

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
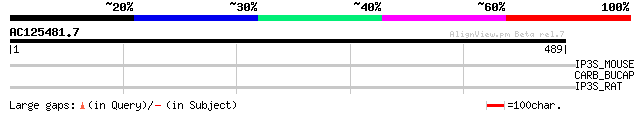
Query= AC125481.7 + phase: 0 /pseudo
(489 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
IP3S_MOUSE (Q9Z329) Inositol 1,4,5-trisphosphate receptor type 2... 33 1.3
CARB_BUCAP (Q8K9Z7) Carbamoyl-phosphate synthase large chain (EC... 32 3.7
IP3S_RAT (P29995) Inositol 1,4,5-trisphosphate receptor type 2 (... 31 8.3
>IP3S_MOUSE (Q9Z329) Inositol 1,4,5-trisphosphate receptor type 2
(Type 2 inositol 1,4,5-trisphosphate receptor) (Type 2
InsP3 receptor) (IP3 receptor isoform 2) (InsP3R2)
(Inositol 1,4,5-trisphosphate type V receptor)
(Fragments)
Length = 1281
Score = 33.5 bits (75), Expect = 1.3
Identities = 31/99 (31%), Positives = 47/99 (47%), Gaps = 18/99 (18%)
Query: 56 TDASNVGQKIVLQPSFAGGRHYMFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFLK 115
++ NVG K+VL P AG + N CK+ + CNT+W+ I F+K
Sbjct: 171 SEGDNVGDKVVLMPVNAGQPLHASNVELLDNPGCKEVN------AVNCNTSWK-ITLFMK 223
Query: 116 -----ERNLKASDRPDIVCRVFKMKLDKLM--DDFEKEE 147
E LK D V R+F + +K + DD+EK++
Sbjct: 224 FSSYREDVLKGGD----VVRLFHAEQEKFLTCDDYEKKQ 258
>CARB_BUCAP (Q8K9Z7) Carbamoyl-phosphate synthase large chain (EC
6.3.5.5) (Carbamoyl-phosphate synthetase ammonia chain)
Length = 1076
Score = 32.0 bits (71), Expect = 3.7
Identities = 16/42 (38%), Positives = 24/42 (57%), Gaps = 4/42 (9%)
Query: 35 NQKTIRCGILNGLHEAMDMGETDASNVGQKIVLQPSFAGGRH 76
N KT +CGI + + EA+ + NVG +++PSF G H
Sbjct: 139 NLKTAKCGIAHNIQEALLV----LKNVGFPCIIRPSFTMGGH 176
>IP3S_RAT (P29995) Inositol 1,4,5-trisphosphate receptor type 2
(Type 2 inositol 1,4,5-trisphosphate receptor) (Type 2
InsP3 receptor) (IP3 receptor isoform 2) (InsP3R2)
Length = 2701
Score = 30.8 bits (68), Expect = 8.3
Identities = 32/100 (32%), Positives = 46/100 (46%), Gaps = 18/100 (18%)
Query: 55 ETDASNVGQKIVLQPSFAGGRHYMFNNCQDAMSICKKYGYPDLFVTIACNTNWREIQDFL 114
E D VG K+VL P AG + N CK+ + CNT+W+ I F+
Sbjct: 172 EGDNIVVGDKVVLMPVNAGQPLHASNVELLDNPGCKEVN------AVNCNTSWK-ITLFM 224
Query: 115 K-----ERNLKASDRPDIVCRVFKMKLDKLM--DDFEKEE 147
K E LK D V R+F + +K + DD+EK++
Sbjct: 225 KFSSYREDVLKGGD----VVRLFHAEQEKFLTCDDYEKKQ 260
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.342 0.150 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,939,391
Number of Sequences: 164201
Number of extensions: 2278093
Number of successful extensions: 6362
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 6358
Number of HSP's gapped (non-prelim): 8
length of query: 489
length of database: 59,974,054
effective HSP length: 114
effective length of query: 375
effective length of database: 41,255,140
effective search space: 15470677500
effective search space used: 15470677500
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 68 (30.8 bits)
Medicago: description of AC125481.7


