSYTG---AWRWMLGVIIIPAILLLI 186
Query: 230 ALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVW 289
+F +P+SPRW + R +A VLL+ + E + L EI+++ SG W
Sbjct: 187 GVFFLPDSPRWFAAKRRFVDAERVLLRLRDTSAEAKRELDEIRESLQVKQSG-------W 239
Query: 290 RELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGI 349
L RR + G+ +Q QQ +G++ +YY+P+I AG + ++ + TV VG+
Sbjct: 240 A-LFKENSNFRRAVFLGVLLQVMQQFTGMNVIMYYAPKIFELAGYTNTTEQMWGTVIVGL 298
Query: 350 TKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGP----LVIALGILF 405
T + +AI L+D+ GRKP L + M A + +G + + P IA+ ++F
Sbjct: 299 TNVLATFIAIGLVDRWGRKPTLTLGFLVMAAGMGVLGTMMHIGIHSPSAQYFAIAMLLMF 358
Query: 406 VCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISF 465
+ G F++ GP+ WVL SEI PL+ R N + + +V +FL++ + +
Sbjct: 359 IVG----FAMSAGPLIWVLCSEIQPLKGRDFGITCSTATNWIANMIVGATFLTMLNTLGN 414
Query: 466 GGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
TF++++A++ L I+ LVPETK SLE IE
Sbjct: 415 ANTFWVYAALNVLFILLTLWLVPETKHVSLEHIE 448
>ARAE_BACSU (P96710) Arabinose-proton symporter (Arabinose
transporter)
Length = 464
Score = 248 bits (634), Expect = 2e-65
Identities = 152/460 (33%), Positives = 242/460 (52%), Gaps = 18/460 (3%)
Query: 48 RNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLL 107
R+ + +VI + A L +L GYD V+SGA+ F+K+ ++ +I + I ++
Sbjct: 16 RSHSMGFVILISCAAGLGGLLYGYDTAVISGAIGFLKDLYSLSPFMEGLVISSIMIGGVV 75
Query: 108 GSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISP 167
G G SD GR+ + AA++F + I L+ L+I R++ G+GIG G +S
Sbjct: 76 GVGISGFLSDRFGRRKILMTAALLFAISAIVSALSQDVSTLIIARIIGGLGIGMGSSLSV 135
Query: 168 IYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAF--SGL---SVHISWRVMLAVGIL 222
YI E +P RGSL++ ++F +GI Y N A SG VH WR MLA G++
Sbjct: 136 TYITEAAPPAIRGSLSSLYQLFTILGISATYFINLAVQRSGTYEWGVHTGWRWMLAYGMV 195
Query: 223 PSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGK 282
PSV L ++PESPRWL + EA +L + N E +E L I+ NS K
Sbjct: 196 PSVIFFLVLLVVPESPRWLAKAGKTNEALKILTRIN-GETVAKEELKNIE------NSLK 248
Query: 283 YEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLA 342
E +L P LR+ L+ G+ + F Q+ G++A YY PEI G + +
Sbjct: 249 IEQMGSLSQLFK--PGLRKALVIGILLALFNQVIGMNAITYYGPEIFKMMGFGQNAGFV- 305
Query: 343 ATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLVIALG 402
T VG+ + +F ++A++LIDKVGRK L+ + M + +G + +++
Sbjct: 306 TTCIVGVVEVIFTVIAVLLIDKVGRKKLMSIGSAFMAIFMILIGTSFYFELTSGIMM--- 362
Query: 403 ILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDA 462
I+ + G VA F V +GP+ W++ SEIFP +RA+A+ + + + + + D+
Sbjct: 363 IVLILGFVAAFCVSVGPITWIMISEIFPNHLRARAAGIATIFLWGANWAIGQFVPMMIDS 422
Query: 463 ISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
TF++F+ I+ L +FV T+ PETK KSLE+IE ++
Sbjct: 423 FGLAYTFWIFAVINILCFLFVVTICPETKNKSLEEIEKLW 462
>YFIG_BACSU (P54723) Hypothetical metabolite transport protein yfiG
Length = 482
Score = 245 bits (626), Expect = 2e-64
Identities = 157/477 (32%), Positives = 255/477 (52%), Gaps = 34/477 (7%)
Query: 60 IFASLNNVLLGYDVGVMSGAVIFIKE--DLKITEVQVEFLIGILSIVSLLGSLGGGRTSD 117
+ ++ +L GYD GV++GA+ F+ L +T V + L + + G++ GGR SD
Sbjct: 26 LVSTFGGLLFGYDTGVINGALPFMATAGQLNLTPVTEGLVASSLLLGAAFGAMFGGRLSD 85
Query: 118 IIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEISPNL 177
GR+ T+ A++F + T +P+ V++ R L G+ +G + P ++AEISP
Sbjct: 86 RHGRRKTILYLALLFIAATLGCTFSPNASVMIAFRFLLGLAVGCASVTVPTFLAEISPAE 145
Query: 178 TRGSLTTFPEIFINVGIMLGYVSNYAFS---GLSVHISWRVMLAVGILPSVFIGFALFII 234
RG + T E+ I +G +L Y N G S ++ WR ML + LP+V + F + I+
Sbjct: 146 RRGRIVTQNELMIVIGQLLAYTFNAIIGSTMGESANV-WRYMLVIATLPAVVLWFGMLIV 204
Query: 235 PESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAA-GFANSGKYEDKPVWRELL 293
PESPRWL + R+ +A VL + ED + ++ + EI+ A G A + D
Sbjct: 205 PESPRWLAAKGRMGDALRVLRQIREDS-QAQQEIKEIKHAIEGTAKKAGFHD-------- 255
Query: 294 SPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTV 353
P +RR+L G+GI QQI+G+++ +YY EIL AG + ++ L+ +A G+ +
Sbjct: 256 FQEPWIRRILFIGIGIAIVQQITGVNSIMYYGTEILREAGFQTEAALIG-NIANGVISVI 314
Query: 354 FILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGP----LVIALGILFVCGN 409
++ I L+ KV R+P+LI IG L +G+ + E P +V++L ILF+
Sbjct: 315 AVIFGIWLLGKVRRRPMLIIGQIGTMTALLLIGILSIVLEGTPALPYVVLSLTILFL--- 371
Query: 410 VAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTF 469
AF + V W++ SEIFP+ VR + + L+ +F + + I TF
Sbjct: 372 -AFQQTAISTVTWLMLSEIFPMHVRGLGMGISTFCLWTANFLIGFTFPILLNHIGMSATF 430
Query: 470 FLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEMELGDVEQLVQNKT 526
F+F A++ LAI+FV VPETKG+SLEQ+E F ++G + +Q +QN+T
Sbjct: 431 FIFVAMNILAILFVKKYVPETKGRSLEQLEHSF-RQYGRR--------ADQEIQNQT 478
>ARAE_KLEOX (P45598) Arabinose-proton symporter (Arabinose
transporter)
Length = 472
Score = 233 bits (594), Expect = 9e-61
Identities = 134/457 (29%), Positives = 242/457 (52%), Gaps = 18/457 (3%)
Query: 48 RNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLL 107
+ TR+ +I A++ +L G D+GV++GA+ FI + ++ E+++ + + + +
Sbjct: 15 QRDTRRMNQFVSIAAAVAGLLFGLDIGVIAGALPFITDHFVLSSRLQEWVVSSMMLGAAI 74
Query: 108 GSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISP 167
G+L G S +GRK+++ + AV+F G + A S ++L++ R++ G+ +G +P
Sbjct: 75 GALFNGWLSFRLGRKYSLMVGAVLFVAGSVGSAFATSVEMLLVARIVLGVAVGIASYTAP 134
Query: 168 IYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFI 227
+Y++E++ RG + + ++ + +GI++ ++S+ AFS +WR ML V LP+V +
Sbjct: 135 LYLSEMASENVRGKMISMYQLMVTLGIVMAFLSDTAFSYSG---NWRAMLGVLALPAVVL 191
Query: 228 GFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKP 287
+ +P SPRWL + R EA VL + ++ + L EI+++ G
Sbjct: 192 IILVIFLPNSPRWLAEKGRHVEAEEVLRMLRDTSEKARDELNEIRESLKLKQGG------ 245
Query: 288 VWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAV 347
W L +RR + G+ +Q QQ +G++ +YY+P I AG + + AT+ V
Sbjct: 246 -W-ALFKVNRNVRRAVFLGMLLQAMQQFTGMNIIMYYAPRIFKMAGFTTTEQQMVATLVV 303
Query: 348 GITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLV-----IALG 402
G+T +A+ +DK GRKP L M +G L F+ G +++G
Sbjct: 304 GLTFMFATFIAVFTVDKAGRKPALKIGFSVMAIGTLVLGYCLMQFDNGTASSGLSWLSVG 363
Query: 403 ILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDA 462
+ +C +A +++ PV W+L SEI PL+ R N V + ++ +FL++ DA
Sbjct: 364 MTMMC--IAGYAMSAAPVVWILCSEIQPLKCRDFGITCSTTTNWVSNMIIGATFLTLLDA 421
Query: 463 ISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
I GTF+L++A++ I F L+PETK +LE IE
Sbjct: 422 IGAAGTFWLYTALNVAFIGVTFWLIPETKNVTLEHIE 458
>ARAE_ECOLI (P09830) Arabinose-proton symporter (Arabinose
transporter)
Length = 472
Score = 233 bits (594), Expect = 9e-61
Identities = 135/454 (29%), Positives = 241/454 (52%), Gaps = 18/454 (3%)
Query: 51 TRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSL 110
TR+ + ++ A++ +L G D+GV++GA+ FI + +T E+++ + + + +G+L
Sbjct: 18 TRRMNMFVSVAAAVAGLLFGLDIGVIAGALPFITDHFVLTSRLQEWVVSSMMLGAAIGAL 77
Query: 111 GGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYI 170
G S +GRK+++ A++F +G I A S ++L+ R++ GI +G +P+Y+
Sbjct: 78 FNGWLSFRLGRKYSLMAGAILFVLGSIGSAFATSVEMLIAARVVLGIAVGIASYTAPLYL 137
Query: 171 AEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFA 230
+E++ RG + + ++ + +GI+L ++S+ AFS +WR ML V LP+V +
Sbjct: 138 SEMASENVRGKMISMYQLMVTLGIVLAFLSDTAFSYSG---NWRAMLGVLALPAVLLIIL 194
Query: 231 LFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWR 290
+ +P SPRWL + R EA VL + ++ E L EI+++ G W
Sbjct: 195 VVFLPNSPRWLAEKGRHIEAEEVLRMLRDTSEKAREELNEIRESLKLKQGG-------W- 246
Query: 291 ELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGIT 350
L +RR + G+ +Q QQ +G++ +YY+P I AG + + AT+ VG+T
Sbjct: 247 ALFKINRNVRRAVFLGMLLQAMQQFTGMNIIMYYAPRIFKMAGFTTTEQQMIATLVVGLT 306
Query: 351 KTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGPLV-----IALGILF 405
+A+ +DK GRKP L M +G L F+ G +++G+
Sbjct: 307 FMFATFIAVFTVDKAGRKPALKIGFSVMALGTLVLGYCLMQFDNGTASSGLSWLSVGMTM 366
Query: 406 VCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISF 465
+C +A +++ PV W+L SEI PL+ R N V + ++ +FL++ D+I
Sbjct: 367 MC--IAGYAMSAAPVVWILCSEIQPLKCRDFGITCSTTTNWVSNMIIGATFLTLLDSIGA 424
Query: 466 GGTFFLFSAISALAIVFVFTLVPETKGKSLEQIE 499
GTF+L++A++ + F L+PETK +LE IE
Sbjct: 425 AGTFWLYTALNIAFVGITFWLIPETKNVTLEHIE 458
>GLCP_SYNY3 (P15729) Glucose transport protein
Length = 468
Score = 227 bits (578), Expect = 7e-59
Identities = 151/463 (32%), Positives = 243/463 (51%), Gaps = 26/463 (5%)
Query: 53 KYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSLLGSLGG 112
K+V+ + A+L L G+D V++GAV +++ + + + + + S LG+ G
Sbjct: 15 KFVLLISGVAALGGFLFGFDTAVINGAVAALQKHFQTDSLLTGLSVSLALLGSALGAFGA 74
Query: 113 GRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAE 172
G +D GR TM LAAV+F + I L + + R+L GIG+G +I+P YIAE
Sbjct: 75 GPIADRHGRIKTMILAAVLFTLSSIGSGLPFTIWDFIFWRVLGGIGVGAASVIAPAYIAE 134
Query: 173 ISPNLTRGSLTTFPEIFINVGIMLGYVSNY---AFSGLSVH-------ISWRVMLAVGIL 222
+SP RG L + ++ I GI + +SN+ +G S +WR M ++
Sbjct: 135 VSPAHLRGRLGSLQQLAIVSGIFIALLSNWFIALMAGGSAQNPWLFGAAAWRWMFWTELI 194
Query: 223 PSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGK 282
P++ G F+IPESPR+LV Q + E+A ++L K + +V R+ EIQ
Sbjct: 195 PALLYGVCAFLIPESPRYLVAQGQGEKAAAILWKV--EGGDVPSRIEEIQATVSL----- 247
Query: 283 YEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLA 342
+ KP + +LLS L ++ G+G+ QQ GI+ YYS + + G ++ LL
Sbjct: 248 -DHKPRFSDLLSRRGGLLPIVWIGMGLSALQQFVGINVIFYYSSVLWRSVGFTEEKSLL- 305
Query: 343 ATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLS----LFEKGPLV 398
TV G + LVAI +DK GRKPLL+ +IGMT L + V + + L
Sbjct: 306 ITVITGFINILTTLVAIAFVDKFGRKPLLLMGSIGMTITLGILSVVFGGATVVNGQPTLT 365
Query: 399 IALGIL-FVCGNVAFFSVGL--GPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMS 455
A GI+ V N+ FS G GP+ WVL E+F ++RA A ++ A + + +++ +
Sbjct: 366 GAAGIIALVTANLYVFSFGFSWGPIVWVLLGEMFNNKIRAAALSVAAGVQWIANFIISTT 425
Query: 456 FLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQI 498
F + D + G + L++ +A++I F++ V ETKGK+LEQ+
Sbjct: 426 FPPLLDTVGLGPAYGLYATSAAISIFFIWFFVKETKGKTLEQM 468
>ITR1_YEAST (P30605) Myo-inositol transporter 1
Length = 584
Score = 218 bits (555), Expect = 3e-56
Identities = 143/493 (29%), Positives = 253/493 (51%), Gaps = 30/493 (6%)
Query: 32 VDDNDDVLHQQQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLK--- 88
V+D DD + S ++I AS++ + GYD G +S A+I I DL
Sbjct: 66 VNDEDDT---SVMITFNQSLSPFIITLTFVASISGFMFGYDTGYISSALISIGTDLDHKV 122
Query: 89 ITEVQVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVL 148
+T + E + S+ +L+ S+ G +DI GRK + + ++F +G I A ++ +
Sbjct: 123 LTYGEKEIVTAATSLGALITSIFAGTAADIFGRKRCLMGSNLMFVIGAILQVSAHTFWQM 182
Query: 149 MIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLS 208
+GRL+ G G+G G +I+P++I+EI+P + RG LT +++ G ++ Y +GL+
Sbjct: 183 AVGRLIMGFGVGIGSLIAPLFISEIAPKMIRGRLTVINSLWLTGGQLVAYGCG---AGLN 239
Query: 209 -VHISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNED-EKEVEE 266
V+ WR+++ + ++P+ L +P++PR+ VM+ + A VL ++ D +E+ E
Sbjct: 240 YVNNGWRILVGLSLIPTAVQFTCLCFLPDTPRYYVMKGDLARATEVLKRSYTDTSEEIIE 299
Query: 267 RLAEIQQAAGFANSGKYEDKPVWREL--LSPPPALRRMLITGLGIQCFQQISGIDATVYY 324
R E + GK + VW + L P+ R LI G G+Q QQ +G ++ +Y+
Sbjct: 300 RKVEELVTLNQSIPGKNVPEKVWNTIKELHTVPSNLRALIIGCGLQAIQQFTGWNSLMYF 359
Query: 325 SPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFC 384
S I G ++ S A ++ V T +F LVA IDK+GR+ +L+ GMT L
Sbjct: 360 SGTIFETVGFKNSS---AVSIIVSGTNFIFTLVAFFSIDKIGRRTILLIGLPGMTMALVV 416
Query: 385 MGVT---LSLFEKGPLVIALG----------ILFVCGNVAFFSVGLGPVCWVLTSEIFPL 431
+ L + G + + + I+F+ AF+++G+G V W SE+FP
Sbjct: 417 CSIAFHFLGIKFDGAVAVVVSSGFSSWGIVIIVFIIVFAAFYALGIGTVPW-QQSELFPQ 475
Query: 432 RVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETK 491
VR ++ N S ++A +FL++ I+ GTF F+ +S L+ +F + PE
Sbjct: 476 NVRGIGTSYATATNWAGSLVIASTFLTMLQNITPAGTFAFFAGLSCLSTIFCYFCYPELS 535
Query: 492 GKSLEQIEMMFEN 504
G LE+++ + ++
Sbjct: 536 GLELEEVQTILKD 548
>XYLE_ECOLI (P09098) D-xylose-proton symporter (D-xylose
transporter)
Length = 491
Score = 217 bits (552), Expect = 7e-56
Identities = 152/491 (30%), Positives = 255/491 (50%), Gaps = 57/491 (11%)
Query: 54 YVIACAIFASLNNVLLGYDVGVMSGAV-----IFIKEDLKITEVQVEFLIGILSIVSLLG 108
Y+ + + A+L +L GYD V+SG V +F+ ++E L+G +L+G
Sbjct: 9 YIFSITLVATLGGLLFGYDTAVISGTVESLNTVFVAPQ-NLSESAANSLLGFCVASALIG 67
Query: 109 SLGGGRT----SDIIGRKWTMALAAVVFQMGGI------------------TMTLAPSYQ 146
+ GG S+ GR+ ++ +AAV+F + G+ + LA
Sbjct: 68 CIIGGALGGYCSNRFGRRDSLKIAAVLFFISGVGSAWPELGFTSINPDNTVPVYLAGYVP 127
Query: 147 VLMIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNY--AF 204
+I R++ GIG+G M+SP+YIAE++P RG L +F + I G +L Y NY A
Sbjct: 128 EFVIYRIIGGIGVGLASMLSPMYIAELAPAHIRGKLVSFNQFAIIFGQLLVYCVNYFIAR 187
Query: 205 SGLSVHIS---WRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDE 261
SG + ++ WR M A +P++ L+ +PESPRWL+ + + E+A +L K +
Sbjct: 188 SGDASWLNTDGWRYMFASECIPALLFLMLLYTVPESPRWLMSRGKQEQAEGILRKIMGNT 247
Query: 262 KEVEERLAEIQQAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDAT 321
+ +Q+ + G+ K R L+ +++ G+ + FQQ GI+
Sbjct: 248 LATQA----VQEIKHSLDHGR---KTGGRLLMFGVG----VIVIGVMLSIFQQFVGINVV 296
Query: 322 VYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTAC 381
+YY+PE+ G LL T+ VG+ F ++AI+ +DK GRKPL I +GM
Sbjct: 297 LYYAPEVFKTLGASTDIALLQ-TIIVGVINLTFTVLAIMTVDKFGRKPLQIIGALGMAIG 355
Query: 382 LFCMGVTLSLFEKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALG 441
+F +G + + + P ++AL L + VA F++ GPVCWVL SEIFP +R +A A+
Sbjct: 356 MFSLGT--AFYTQAPGIVAL--LSMLFYVAAFAMSWGPVCWVLLSEIFPNAIRGKALAIA 411
Query: 442 AVANRVCSGLVAMSFLSVSDAISF-------GGTFFLFSAISALAIVFVFTLVPETKGKS 494
A + + V+ +F + D S+ G +++++ + LA +F++ VPETKGK+
Sbjct: 412 VAAQWLANYFVSWTF-PMMDKNSWLVAHFHNGFSYWIYGCMGVLAALFMWKFVPETKGKT 470
Query: 495 LEQIEMMFENE 505
LE++E ++E E
Sbjct: 471 LEELEALWEPE 481
>ITR2_YEAST (P30606) Myo-inositol transporter 2
Length = 612
Score = 216 bits (550), Expect = 1e-55
Identities = 141/495 (28%), Positives = 259/495 (51%), Gaps = 34/495 (6%)
Query: 32 VDDNDDVLHQQQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLK--- 88
V+D DD + S ++I AS++ + GYD G +S A+I I DL
Sbjct: 92 VNDEDDT---SVIITFNQSISPFIITLTFVASISGFMFGYDTGYISSALISINRDLDNKV 148
Query: 89 ITEVQVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVL 148
+T + E + S+ +L+ S+G G +D+ GR+ + + ++F +G I A + +
Sbjct: 149 LTYGEKELITAATSLGALITSVGAGTAADVFGRRPCLMFSNLMFLIGAILQITAHKFWQM 208
Query: 149 MIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLS 208
GRL+ G G+G G +ISP++I+EI+P + RG LT +++ G ++ Y +GL+
Sbjct: 209 AAGRLIMGFGVGIGSLISPLFISEIAPKMIRGRLTVINSLWLTGGQLIAYGCG---AGLN 265
Query: 209 -VHISWRVMLAVGILPSVFIGFALF-IIPESPRWLVMQNRIEEARSVLLKT--NEDEKEV 264
V WR+++ + ++P+V + F+ F +P++PR+ VM+ ++ A+ VL ++ N +++ +
Sbjct: 266 HVKNGWRILVGLSLIPTV-LQFSFFCFLPDTPRYYVMKGDLKRAKMVLKRSYVNTEDEII 324
Query: 265 EERLAEIQQAAGFANSGKYEDKPVWREL--LSPPPALRRMLITGLGIQCFQQISGIDATV 322
++++ E+ + + GK W + L P+ R LI G G+Q QQ +G ++ +
Sbjct: 325 DQKVEEL-SSLNQSIPGKNPITKFWNMVKELHTVPSNFRALIIGCGLQAIQQFTGWNSLM 383
Query: 323 YYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACL 382
Y+S I G ++ S A ++ V T VF L+A IDK+GR+ +L+ GMT L
Sbjct: 384 YFSGTIFETVGFKNSS---AVSIIVSGTNFVFTLIAFFCIDKIGRRYILLIGLPGMTVAL 440
Query: 383 FCMGVTLSL----FEKGPLVIA---------LGILFVCGNVAFFSVGLGPVCWVLTSEIF 429
+ F V+A + I+F+ AF+++G+G V W SE+F
Sbjct: 441 VICAIAFHFLGIKFNGADAVVASDGFSSWGIVIIVFIIVYAAFYALGIGTVPW-QQSELF 499
Query: 430 PLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPE 489
P VR ++ N S ++A +FL++ I+ GTF F+ ++ L+ +F + PE
Sbjct: 500 PQNVRGVGTSYATATNWAGSLVIASTFLTMLQNITPTGTFSFFAGVACLSTIFCYFCYPE 559
Query: 490 TKGKSLEQIEMMFEN 504
G LE+++ + ++
Sbjct: 560 LSGLELEEVQTILKD 574
>GTR1_LEIDO (Q01440) Membrane transporter D1
Length = 547
Score = 214 bits (546), Expect = 3e-55
Identities = 146/478 (30%), Positives = 242/478 (50%), Gaps = 34/478 (7%)
Query: 52 RKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEV--QVEFLIGILSIVSLLGS 109
R V+ CA +L L GYD GV++ A+ +K+ +E Q ++ I + +G+
Sbjct: 2 RASVMLCA---ALGGFLFGYDTGVINAALFQMKDHFGFSEHSWQYALIVAIAIAGAFVGA 58
Query: 110 LGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIY 169
G S GR+ +A+A +F +G + M AP+ +V+++ R++ G+ IG P+Y
Sbjct: 59 FISGFISAAFGRRPCIAVADALFVIGSVLMGAAPNVEVVLVSRVIVGLAIGISSATIPVY 118
Query: 170 IAEISPNLTRGSLTTFPEIFINVG--IMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFI 227
+AE++ RG+ +F+ G + G+ + S +I WRV + +G LP+V
Sbjct: 119 LAEVTSPKHRGATIVLNNLFLTGGQFVAAGFTAIMVVF-TSKNIGWRVAIGIGALPAVVQ 177
Query: 228 GFAL-FIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDK 286
F L F +PESPRWL+ + + A++V D+ EV+ L E Q+ S + + +
Sbjct: 178 AFCLLFFLPESPRWLLSKGHADRAKAVA-----DKFEVD--LCEFQEGDELP-SVRIDYR 229
Query: 287 PVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVA 346
P+ +R ++ G+Q QQ SGI+ +YYS IL AG D + ++
Sbjct: 230 PLMAR------DMRFRVVLSSGLQIIQQFSGINTIMYYSSVILYDAGFRDAIMPVVLSIP 283
Query: 347 VGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACL-------FCMGVTLSLFEKGPLVI 399
+ +F VAI +D+ GR+ +L+ S G L F +G +S G L +
Sbjct: 284 LAFMNALFTAVAIFTVDRFGRRRMLLISVFGCLVLLVVIAIIGFFIGTRISYSVGGGLFL 343
Query: 400 ALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSV 459
AL +F+ A ++ G+G + WV+ EIFP +R A+++ +AN + LV+ F +
Sbjct: 344 ALLAVFL----ALYAPGIGCIPWVIMGEIFPTHLRTSAASVATMANWGANVLVSQVFPIL 399
Query: 460 SDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKEMELGD 517
AI GGTF + S + AL +FV+ ETKG +LEQI+ MF G + E G+
Sbjct: 400 MGAIGVGGTFTIISGLMALGCIFVYFFAVETKGLTLEQIDNMFRKRAGLPPRFHEEGE 457
>ITR2_SCHPO (P87110) Myo-inositol transporter 2
Length = 557
Score = 214 bits (545), Expect = 4e-55
Identities = 138/467 (29%), Positives = 240/467 (50%), Gaps = 26/467 (5%)
Query: 62 ASLNNVLLGYDVGVMSGAVIFIKEDLK--ITEVQVEFLIGILSIVSLLGSLGGGRTSDII 119
A ++ +L GYD GV+SGA+ + DL ++ Q E + S +L+ + G +D +
Sbjct: 88 AGISGLLFGYDTGVISGALAVLGSDLGHVLSSGQKELITSATSFAALISATTSGWLADWV 147
Query: 120 GRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYIAEISPNLTR 179
GRK + A +F +G + M + + ++++GR + G GIG +I P+YI E++P R
Sbjct: 148 GRKRLLLCADAIFVIGSVIMAASRNVAMMVVGRFIVGYGIGLTSLIVPMYITELAPARLR 207
Query: 180 GSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFALFIIPESPR 239
G L +FI G ++ Y N AF VH WR+M +G P++ +LF PESPR
Sbjct: 208 GRLVIIYVVFITGGQLIAYSLNAAFE--HVHQGWRIMFGIGAAPALGQLISLFWTPESPR 265
Query: 240 WLVMQNRIEEARSVLLKTNEDEK--EVEERLAEIQQA--AGFANSGKYEDKPVWRELLSP 295
+L+ N +E+ +L + + + K E+ +++ IQ+ F K++ ++L
Sbjct: 266 YLLRHNHVEKVYKILSRIHPEAKPAEIAYKVSLIQEGVKVDFPEGNKFQHFFHSLKVLFT 325
Query: 296 PPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFI 355
P+ RR L G +Q FQQ SG +A Y+S I + G ++ ++ ++ VG T VF
Sbjct: 326 VPSNRRSLFIGCFLQWFQQFSGTNAIQYFSAIIFQSVGFKNS---ISVSIVVGATNFVFT 382
Query: 356 LVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGP----------LVIALGILF 405
+VA + ID++GR+ +L+ ++ M A L + +V+A I+F
Sbjct: 383 IVAFMFIDRIGRRRILLCTSAVMIAGLALCAIAYHFLPADTTQNTNSGWQYVVLASIIIF 442
Query: 406 VCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISF 465
+A ++ G+G + W +E+FP+ VRA + N V + +++ SFL++ ++I+
Sbjct: 443 ----LASYASGIGNIPW-QQAELFPMEVRALGAGFSTAINWVGNLIISASFLTMMESITP 497
Query: 466 GGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFENEHGSQGKE 512
GTF LF+ + +V + PE G S+E I + E KE
Sbjct: 498 TGTFALFAGFCFVGLVTSYFTYPELAGMSIENIHKLLEKGFWQAVKE 544
>HXT5_YEAST (P38695) Probable glucose transporter HXT5
Length = 592
Score = 212 bits (539), Expect = 2e-54
Identities = 137/483 (28%), Positives = 239/483 (49%), Gaps = 26/483 (5%)
Query: 42 QQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVI---FIKE--------DLKIT 90
+QLE K S +V C + + + G+D G +SG V FI+ ++
Sbjct: 73 KQLEKKSKSDLLFVSVCCLMVAFGGFVFGWDTGTISGFVRQTDFIRRFGSTRANGTTYLS 132
Query: 91 EVQVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLA-PSYQVLM 149
+V+ ++ I +I +G + + D+ GRK + V++ +G I + +
Sbjct: 133 DVRTGLMVSIFNIGCAIGGIVLSKLGDMYGRKIGLMTVVVIYSIGIIIQIASIDKWYQYF 192
Query: 150 IGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSV 209
IGR+++G+G+G +++P+ I+E+SP RG+L + ++ I GI LGY +N+ S
Sbjct: 193 IGRIISGLGVGGITVLAPMLISEVSPKQLRGTLVSCYQLMITFGIFLGYCTNFGTKNYSN 252
Query: 210 HISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLA 269
+ WRV L + S+F+ + +PESPR+LV +IEEA+ L + N+ ++
Sbjct: 253 SVQWRVPLGLCFAWSIFMIVGMTFVPESPRYLVEVGKIEEAKRSLARANKTTEDSPLVTL 312
Query: 270 EIQQAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEIL 329
E++ + + W EL++ P + R + G+ IQ QQ++G + YY I
Sbjct: 313 EMENYQSSIEAERLAGSASWGELVTGKPQMFRRTLMGMMIQSLQQLTGDNYFFYYGTTIF 372
Query: 330 MAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLF---CMG 386
A G+ED + +G+ V ++ +D+ GR+ L+ +GM C +G
Sbjct: 373 QAVGLEDS---FETAIVLGVVNFVSTFFSLYTVDRFGRRNCLLWGCVGMICCYVVYASVG 429
Query: 387 VTLSLFEKG---PLVIALG---ILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASAL 440
VT L+ G P G I+F C + F+ PV +VL SE +PLRVR +A ++
Sbjct: 430 VT-RLWPNGQDQPSSKGAGNCMIVFACFYIFCFATTWAPVAYVLISESYPLRVRGKAMSI 488
Query: 441 GAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEM 500
+ N + L++ ++ AI+F ++F A +VF VPETKG +LE++
Sbjct: 489 ASACNWIWGFLISFFTPFITSAINF-YYGYVFMGCMVFAYFYVFFFVPETKGLTLEEVNE 547
Query: 501 MFE 503
M+E
Sbjct: 548 MYE 550
>MYCT_HUMAN (Q96QE2) Proton myo-inositol cotransporter
(H(+)-myo-inositol cotransporter) (Hmit)
Length = 629
Score = 206 bits (524), Expect = 1e-52
Identities = 118/347 (34%), Positives = 184/347 (53%), Gaps = 15/347 (4%)
Query: 47 KRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVEFLIGILSIVSL 106
+++ T +V A+F++L L GYD GV+SGA++ +K L + + E L+ +
Sbjct: 54 QQDETPAFVYVVAVFSALGGFLFGYDTGVVSGAMLLLKRQLSLDALWQELLVSSTVGAAA 113
Query: 107 LGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMIS 166
+ +L GG + + GR+ + LA+ +F G + A + + L+ GRL+ G+GIG M
Sbjct: 114 VSALAGGALNGVFGRRAAILLASALFTAGSAVLAAANNKETLLAGRLVVGLGIGIASMTV 173
Query: 167 PIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVF 226
P+YIAE+SP RG L T +FI G V + AFS L WR ML + +P+V
Sbjct: 174 PVYIAEVSPPNLRGRLVTINTLFITGGQFFASVVDGAFSYLQKD-GWRYMLGLAXVPAVI 232
Query: 227 IGFALFIIPESPRWLVMQNRIEEARSVLLK------TNEDEKEVEERLAEIQQAAGFANS 280
F +PESPRWL+ + + ++AR +L + +E+ ++ + E ++ G A
Sbjct: 233 QFFGFLFLPESPRWLIQKGQTQKARRILSQMRGNQTIDEEYDSIKNNIEEEEKEVGSAG- 291
Query: 281 GKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKL 340
PV +LS PP RR LI G G+Q FQQ+SGI+ +YYS IL +G+ED
Sbjct: 292 ------PVICRMLSYPPT-RRALIVGCGLQMFQQLSGINTIMYYSATILQMSGVEDDRLA 344
Query: 341 LAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGV 387
+ T +F LV + L++KVGR+ L S G T L + +
Sbjct: 345 IWLASVTAFTNFIFTLVGVWLVEKVGRRKLTFGSLAGTTVALIILAL 391
Score = 86.7 bits (213), Expect = 1e-16
Identities = 36/95 (37%), Positives = 67/95 (69%)
Query: 410 VAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTF 469
+ FF+ G+GP+ W + SEI+PL R+ +A + N + + LV+++FL ++ +++ G F
Sbjct: 498 LVFFAPGMGPMPWTVNSEIYPLWARSTGNACSSGINWIFNVLVSLTFLHTAEYLTYYGAF 557
Query: 470 FLFSAISALAIVFVFTLVPETKGKSLEQIEMMFEN 504
FL++ +A+ ++F++ +PETKGK LE+IE +F+N
Sbjct: 558 FLYAGFAAVGLLFIYGCLPETKGKKLEEIESLFDN 592
>GTR4_RAT (P19357) Solute carrier family 2, facilitated glucose
transporter, member 4 (Glucose transporter type 4,
insulin-responsive)
Length = 509
Score = 206 bits (523), Expect = 2e-52
Identities = 144/470 (30%), Positives = 235/470 (49%), Gaps = 33/470 (7%)
Query: 56 IACAIFAS-LNNVLLGYDVGVMSGAVIFIKEDLKITEVQVE------------------F 96
+ A+F++ L ++ GY++GV++ I++ T + +
Sbjct: 24 LVLAVFSAVLGSLQFGYNIGVINAPQKVIEQSYNATWLGRQGPGGPDSIPQGTLTTLWAL 83
Query: 97 LIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLA---PSYQVLMIGRL 153
+ I S+ ++ S G S +GRK M V+ +GG M LA SY++L++GR
Sbjct: 84 SVAIFSVGGMISSFLIGIISQWLGRKRAMLANNVLAVLGGALMGLANAAASYEILILGRF 143
Query: 154 LAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISW 213
L G G + P+Y+ EI+P RG+L T ++ I +GI++ V S L W
Sbjct: 144 LIGAYSGLTSGLVPMYVGEIAPTHLRGALGTLNQLAIVIGILVAQVLGLE-SMLGTATLW 202
Query: 214 RVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQ 273
++LA+ +LP++ L PESPR+L + +E LK +V + LAE++
Sbjct: 203 PLLLAITVLPALLQLLLLPFCPESPRYLYIIRNLEGPARKSLKRLTGWADVSDALAELKD 262
Query: 274 AAGFANSGKYE-DKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAA 332
K E ++P+ L R+ LI + +Q QQ+SGI+A YYS I A
Sbjct: 263 -----EKRKLERERPLSLLQLLGSRTHRQPLIIAVVLQLSQQLSGINAVFYYSTSIFELA 317
Query: 333 GIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLF 392
G+E + AT+ G+ TVF LV+++L+++ GR+ L + GM C M V L L
Sbjct: 318 GVEQPAY---ATIGAGVVNTVFTLVSVLLVERAGRRTLHLLGLAGMCGCAILMTVALLLL 374
Query: 393 EKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCSGLV 452
E+ P + + I+ + G VAFF +G GP+ W + +E+F R A A+ +N C+ +V
Sbjct: 375 ERVPSMSYVSIVAIFGFVAFFEIGPGPIPWFIVAELFSQGPRPAAMAVAGFSNWTCNFIV 434
Query: 453 AMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
M F V+DA+ F LF+ + +F F VPET+G++ +QI F
Sbjct: 435 GMGFQYVADAMG-PYVFLLFAVLLLGFFIFTFLRVPETRGRTFDQISATF 483
>HXT2_YEAST (P23585) High-affinity glucose transporter HXT2
Length = 541
Score = 205 bits (521), Expect = 3e-52
Identities = 141/473 (29%), Positives = 237/473 (49%), Gaps = 34/473 (7%)
Query: 55 VIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLK-------------ITEVQVEFLIGIL 101
VI + + + G+D G +SG V + D K +++V+ ++GI
Sbjct: 56 VICLCLMIAFGGFVFGWDTGTISGFVN--QTDFKRRFGQMKSDGTYYLSDVRTGLIVGIF 113
Query: 102 SIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPS---YQVLMIGRLLAGIG 158
+I G L GR D+ GR+ + +V+ +G I + +A S YQ IGR+++G+G
Sbjct: 114 NIGCAFGGLTLGRLGDMYGRRIGLMCVVLVYIVG-IVIQIASSDKWYQYF-IGRIISGMG 171
Query: 159 IGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLA 218
+G ++SP I+E +P RG+ +F ++ I +GI LGY +NY S + WRV L
Sbjct: 172 VGGIAVLSPTLISETAPKHIRGTCVSFYQLMITLGIFLGYCTNYGTKDYSNSVQWRVPLG 231
Query: 219 VGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFA 278
+ ++F+ + ++PESPR+LV + R E+A+ L K+N+ E +AE+
Sbjct: 232 LNFAFAIFMIAGMLMVPESPRFLVEKGRYEDAKRSLAKSNKVTIEDPSIVAEMDTIMANV 291
Query: 279 NSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKS 338
+ + W EL S A+ +I G+ IQ QQ++G + YY I A G++D
Sbjct: 292 ETERLAGNASWGELFSNKGAILPRVIMGIMIQSLQQLTGNNYFFYYGTTIFNAVGMKDS- 350
Query: 339 KLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACLF---CMGVTLSLFEKG 395
++ +GI VA+ +DK GR+ L+ + M C +GVT SL+ G
Sbjct: 351 --FQTSIVLGIVNFASTFVALYTVDKFGRRKCLLGGSASMAICFVIFSTVGVT-SLYPNG 407
Query: 396 ---PLVIALG---ILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCS 449
P A G I+F C + FF++ P+ +V+ +E +PLRV+ +A A+ AN +
Sbjct: 408 KDQPSSKAAGNVMIVFTCLFIFFFAISWAPIAYVIVAESYPLRVKNRAMAIAVGANWIWG 467
Query: 450 GLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
L+ ++ AI F ++F + +VF V ETKG +LE++ M+
Sbjct: 468 FLIGFFTPFITSAIGF-SYGYVFMGCLVFSFFYVFFFVCETKGLTLEEVNEMY 519
>GTR4_MOUSE (P14142) Solute carrier family 2, facilitated glucose
transporter, member 4 (Glucose transporter type 4,
insulin-responsive) (GT2)
Length = 509
Score = 204 bits (518), Expect = 6e-52
Identities = 144/480 (30%), Positives = 234/480 (48%), Gaps = 32/480 (6%)
Query: 45 EDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVIFIKEDLKITEVQVE--------- 95
E R ++ A L ++ GY++GV++ I++ T + +
Sbjct: 14 EPPRQRVTGTLVLAVFSAVLGSLQFGYNIGVINAPQKVIEQSYNATWLGRQGPGGPDSIP 73
Query: 96 ---------FLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMTLA---P 143
+ I S+ ++ S G S +GRK M V+ +GG M LA
Sbjct: 74 QGTLTTLWALSVAIFSVGGMISSFLIGIISQWLGRKRAMLANNVLAVLGGALMGLANAVA 133
Query: 144 SYQVLMIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYVSNYA 203
SY++L++GR L G G + P+Y+ EI+P RG+L T ++ I +GI++ V
Sbjct: 134 SYEILILGRFLIGAYSGLTSGLVPMYVGEIAPTHLRGALGTLNQLAIVIGILVAQVLGLE 193
Query: 204 FSGLSVHISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNEDEKE 263
S L W ++LA+ +LP++ L PESPR+L + +E LK +
Sbjct: 194 -SMLGTATLWPLLLALTVLPALLQLILLPFCPESPRYLYIIRNLEGPARKSLKRLTGWAD 252
Query: 264 VEERLAEIQQAAGFANSGKYE-DKPVWRELLSPPPALRRMLITGLGIQCFQQISGIDATV 322
V + LAE++ K E ++P+ L R+ LI + +Q QQ+SGI+A
Sbjct: 253 VSDALAELKD-----EKRKLERERPMSLLQLLGSRTHRQPLIIAVVLQLSQQLSGINAVF 307
Query: 323 YYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMTACL 382
YYS I +AG+ + AT+ G+ TVF LV+++L+++ GR+ L + GM C
Sbjct: 308 YYSTSIFESAGVGQPAY---ATIGAGVVNTVFTLVSVLLVERAGRRTLHLLGLAGMCGCA 364
Query: 383 FCMGVTLSLFEKGPLVIALGILFVCGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGA 442
M V L L E+ P + + I+ + G VAFF +G GP+ W + +E+F R A A+
Sbjct: 365 ILMTVALLLLERVPAMSYVSIVAIFGFVAFFEIGPGPIPWFIVAELFSQGPRPAAMAVAG 424
Query: 443 VANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMF 502
+N C+ +V M F V+DA+ F LF+ + +F F VPET+G++ +QI F
Sbjct: 425 FSNWTCNFIVGMGFQYVADAMG-PYVFLLFAVLLLGFFIFTFLKVPETRGRTFDQISAAF 483
>GTR8_HUMAN (Q9NY64) Solute carrier family 2, facilitated glucose
transporter, member 8 (Glucose transporter type 8)
(Glucose transporter type X1)
Length = 477
Score = 201 bits (511), Expect = 4e-51
Identities = 135/474 (28%), Positives = 238/474 (49%), Gaps = 49/474 (10%)
Query: 55 VIACAIFASLNNVLLGYDVGVMSGAVIFIKEDL----KITEVQVEFLIGILSIVSLLGSL 110
V A A+L + G+ +G S A+ ++ ++ + + ++++ + G +
Sbjct: 26 VFLAAFAAALGPLSFGFALGYSSPAIPSLQRAAPPAPRLDDAAASWFGAVVTLGAAAGGV 85
Query: 111 GGGRTSDIIGRKWTMALAAVVFQMGGITMTLAPSYQVLMIGRLLAGIGIGFGVMISPIYI 170
GG D GRK ++ L +V F G +T A +L+ GRLL G+ G +++P+YI
Sbjct: 86 LGGWLVDRAGRKLSLLLCSVPFVAGFAVITAAQDVWMLLGGRLLTGLACGVASLVAPVYI 145
Query: 171 AEISPNLTRGSLTTFPEIFINVGIMLGYVSNYAFSGLSVHISWRVMLAVGILPSVFIGFA 230
+EI+ RG L + ++ + VGI+L Y++ + + WR + +G +P +
Sbjct: 146 SEIAYPAVRGLLGSCVQLMVVVGILLAYLAGWV-------LEWRWLAVLGCVPPSLMLLL 198
Query: 231 LFIIPESPRWLVMQNRIEEARSVLLKTNEDEKEVEERLAEIQQAAGFANSGKYEDKPVWR 290
+ +PE+PR+L+ Q+R +EA + L E+ E+ +Q+ A
Sbjct: 199 MCFMPETPRFLLTQHRRQEAMAALRFLWGSEQGWEDPPIGAEQSFHLA------------ 246
Query: 291 ELLSPPPALRRMLITGLGIQCFQQISGIDATVYYSPEILMAAGIEDKSKLLAATVAVGIT 350
L P + + I G+ + FQQ+SG++A ++Y+ I A +D S A+V VG+
Sbjct: 247 --LLRQPGIYKPFIIGVSLMAFQQLSGVNAVMFYAETIFEEAKFKDSS---LASVVVGVI 301
Query: 351 KTVFILVAIVLIDKVGRKPLLITSTIGMTACLFCMGVTLSLFEKGP-------------- 396
+ +F VA +++D+ GR+ LL+ S + M G L + GP
Sbjct: 302 QVLFTAVAALIMDRAGRRLLLVLSGVVMVFSTSAFGAYFKLTQGGPGNSSHVAISAPVSA 361
Query: 397 --LVIALGILFV-----CGNVAFFSVGLGPVCWVLTSEIFPLRVRAQASALGAVANRVCS 449
+ ++G+ ++ C +A F+VG GP+ W+L SEIFPL V+ A+ + + N + +
Sbjct: 362 QPVDASVGLAWLAVGSMCLFIAGFAVGWGPIPWLLMSEIFPLHVKGVATGICVLTNWLMA 421
Query: 450 GLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVPETKGKSLEQIEMMFE 503
LV F S+ + + G F+L SA +++F F+ VPETKGK+LEQI FE
Sbjct: 422 FLVTKEFSSLMEVLRPYGAFWLASAFCIFSVLFTFSCVPETKGKTLEQITAHFE 475
>KHT2_KLULA (P53387) Hexose transporter 2
Length = 566
Score = 201 bits (510), Expect = 5e-51
Identities = 138/495 (27%), Positives = 245/495 (48%), Gaps = 28/495 (5%)
Query: 32 VDDNDDVLHQQQLEDKRNSTRKYVIACAIFASLNNVLLGYDVGVMSGAVI---FIKE--- 85
VDD++ + +L K S V + + + G+D G +SG V FI+
Sbjct: 39 VDDDNHSVDAIELPKKPRSAYITVSILCLMVAFGGFVFGWDTGTISGFVNQTDFIRRFGQ 98
Query: 86 -----DLKITEVQVEFLIGILSIVSLLGSLGGGRTSDIIGRKWTMALAAVVFQMGGITMT 140
++ V+ ++ I +I +G + + D+ GR+ + + +++ +G I
Sbjct: 99 EKADGSHYLSNVRTGLIVSIFNIGCAIGGIILSKLGDMYGRRIGLMIVVLIYVVGIIIQI 158
Query: 141 LA-PSYQVLMIGRLLAGIGIGFGVMISPIYIAEISPNLTRGSLTTFPEIFINVGIMLGYV 199
+ + IGR+++G+G+G ++SP+ I+E +P RG+L +F ++ I GI LGY
Sbjct: 159 ASIDKWYQYFIGRIISGLGVGGISVLSPMLISETAPKHIRGTLVSFYQLMITFGIFLGYC 218
Query: 200 SNYAFSGLSVHISWRVMLAVGILPSVFIGFALFIIPESPRWLVMQNRIEEARSVLLKTNE 259
+NY S + WRV L + ++F+ + +PESPR+LV ++RI+EA+ + K+N+
Sbjct: 219 TNYGTKTYSNSVQWRVPLGLCFAWAIFMITGMLFVPESPRFLVEKDRIDEAKRSIAKSNK 278
Query: 260 DEKEVEERLAEIQQAAGFANSGKYEDKPVWRELLSPPPALRRMLITGLGIQCFQQISGID 319
E AE+ + + +EL S + + LI G+ IQ FQQ++G +
Sbjct: 279 VSYEDPAVQAEVDLICAGVEAERLAGSASIKELFSTKTKVFQRLIMGMLIQSFQQLTGNN 338
Query: 320 ATVYYSPEILMAAGIEDKSKLLAATVAVGITKTVFILVAIVLIDKVGRKPLLITSTIGMT 379
YY I + G++D ++ +GI VAI ++DK GR+ L+ MT
Sbjct: 339 YFFYYGTTIFNSVGMDDS---FETSIVLGIVNFASTFVAIYVVDKFGRRKCLLWGAAAMT 395
Query: 380 ACLF---CMGVTLSLFEKG---PLVIALG-----ILFVCGNVAFFSVGLGPVCWVLTSEI 428
AC+ +GVT L+ G P + G I+F C + F+ P+ +V+ +E
Sbjct: 396 ACMVVFASVGVT-RLWPDGANHPETASKGAGNCMIVFACFYIFCFATSWAPIAYVVVAES 454
Query: 429 FPLRVRAQASALGAVANRVCSGLVAMSFLSVSDAISFGGTFFLFSAISALAIVFVFTLVP 488
+PLRV+A+ A+ +N + L ++ AI F + + A+ +VF VP
Sbjct: 455 YPLRVKAKCMAIATASNWIWGFLNGFFTPFITSAIHFYYGYVFMGCLVAM-FFYVFFFVP 513
Query: 489 ETKGKSLEQIEMMFE 503
ETKG +LE+++ M+E
Sbjct: 514 ETKGLTLEEVQEMWE 528
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.140 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 57,781,376
Number of Sequences: 164201
Number of extensions: 2410015
Number of successful extensions: 8183
Number of sequences better than 10.0: 280
Number of HSP's better than 10.0 without gapping: 196
Number of HSP's successfully gapped in prelim test: 84
Number of HSP's that attempted gapping in prelim test: 7361
Number of HSP's gapped (non-prelim): 399
length of query: 530
length of database: 59,974,054
effective HSP length: 115
effective length of query: 415
effective length of database: 41,090,939
effective search space: 17052739685
effective search space used: 17052739685
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 68 (30.8 bits)
Medicago: description of AC126008.6


