

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125480.1 - phase: 0 /pseudo
(250 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
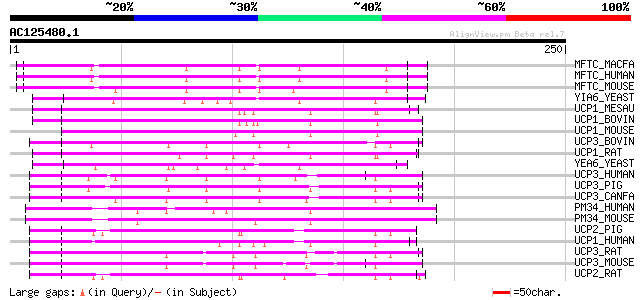
MFTC_MACFA (Q95J75) Mitochondrial folate transporter/carrier (Qm... 108 2e-23
MFTC_HUMAN (Q9H2D1) Mitochondrial folate transporter/carrier 108 2e-23
MFTC_MOUSE (Q8BMG8) Mitochondrial folate transporter/carrier 107 2e-23
YIA6_YEAST (P40556) Putative mitochondrial carrier YIL006W 91 3e-18
UCP1_MESAU (P04575) Mitochondrial brown fat uncoupling protein 1... 82 9e-16
UCP1_BOVIN (P10861) Mitochondrial brown fat uncoupling protein 1... 79 8e-15
UCP1_MOUSE (P12242) Mitochondrial brown fat uncoupling protein 1... 77 5e-14
UCP3_BOVIN (O77792) Mitochondrial uncoupling protein 3 (UCP 3) 76 7e-14
UCP1_RAT (P04633) Mitochondrial brown fat uncoupling protein 1 (... 76 7e-14
YEA6_YEAST (P39953) Putative mitochondrial carrier YEL006W 76 9e-14
UCP3_HUMAN (P55916) Mitochondrial uncoupling protein 3 (UCP 3) 75 1e-13
UCP3_PIG (O97649) Mitochondrial uncoupling protein 3 (UCP 3) 75 1e-13
UCP3_CANFA (Q9N2I9) Mitochondrial uncoupling protein 3 (UCP 3) 73 7e-13
PM34_HUMAN (O43808) Peroxisomal membrane protein PMP34 (34 kDa p... 73 7e-13
PM34_MOUSE (O70579) Peroxisomal membrane protein PMP34 (34 kDa p... 71 2e-12
UCP2_PIG (O97562) Mitochondrial uncoupling protein 2 (UCP 2) 71 3e-12
UCP1_HUMAN (P25874) Mitochondrial brown fat uncoupling protein 1... 70 4e-12
UCP3_RAT (P56499) Mitochondrial uncoupling protein 3 (UCP 3) 70 5e-12
UCP3_MOUSE (P56501) Mitochondrial uncoupling protein 3 (UCP 3) 70 6e-12
UCP2_RAT (P56500) Mitochondrial uncoupling protein 2 (UCP 2) 70 6e-12
>MFTC_MACFA (Q95J75) Mitochondrial folate transporter/carrier
(QmoA-10785/QtrA-13024)
Length = 315
Score = 108 bits (269), Expect = 2e-23
Identities = 65/190 (34%), Positives = 94/190 (49%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD+V+ R V DG P+YN H + TI + +GL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDLVKIRFAVSDG--LELRPKYNGILHCLTTIWKLDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAERLEATEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++D I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYDAVINSPHRQYKGMFDTLVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 70.9 bits (172), Expect = 3e-12
Identities = 56/197 (28%), Positives = 90/197 (45%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + +D + S + +Y + I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYDAVINSPHRQYKGMFDTLVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + L
Sbjct: 182 LYKGFVPGLFGTSHGALQFMAYELLKLKYNQHINRLPEAQLSTVEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q YSG+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQHMF-----YSGVIDVITKTWRKEGIGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + +++L+ K
Sbjct: 297 FVVYENVSHFLLDLREK 313
>MFTC_HUMAN (Q9H2D1) Mitochondrial folate transporter/carrier
Length = 315
Score = 108 bits (269), Expect = 2e-23
Identities = 64/190 (33%), Positives = 94/190 (48%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E +A ++ G +PLD+V+ R V DG P+YN H + TI + +GL
Sbjct: 21 VRYENLIAGVSGGVLSNLALHPLDLVKIRFAVSDG--LELRPKYNGILHCLTTIWKLDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNIWGAGLSWGLYFFFYNAIKSYKTEGRAERLEATEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQTPL---HHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++D I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYDAVVNSPHRQYKGMFDTLVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 70.1 bits (170), Expect = 5e-12
Identities = 56/197 (28%), Positives = 90/197 (45%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQV-HDGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + +D + S + +Y + I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYDAVVNSPHRQYKGMFDTLVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + L
Sbjct: 182 LYKGFVPGLFGTSHGALQFMAYELLKLKYNQHINRLPEAQLSTVEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q YSG+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQHMF-----YSGVIDVITKTWRKEGVGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + +++L+ K
Sbjct: 297 FVVYENVSHFLLDLREK 313
>MFTC_MOUSE (Q8BMG8) Mitochondrial folate transporter/carrier
Length = 316
Score = 107 bits (268), Expect = 2e-23
Identities = 65/190 (34%), Positives = 94/190 (49%), Gaps = 17/190 (8%)
Query: 4 LQWEYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGL 63
+++E VA ++ G +PLD+V+ R V DG P+Y H + TI + +GL
Sbjct: 21 VRYENLVAGVSGGVLSNLALHPLDLVKIRFAVSDG--LEVRPKYKGILHCLATIWKVDGL 78
Query: 64 KGLYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLC 112
+GLY G + GA +SW L F+Y +K + E+ MT+C+
Sbjct: 79 RGLYQGVTPNVWGAGLSWGLYFFFYNAIKSYKTEGRAEQLEPLEYLVSAAEAGAMTLCI- 137
Query: 113 ANPAFVVKTRLQLQ---TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAA 169
NP +V KTRL LQ R Y G++DA I + EG Y+G VPG F S A
Sbjct: 138 TNPLWVTKTRLMLQYGGVASPSQRQYKGMFDALVKIYKYEGVRGLYKGFVPGLFGTSHGA 197
Query: 170 IQFIVYEQLR 179
+QF+ YE L+
Sbjct: 198 LQFMAYELLK 207
Score = 69.7 bits (169), Expect = 6e-12
Identities = 58/197 (29%), Positives = 89/197 (44%), Gaps = 20/197 (10%)
Query: 7 EYPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPR-YNNTTHAIFTIARKEGLKG 65
EY V++ G + PL V +TRL + G + S R Y A+ I + EG++G
Sbjct: 122 EYLVSAAEAGAMTLCITNPLWVTKTRLMLQYGGVASPSQRQYKGMFDALVKIYKYEGVRG 181
Query: 66 LYAGFPAGLLGAT------ISWSLLVFYYGIVKDRHARSKVEKSGALMTVCL-------C 112
LY GF GL G + +++ LL Y +R +++ + + L
Sbjct: 182 LYKGFVPGLFGTSHGALQFMAYELLKLKYNKHINRLPEAQLSTAEYISVAALSKIFAVAA 241
Query: 113 ANPAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQA-AIQ 171
P VV+ RLQ Q H Y G+ D R+EG FY+GI P ++ A I
Sbjct: 242 TYPYQVVRARLQDQ----HVS-YGGVTDVITKTWRKEGIGGFYKGIAPNLIRVTPACCIT 296
Query: 172 FIVYEQLRKTVVNLKTK 188
F+VYE + + +L+ K
Sbjct: 297 FVVYENVSHFLYDLREK 313
>YIA6_YEAST (P40556) Putative mitochondrial carrier YIL006W
Length = 373
Score = 90.9 bits (224), Expect = 3e-18
Identities = 61/166 (36%), Positives = 83/166 (49%), Gaps = 12/166 (7%)
Query: 25 PLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSLL 84
PLDV +TRLQ + + P Y + TI R EG +GLY G +LG +W +
Sbjct: 97 PLDVAKTRLQAQGLQTRFENPYYRGIMGTLSTIVRDEGPRGLYKGLVPIVLGYFPTWMIY 156
Query: 85 V--------FYYGIVK--DRHARSKVEKSGALMTVCLCANPAFVVKTRLQLQTPL-HHAR 133
F++GI D A+S + + L NP +VVKTRL LQ+ L H
Sbjct: 157 FSVYEFSKKFFHGIFPQFDFVAQSCAAITAGAASTTL-TNPIWVVKTRLMLQSNLGEHPT 215
Query: 134 PYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVYEQLR 179
Y G +DAFR + +EGF A Y G+VP + AI F +YE L+
Sbjct: 216 HYKGTFDAFRKLFYQEGFKALYAGLVPSLLGLFHVAIHFPIYEDLK 261
Score = 52.0 bits (123), Expect = 1e-06
Identities = 49/192 (25%), Positives = 81/192 (41%), Gaps = 18/192 (9%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYP-RYNNTTHAIFTIARKEGLKGLYAG 69
A++T G A T P+ VV+TRL + ++P Y T A + +EG K LYAG
Sbjct: 182 AAITAGAASTTLTNPIWVVKTRLMLQSN--LGEHPTHYKGTFDAFRKLFYQEGFKALYAG 239
Query: 70 FPAGLLGA-------TISWSLLVFYYGIVKDRHARS------KVEKSGALMTVCLCANPA 116
LLG I L V ++ ++ + S + S + M P
Sbjct: 240 LVPSLLGLFHVAIHFPIYEDLKVRFHCYSRENNTNSINLQRLIMASSVSKMIASAVTYPH 299
Query: 117 FVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVY 175
+++TR+QL++ + + L+ + +EG FY G I +AI + +
Sbjct: 300 EILRTRMQLKSDIPDS-IQRRLFPLIKATYAQEGLKGFYSGFTTNLVRTIPASAITLVSF 358
Query: 176 EQLRKTVVNLKT 187
E R + N+ T
Sbjct: 359 EYFRNRLENIST 370
>UCP1_MESAU (P04575) Mitochondrial brown fat uncoupling protein 1
(UCP 1) (Thermogenin)
Length = 306
Score = 82.4 bits (202), Expect = 9e-16
Identities = 55/173 (31%), Positives = 81/173 (46%), Gaps = 12/173 (6%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+PLD + RLQ+ S RY I T+A+ EGL LY+G PAG+ SL
Sbjct: 31 FPLDTAKVRLQIQGEGQISSTIRYKGVLGTITTLAKTEGLPKLYSGLPAGIQRQISFASL 90
Query: 84 LVFYYGIVKDRHARSKVEK-------SGALMT---VCLCANPAFVVKTRLQLQTPLHHAR 133
+ Y V++ + K S LMT P VVK RLQ Q+ LH +
Sbjct: 91 RIGLYDTVQEYFSSGKETPPTLGNRISAGLMTGGVAVFIGQPTEVVKVRLQAQSHLHGIK 150
Query: 134 P-YSGLYDAFRTIKREEGFSAFYRGIVPGFFL-ISQAAIQFIVYEQLRKTVVN 184
P Y+G Y+A+R I E FS ++G P + ++ + Y+ ++ +VN
Sbjct: 151 PRYTGTYNAYRIIATTESFSTLWKGTTPNLLRNVIINCVELVTYDLMKGALVN 203
Score = 60.1 bits (144), Expect = 5e-09
Identities = 54/181 (29%), Positives = 76/181 (41%), Gaps = 16/181 (8%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGF 70
A L TG + P +VV+ RLQ L PRY T +A IA E L+ G
Sbjct: 118 AGLMTGGVAVFIGQPTEVVKVRLQAQS-HLHGIKPRYTGTYNAYRIIATTESFSTLWKGT 176
Query: 71 PAGLLGATISWSLLVFYYGIVKDRHARSKVEKSG----------ALMTVCLCANPAFVVK 120
LL I + + Y ++K +++ A A+PA VVK
Sbjct: 177 TPNLLRNVIINCVELVTYDLMKGALVNNQILADDVPCHLLSAFVAGFCTTFLASPADVVK 236
Query: 121 TRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLR 179
TR P Y + T+ +EG +AF++G VP F L S I F+ +EQL+
Sbjct: 237 TRFINSLP----GQYPSVPSCAMTMLTKEGPTAFFKGFVPSFLRLASWNVIMFVCFEQLK 292
Query: 180 K 180
K
Sbjct: 293 K 293
>UCP1_BOVIN (P10861) Mitochondrial brown fat uncoupling protein 1
(UCP 1) (Thermogenin) (Fragment)
Length = 288
Score = 79.3 bits (194), Expect = 8e-15
Identities = 55/173 (31%), Positives = 81/173 (46%), Gaps = 10/173 (5%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+PLD + RLQ+ L S RY I T+A+ EG LY+G PAGL SL
Sbjct: 16 FPLDTAKVRLQIQGECLISSAIRYKGVLGTIITLAKTEGPVKLYSGLPAGLQRQISLASL 75
Query: 84 LVFYYGIVKDRHARSKVEKSGA-----LMT---VCLCANPAFVVKTRLQLQTPLHHARP- 134
+ Y V++ K G+ LMT P VVK RLQ Q+ LH +P
Sbjct: 76 RIGLYDTVQEFFTTGKEASLGSKISAGLMTGGVAVFIGQPTEVVKVRLQAQSHLHGPKPR 135
Query: 135 YSGLYDAFRTIKREEGFSAFYRGIVPGFFL-ISQAAIQFIVYEQLRKTVVNLK 186
Y+G Y+A+R I EG + ++G P + + + Y+ +++ +V K
Sbjct: 136 YTGTYNAYRIIATTEGLTGLWKGTSPNLTTNVIINCTELVTYDLMKEALVKNK 188
Score = 63.2 bits (152), Expect = 6e-10
Identities = 51/186 (27%), Positives = 84/186 (44%), Gaps = 15/186 (8%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGF 70
A L TG + P +VV+ RLQ L PRY T +A IA EGL GL+ G
Sbjct: 101 AGLMTGGVAVFIGQPTEVVKVRLQAQS-HLHGPKPRYTGTYNAYRIIATTEGLTGLWKGT 159
Query: 71 PAGLLGATISWSLLVFYYGIVKDRHARSKVEK--------SGALMTVC--LCANPAFVVK 120
L I + Y ++K+ ++K+ S + C + ++P VVK
Sbjct: 160 SPNLTTNVIINCTELVTYDLMKEALVKNKLLADDVPCHFVSAVVAGFCTTVLSSPVDVVK 219
Query: 121 TRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVYEQLRK 180
TR +P + + + + + EG SAF++G VP F + I F+ +E+L++
Sbjct: 220 TRFVNSSPGQN----TSVPNCAMMMLTREGPSAFFKGFVPSFLRLGSWNIMFVCFERLKQ 275
Query: 181 TVVNLK 186
++ +
Sbjct: 276 ELMKCR 281
>UCP1_MOUSE (P12242) Mitochondrial brown fat uncoupling protein 1
(UCP 1) (Thermogenin)
Length = 306
Score = 76.6 bits (187), Expect = 5e-14
Identities = 53/175 (30%), Positives = 81/175 (46%), Gaps = 12/175 (6%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+PLD + RLQ+ S RY I T+A+ EGL LY+G PAG+ SL
Sbjct: 31 FPLDTAKVRLQIQGEGQASSTIRYKGVLGTITTLAKTEGLPKLYSGLPAGIQRQISFASL 90
Query: 84 LVFYYGIVKDRHARSKV-------EKSGALMT---VCLCANPAFVVKTRLQLQTPLHHAR 133
+ Y V++ + + + S LMT P VVK R+Q Q+ LH +
Sbjct: 91 RIGLYDSVQEYFSSGRETPASLGNKISAGLMTGGVAVFIGQPTEVVKVRMQAQSHLHGIK 150
Query: 134 P-YSGLYDAFRTIKREEGFSAFYRGIVPGFFL-ISQAAIQFIVYEQLRKTVVNLK 186
P Y+G Y+A+R I E S ++G P + + + Y+ ++ +VN K
Sbjct: 151 PRYTGTYNAYRVIATTESLSTLWKGTTPNLMRNVIINCTELVTYDLMKGALVNNK 205
Score = 34.3 bits (77), Expect = 0.29
Identities = 24/77 (31%), Positives = 38/77 (49%), Gaps = 7/77 (9%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+++L GF P+DVV+TR L QYP + +++T KEG + G
Sbjct: 216 LSALVAGFCTTLLASPVDVVKTRF---INSLPGQYPSVPSCAMSMYT---KEGPTAFFKG 269
Query: 70 FPAGLLGATISWSLLVF 86
F A L SW++++F
Sbjct: 270 FVASFLRLG-SWNVIMF 285
>UCP3_BOVIN (O77792) Mitochondrial uncoupling protein 3 (UCP 3)
Length = 311
Score = 76.3 bits (186), Expect = 7e-14
Identities = 57/188 (30%), Positives = 87/188 (45%), Gaps = 15/188 (7%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A TTG +T P DVV+ R Q +Y+ T A TIAR+EG++GL+ G
Sbjct: 121 LAGCTTGAMAVTCAQPTDVVKIRFQASMHTGLGGNRKYSGTMDAYRTIAREEGVRGLWKG 180
Query: 70 F-PAGLLGATISWSLLVFY---------YGIVKDRHARSKVEKSGALMTVCLCANPAFVV 119
P A ++ +V Y Y ++ D V GA L A+P VV
Sbjct: 181 ILPNITRNAIVNCGEMVTYDIIKEKLLDYHLLTDNFPCHFVSAFGAGFCATLVASPVDVV 240
Query: 120 KTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQL 178
KTR P + P+ D + +EG +AFY+G P F L S + F+ YEQ+
Sbjct: 241 KTRYMNSPPGQYHSPF----DCMLKMVTQEGPTAFYKGFTPSFLRLGSWNVVMFVTYEQM 296
Query: 179 RKTVVNLK 186
++ ++ ++
Sbjct: 297 KRALMKVQ 304
Score = 73.9 bits (180), Expect = 3e-13
Identities = 54/180 (30%), Positives = 86/180 (47%), Gaps = 22/180 (12%)
Query: 24 YPLDVVQTRLQV---HDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATIS 80
+PLD + RLQ+ + L ++ +Y I T+ R EG + LY+G AGL
Sbjct: 32 FPLDTAKVRLQIQGENQAALAARSAQYRGVLGTILTMVRTEGPRSLYSGLVAGLQRQMSF 91
Query: 81 WSLLVFYYGIVKDRHARSKVEKSGALMTVCL----------CANPAFVVKTRLQ--LQTP 128
S+ + Y VK + + S + + CA P VVK R Q + T
Sbjct: 92 ASIRIGLYDSVKQFYTPKGSDHSSIITRILAGCTTGAMAVTCAQPTDVVKIRFQASMHTG 151
Query: 129 LHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI----QFIVYEQLRKTVVN 184
L R YSG DA+RTI REEG ++GI+P I++ AI + + Y+ +++ +++
Sbjct: 152 LGGNRKYSGTMDAYRTIAREEGVRGLWKGILPN---ITRNAIVNCGEMVTYDIIKEKLLD 208
>UCP1_RAT (P04633) Mitochondrial brown fat uncoupling protein 1 (UCP
1) (Thermogenin)
Length = 306
Score = 76.3 bits (186), Expect = 7e-14
Identities = 52/173 (30%), Positives = 80/173 (46%), Gaps = 12/173 (6%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+PLD + RLQ+ S RY I T+A+ EGL LY+G PAG+ SL
Sbjct: 31 FPLDTAKVRLQIQGEGQASSTIRYKGVLGTITTLAKTEGLPKLYSGLPAGIQRQISFASL 90
Query: 84 LVFYYGIVKDRHARSK-------VEKSGALMT---VCLCANPAFVVKTRLQLQTPLHHAR 133
+ Y V++ + + + S LMT P VVK R+Q Q+ LH +
Sbjct: 91 RIGLYDTVQEYFSSGRETPASLGSKISAGLMTGGVAVFIGQPTEVVKVRMQAQSHLHGIK 150
Query: 134 P-YSGLYDAFRTIKREEGFSAFYRGIVPGFFL-ISQAAIQFIVYEQLRKTVVN 184
P Y+G Y+A+R I E S ++G P + + + Y+ ++ +VN
Sbjct: 151 PRYTGTYNAYRVIATTESLSTLWKGTTPNLMRNVIINCTELVTYDLMKGALVN 203
Score = 57.0 bits (136), Expect = 4e-08
Identities = 55/184 (29%), Positives = 79/184 (42%), Gaps = 16/184 (8%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGF 70
A L TG + P +VV+ R+Q L PRY T +A IA E L L+ G
Sbjct: 118 AGLMTGGVAVFIGQPTEVVKVRMQAQS-HLHGIKPRYTGTYNAYRVIATTESLSTLWKGT 176
Query: 71 PAGLL-GATISWSLLVFY---------YGIVKDRHARSKVEKSGALMTVCLCANPAFVVK 120
L+ I+ + LV Y + I+ D + A L A+P VVK
Sbjct: 177 TPNLMRNVIINCTELVTYDLMKGALVNHHILADDVPCHLLSALVAGFCTTLLASPVDVVK 236
Query: 121 TRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLR 179
TR P Y + T+ +EG +AF++G P F L S I F+ +EQL+
Sbjct: 237 TRFINSLP----GQYPSVPSCAMTMYTKEGPAAFFKGFAPSFLRLGSWNVIMFVCFEQLK 292
Query: 180 KTVV 183
K ++
Sbjct: 293 KELM 296
Score = 32.3 bits (72), Expect = 1.1
Identities = 23/77 (29%), Positives = 36/77 (45%), Gaps = 7/77 (9%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+++L GF P+DVV+TR L QYP + ++T KEG + G
Sbjct: 216 LSALVAGFCTTLLASPVDVVKTRF---INSLPGQYPSVPSCAMTMYT---KEGPAAFFKG 269
Query: 70 FPAGLLGATISWSLLVF 86
F L SW++++F
Sbjct: 270 FAPSFLRLG-SWNVIMF 285
>YEA6_YEAST (P39953) Putative mitochondrial carrier YEL006W
Length = 335
Score = 75.9 bits (185), Expect = 9e-14
Identities = 56/168 (33%), Positives = 75/168 (44%), Gaps = 15/168 (8%)
Query: 25 PLDVVQTRLQVHD-GRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
P DV +TRLQ + Q Y TI + EG GLY G +LG + +
Sbjct: 58 PFDVAKTRLQAQGLQNMTHQSQHYKGFFGTFATIFKDEGAAGLYKGLQPTVLGYIPTLMI 117
Query: 84 LVFYYGIVKDRHA-----------RSKVEKSGALMTVCLCANPAFVVKTRLQLQTPL-HH 131
Y + S +GA+ TV NP +VVKTRL LQT + +
Sbjct: 118 YFSVYDFCRKYSVDIFPHSPFLSNASSAITAGAISTVA--TNPIWVVKTRLMLQTGIGKY 175
Query: 132 ARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIVYEQLR 179
+ Y G D FR I ++EG A Y G+VP + AIQF +YE L+
Sbjct: 176 STHYKGTIDTFRKIIQQEGAKALYAGLVPALLGMLNVAIQFPLYENLK 223
Score = 47.4 bits (111), Expect = 3e-05
Identities = 47/180 (26%), Positives = 74/180 (41%), Gaps = 18/180 (10%)
Query: 11 ASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGF 70
+++T G P+ VV+TRL + G + Y T I ++EG K LYAG
Sbjct: 144 SAITAGAISTVATNPIWVVKTRLMLQTG-IGKYSTHYKGTIDTFRKIIQQEGAKALYAGL 202
Query: 71 -PA--GLLGATISWSL---LVFYYGIVKDRHARSKVEKSG----------ALMTVCLCAN 114
PA G+L I + L L +G + + V S + M
Sbjct: 203 VPALLGMLNVAIQFPLYENLKIRFGYSESTDVSTDVTSSNFQKLILASMLSKMVASTVTY 262
Query: 115 PAFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFIV 174
P +++TR+QL++ L + L + R+EGF+ FY G AA+ +V
Sbjct: 263 PHEILRTRMQLKSDLPNT-VQRHLLPLIKITYRQEGFAGFYSGFATNLVRTVPAAVVTLV 321
>UCP3_HUMAN (P55916) Mitochondrial uncoupling protein 3 (UCP 3)
Length = 312
Score = 75.5 bits (184), Expect = 1e-13
Identities = 60/190 (31%), Positives = 92/190 (47%), Gaps = 18/190 (9%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQ--VHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLY 67
+A TTG +T P DVV+ R Q +H G S +Y+ T A TIAR+EG++GL+
Sbjct: 121 LAGCTTGAMAVTCAQPTDVVKVRFQASIHLGPSRSDR-KYSGTMDAYRTIAREEGVRGLW 179
Query: 68 AG-FPAGLLGATISWSLLVFY---------YGIVKDRHARSKVEKSGALMTVCLCANPAF 117
G P + A ++ + +V Y Y ++ D V GA + A+P
Sbjct: 180 KGTLPNIMRNAIVNCAEVVTYDILKEKLLDYHLLTDNFPCHFVSAFGAGFCATVVASPVD 239
Query: 118 VVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYE 176
VVKTR P + P D + +EG +AFY+G P F L S + F+ YE
Sbjct: 240 VVKTRYMNSPPGQYFSP----LDCMIKMVAQEGPTAFYKGFTPSFLRLGSWNVVMFVTYE 295
Query: 177 QLRKTVVNLK 186
QL++ ++ ++
Sbjct: 296 QLKRALMKVQ 305
Score = 63.2 bits (152), Expect = 6e-10
Identities = 48/153 (31%), Positives = 61/153 (39%), Gaps = 16/153 (10%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPR---YNNTTHAIFTIARKEGLKGLYAGFPAGLLGATIS 80
+PLD + RLQ+ Q R Y I T+ R EG Y G AGL
Sbjct: 32 FPLDTAKVRLQIQGENQAVQTARLVQYRGVLGTILTMVRTEGPCSPYNGLVAGLQRQMSF 91
Query: 81 WSLLVFYYGIVKDRHARSKVEKSGALMTVCL----------CANPAFVVKTRLQLQT--- 127
S+ + Y VK + + S + CA P VVK R Q
Sbjct: 92 ASIRIGLYDSVKQVYTPKGADNSSLTTRILAGCTTGAMAVTCAQPTDVVKVRFQASIHLG 151
Query: 128 PLHHARPYSGLYDAFRTIKREEGFSAFYRGIVP 160
P R YSG DA+RTI REEG ++G +P
Sbjct: 152 PSRSDRKYSGTMDAYRTIAREEGVRGLWKGTLP 184
>UCP3_PIG (O97649) Mitochondrial uncoupling protein 3 (UCP 3)
Length = 308
Score = 75.1 bits (183), Expect = 1e-13
Identities = 59/190 (31%), Positives = 92/190 (48%), Gaps = 19/190 (10%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQ--VHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLY 67
+A TTG +T P DVV+ R Q +H G ++ +Y+ T A TIAR+EG++GL+
Sbjct: 118 LAGCTTGAMAVTCAQPTDVVKVRFQASIHAGPRSNR--KYSGTMDAYRTIAREEGVRGLW 175
Query: 68 AGF-PAGLLGATISWSLLVFY---------YGIVKDRHARSKVEKSGALMTVCLCANPAF 117
G P A ++ + +V Y Y ++ D V GA + A+P
Sbjct: 176 KGILPNITRNAIVNCAEMVTYDVIKEKVLDYHLLTDNLPCHFVSAFGAGFCATVVASPVD 235
Query: 118 VVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYE 176
VVKTR P + P D + +EG +AFY+G P F L S + F+ YE
Sbjct: 236 VVKTRYMNSPPGQYQNPL----DCMLKMVTQEGPTAFYKGFTPSFLRLGSWNVVMFVSYE 291
Query: 177 QLRKTVVNLK 186
QL++ ++ ++
Sbjct: 292 QLKRALMKVQ 301
Score = 70.1 bits (170), Expect = 5e-12
Identities = 53/180 (29%), Positives = 80/180 (44%), Gaps = 25/180 (13%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+PLD + RLQ+ ++ +Y I T+ R EG + Y G AGL S+
Sbjct: 32 FPLDTAKVRLQIQGENQAARSAQYRGVLGTILTMVRNEGPRSPYNGLVAGLQRQMSFASI 91
Query: 84 LVFYYGIVKDRHARSKVEKSGALMTVCL----------CANPAFVVKTRLQLQTPLHHAR 133
+ Y VK + + S + CA P VVK R Q HA
Sbjct: 92 RIGLYDSVKQLYTPKGSDHSSITTRILAGCTTGAMAVTCAQPTDVVKVRFQASI---HAG 148
Query: 134 P-----YSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI----QFIVYEQLRKTVVN 184
P YSG DA+RTI REEG ++GI+P I++ AI + + Y+ +++ V++
Sbjct: 149 PRSNRKYSGTMDAYRTIAREEGVRGLWKGILPN---ITRNAIVNCAEMVTYDVIKEKVLD 205
>UCP3_CANFA (Q9N2I9) Mitochondrial uncoupling protein 3 (UCP 3)
Length = 311
Score = 72.8 bits (177), Expect = 7e-13
Identities = 54/188 (28%), Positives = 87/188 (45%), Gaps = 15/188 (7%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A TTG ++ P DVV+ R Q +Y+ T A TIAR+EG++GL+ G
Sbjct: 121 LAGCTTGAMAVSCAQPTDVVKVRFQASIHLGAGSNRKYSGTMDAYRTIAREEGVRGLWKG 180
Query: 70 -FPAGLLGATISWSLLVFY---------YGIVKDRHARSKVEKSGALMTVCLCANPAFVV 119
P A ++ + +V Y Y ++ D + GA + A+P VV
Sbjct: 181 TLPNITRNAIVNCAEMVTYDIIKEKLLDYHLLTDNFPCHLISAFGAGFCATVVASPVDVV 240
Query: 120 KTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQL 178
KTR P + P D + +EG +AFY+G P F L + + F+ YEQL
Sbjct: 241 KTRYMNSPPGQYCSPL----DCMLKMVTQEGPTAFYKGFTPSFLRLGTWNVVMFVTYEQL 296
Query: 179 RKTVVNLK 186
++ ++ ++
Sbjct: 297 KRALMKVQ 304
Score = 64.3 bits (155), Expect = 3e-10
Identities = 52/180 (28%), Positives = 78/180 (42%), Gaps = 22/180 (12%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPR---YNNTTHAIFTIARKEGLKGLYAGFPAGLLGATIS 80
+PLD + RLQ+ +Q R Y I T+ R EG + Y G AGL
Sbjct: 32 FPLDTAKVRLQIQGENQATQAARRIQYRGVLGTILTMVRTEGPRSPYNGLVAGLQRQMSF 91
Query: 81 WSLLVFYYGIVKDRHARSKVEKSGALMTVCL----------CANPAFVVKTRLQLQTPLH 130
S+ + Y VK + + S + CA P VVK R Q L
Sbjct: 92 ASIRIGLYDSVKQFYTPKGSDHSSITTRILAGCTTGAMAVSCAQPTDVVKVRFQASIHLG 151
Query: 131 HA--RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI----QFIVYEQLRKTVVN 184
R YSG DA+RTI REEG ++G +P I++ AI + + Y+ +++ +++
Sbjct: 152 AGSNRKYSGTMDAYRTIAREEGVRGLWKGTLPN---ITRNAIVNCAEMVTYDIIKEKLLD 208
>PM34_HUMAN (O43808) Peroxisomal membrane protein PMP34 (34 kDa
peroxisomal membrane protein) (Solute carrier family 25,
member 17)
Length = 307
Score = 72.8 bits (177), Expect = 7e-13
Identities = 62/202 (30%), Positives = 94/202 (45%), Gaps = 27/202 (13%)
Query: 8 YPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFT-IARKEGLKGL 66
+ VA +T +PLD + RLQV + R + TTH + I ++EGL
Sbjct: 12 HAVAGAVGSVTAMTVFFPLDTARLRLQVDE-------KRKSKTTHMVLLEIIKEEGLLAP 64
Query: 67 YAG-FPAGLLGATISWSLLVFYYGI-------VKDRHA---RSKVEKSGALMTVCLCANP 115
Y G FP + +++ S V++Y VK +H+ + V A + L P
Sbjct: 65 YRGWFP---VISSLCCSNFVYFYTFNSLKALWVKGQHSTTGKDLVVGFVAGVVNVLLTTP 121
Query: 116 AFVVKTRLQLQTPLHHARP-----YSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI 170
+VV TRL+LQ Y G+ DAF I R+EG SA + G P L+ AI
Sbjct: 122 LWVVNTRLKLQGAKFRNEDIVPTNYKGIIDAFHQIIRDEGISALWNGTFPSLLLVFNPAI 181
Query: 171 QFIVYEQLRKTVVNLKTKGSKI 192
QF+ YE L++ ++ + K S +
Sbjct: 182 QFMFYEGLKRQLLKKRMKLSSL 203
>PM34_MOUSE (O70579) Peroxisomal membrane protein PMP34 (34 kDa
peroxisomal membrane protein) (Solute carrier family 25,
member 17)
Length = 307
Score = 71.2 bits (173), Expect = 2e-12
Identities = 56/199 (28%), Positives = 88/199 (44%), Gaps = 21/199 (10%)
Query: 8 YPVASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFT-IARKEGLKGL 66
+ VA +T +PLD + RLQV + R + TTHA+ I ++EGL
Sbjct: 12 HAVAGAVGSVTAMTVFFPLDTARLRLQVDE-------KRKSKTTHAVLLEIIKEEGLLAP 64
Query: 67 YAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKSGALMTV--------CLCANPAFV 118
Y G+ + S + + + +K + + +G + V L P +V
Sbjct: 65 YRGWFPVISSLCCSNFVYFYTFNSLKAVWVKGQRSSTGKDLVVGFVAGVVNVLLTTPLWV 124
Query: 119 VKTRLQLQTPLHHARP-----YSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAIQFI 173
V TRL+LQ Y G+ DAF I R+EG A + G P L+ AIQF+
Sbjct: 125 VNTRLKLQGAKFRNEDIIPTNYKGIIDAFHQIIRDEGILALWNGTFPSLLLVFNPAIQFM 184
Query: 174 VYEQLRKTVVNLKTKGSKI 192
YE L++ ++ + K S +
Sbjct: 185 FYEGLKRQLLKKRMKLSSL 203
>UCP2_PIG (O97562) Mitochondrial uncoupling protein 2 (UCP 2)
Length = 309
Score = 70.9 bits (172), Expect = 3e-12
Identities = 58/189 (30%), Positives = 84/189 (43%), Gaps = 25/189 (13%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVH----DGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
+A TTG + P DVV+ R Q GR RY +T A TIAR+EGL+G
Sbjct: 121 LAGSTTGALAVAVAQPTDVVKVRFQAQARAGGGR------RYRSTVDAYKTIAREEGLRG 174
Query: 66 LYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKS----------GALMTVCLCANP 115
L+ G + I + Y ++KD ++ + GA + A+P
Sbjct: 175 LWKGTSPNVARNAIVNCAELVTYDLIKDTLLKADLMTDDLPCHFTSAFGAGFCTTVIASP 234
Query: 116 AFVVKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIV 174
VVKTR P YS T+ ++EG AFY+G P F L S + F+
Sbjct: 235 VDVVKTRYMNSAP----GQYSSAGHCALTMLQKEGPRAFYKGFTPSFLRLGSWNVVMFVT 290
Query: 175 YEQLRKTVV 183
YEQL++ ++
Sbjct: 291 YEQLKRALM 299
Score = 61.6 bits (148), Expect = 2e-09
Identities = 50/179 (27%), Positives = 76/179 (41%), Gaps = 24/179 (13%)
Query: 24 YPLDVVQTRLQVHDGRL----FSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATI 79
+PLD + RLQ+ R + +Y I T+ R EG + LY G AGL
Sbjct: 32 FPLDTAKVRLQIQGERRGPVQAAASAQYRGVLGTILTMVRNEGPRSLYNGLVAGLQRQMS 91
Query: 80 SWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLCANPAFVVKTRLQLQTP 128
S+ + Y VK + + +GAL A P VVK R Q Q
Sbjct: 92 FASVRIGLYDSVKHFYTKGSEHAGIGSRLLAGSTTGALAVAV--AQPTDVVKVRFQAQAR 149
Query: 129 LHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI----QFIVYEQLRKTVV 183
R Y DA++TI REEG ++G P +++ AI + + Y+ ++ T++
Sbjct: 150 AGGGRRYRSTVDAYKTIAREEGLRGLWKGTSPN---VARNAIVNCAELVTYDLIKDTLL 205
>UCP1_HUMAN (P25874) Mitochondrial brown fat uncoupling protein 1
(UCP 1) (Thermogenin)
Length = 307
Score = 70.5 bits (171), Expect = 4e-12
Identities = 53/172 (30%), Positives = 77/172 (43%), Gaps = 12/172 (6%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+PLD + RLQV S RY I + + EG LY+G PAGL S SL
Sbjct: 32 FPLDTAKVRLQVQGECPTSSVIRYKGVLGTITAVVKTEGRMKLYSGLPAGLQRQISSASL 91
Query: 84 LVFYYGIVKD------RHARSKVEKSGALMT----VCLCANPAFVVKTRLQLQTPLHHAR 133
+ Y V++ A S K A +T P VVK RLQ Q+ LH +
Sbjct: 92 RIGLYDTVQEFLTAGKETAPSLGSKILAGLTTGGVAVFIGQPTEVVKVRLQAQSHLHGIK 151
Query: 134 P-YSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQLRKTVV 183
P Y+G Y+A+R I EG + ++G P + + + Y+ +++ V
Sbjct: 152 PRYTGTYNAYRIIATTEGLTGLWKGTTPNLMRSVIINCTELVTYDLMKEAFV 203
Score = 67.0 bits (162), Expect = 4e-11
Identities = 55/182 (30%), Positives = 83/182 (45%), Gaps = 16/182 (8%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A LTTG + P +VV+ RLQ L PRY T +A IA EGL GL+ G
Sbjct: 118 LAGLTTGGVAVFIGQPTEVVKVRLQAQS-HLHGIKPRYTGTYNAYRIIATTEGLTGLWKG 176
Query: 70 FPAGLLGATISWSLLVFYYGIVKDRHARSKVEK--------SGALMTVCLCA--NPAFVV 119
L+ + I + Y ++K+ ++ + S + C A +P VV
Sbjct: 177 TTPNLMRSVIINCTELVTYDLMKEAFVKNNILADDVPCHLVSALIAGFCATAMSSPVDVV 236
Query: 120 KTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQL 178
KTR P Y + + + EG +AF++G+VP F L S I F+ +EQL
Sbjct: 237 KTRFINSPP----GQYKSVPNCAMKVFTNEGPTAFFKGLVPSFLRLGSWNVIMFVCFEQL 292
Query: 179 RK 180
++
Sbjct: 293 KR 294
>UCP3_RAT (P56499) Mitochondrial uncoupling protein 3 (UCP 3)
Length = 308
Score = 70.1 bits (170), Expect = 5e-12
Identities = 59/189 (31%), Positives = 92/189 (48%), Gaps = 17/189 (8%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A TTG +T P DVV+ R Q +Y T A TIAR+EG++GL+ G
Sbjct: 118 LAGCTTGAMAVTCAQPTDVVKVRFQAMIRLGTGGERKYRGTMDAYRTIAREEGVRGLWKG 177
Query: 70 -FPAGLLGATISWSLLVFYYGIVKDRHARSK----------VEKSGALMTVCLCANPAFV 118
+P A ++ + +V Y I+K++ S V GA + A+P V
Sbjct: 178 TWPNITRNAIVNCAEMVTY-DIIKEKLLDSHLFTDNFPCHFVSAFGAGFCATVVASPVDV 236
Query: 119 VKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQ 177
VKTR P + P L+ R + +EG +AFY+G +P F L S + F+ YEQ
Sbjct: 237 VKTRYMNAPPGRYRSP---LHCMLRMVA-QEGPTAFYKGFMPSFLRLGSWNVMMFVTYEQ 292
Query: 178 LRKTVVNLK 186
L++ ++ ++
Sbjct: 293 LKRALMKVQ 301
Score = 65.1 bits (157), Expect = 2e-10
Identities = 50/177 (28%), Positives = 77/177 (43%), Gaps = 19/177 (10%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+PLD + RLQ+ Q +Y I T+ R EG + Y+G AGL S+
Sbjct: 32 FPLDTAKVRLQIQGENPGVQSVQYRGVLGTILTMVRTEGPRSPYSGLVAGLHRQMSFASI 91
Query: 84 LVFYYGIVKDRHARSKVEKSGALMTV----------CLCANPAFVVKTRLQLQTPLHHA- 132
+ Y VK + + S + + CA P VVK R Q L
Sbjct: 92 RIGLYDSVKQFYTPKGTDHSSVAIRILAGCTTGAMAVTCAQPTDVVKVRFQAMIRLGTGG 151
Query: 133 -RPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI----QFIVYEQLRKTVVN 184
R Y G DA+RTI REEG ++G P I++ AI + + Y+ +++ +++
Sbjct: 152 ERKYRGTMDAYRTIAREEGVRGLWKGTWPN---ITRNAIVNCAEMVTYDIIKEKLLD 205
>UCP3_MOUSE (P56501) Mitochondrial uncoupling protein 3 (UCP 3)
Length = 308
Score = 69.7 bits (169), Expect = 6e-12
Identities = 60/189 (31%), Positives = 94/189 (48%), Gaps = 17/189 (8%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAG 69
+A TTG +T P DVV+ R Q +Y T A TIAR+EG++GL+ G
Sbjct: 118 LAGCTTGAMAVTCAQPTDVVKVRFQAMIRLGTGGERKYRGTMDAYRTIAREEGVRGLWKG 177
Query: 70 -FPAGLLGATISWSLLVFYYGIVKDRHARSK----------VEKSGALMTVCLCANPAFV 118
+P A ++ + +V Y I+K++ S V GA + A+P V
Sbjct: 178 TWPNITRNAIVNCAEMVTY-DIIKEKLLESHLFTDNFPCHFVSAFGAGFCATVVASPVDV 236
Query: 119 VKTRLQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFIVYEQ 177
VKTR + PL R S L+ + + +EG +AFY+G VP F L + + F+ YEQ
Sbjct: 237 VKTRY-MNAPLGRYR--SPLHCMLKMVA-QEGPTAFYKGFVPSFLRLGAWNVMMFVTYEQ 292
Query: 178 LRKTVVNLK 186
L++ ++ ++
Sbjct: 293 LKRALMKVQ 301
Score = 65.9 bits (159), Expect = 9e-11
Identities = 46/149 (30%), Positives = 63/149 (41%), Gaps = 12/149 (8%)
Query: 24 YPLDVVQTRLQVHDGRLFSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATISWSL 83
+PLD + RLQ+ +Q +Y I T+ R EG + Y+G AGL S+
Sbjct: 32 FPLDTAKVRLQIQGENPGAQSVQYRGVLGTILTMVRTEGPRSPYSGLVAGLHRQMSFASI 91
Query: 84 LVFYYGIVKDRHARSKVEKSGALMTV----------CLCANPAFVVKTRLQLQTPLHHA- 132
+ Y VK + + S + + CA P VVK R Q L
Sbjct: 92 RIGLYDSVKQFYTPKGADHSSVAIRILAGCTTGAMAVTCAQPTDVVKVRFQAMIRLGTGG 151
Query: 133 -RPYSGLYDAFRTIKREEGFSAFYRGIVP 160
R Y G DA+RTI REEG ++G P
Sbjct: 152 ERKYRGTMDAYRTIAREEGVRGLWKGTWP 180
Score = 29.3 bits (64), Expect = 9.3
Identities = 21/69 (30%), Positives = 35/69 (50%), Gaps = 3/69 (4%)
Query: 115 PAFVVKTRLQLQ--TPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQ 171
P K RLQ+Q P + Y G+ T+ R EG + Y G+V G +S A+I+
Sbjct: 33 PLDTAKVRLQIQGENPGAQSVQYRGVLGTILTMVRTEGPRSPYSGLVAGLHRQMSFASIR 92
Query: 172 FIVYEQLRK 180
+Y+ +++
Sbjct: 93 IGLYDSVKQ 101
>UCP2_RAT (P56500) Mitochondrial uncoupling protein 2 (UCP 2)
Length = 309
Score = 69.7 bits (169), Expect = 6e-12
Identities = 57/190 (30%), Positives = 88/190 (46%), Gaps = 27/190 (14%)
Query: 10 VASLTTGFAFITFQYPLDVVQTRLQVH----DGRLFSQYPRYNNTTHAIFTIARKEGLKG 65
+A TTG + P DVV+ R Q GR RY +T A TIAR+EG++G
Sbjct: 121 LAGSTTGALAVAVAQPTDVVKVRFQAQARAGGGR------RYQSTVEAYKTIAREEGIRG 174
Query: 66 LYAGFPAGLLGATISWSLLVFYYGIVKDRHARSKVEKS----------GALMTVCLCANP 115
L+ G + I + Y ++KD ++ + GA + A+P
Sbjct: 175 LWKGTSPNVARNAIVNCTELVTYDLIKDTLLKANLMTDDLPCHFTSAFGAGFCTTVIASP 234
Query: 116 AFVVKTR-LQLQTPLHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFF-LISQAAIQFI 173
VVKTR + +H+ + L T+ R+EG AFY+G +P F L S + F+
Sbjct: 235 VDVVKTRYMNSALGQYHSAGHCAL-----TMLRKEGPRAFYKGFMPSFLRLGSWNVVMFV 289
Query: 174 VYEQLRKTVV 183
YEQL++ ++
Sbjct: 290 TYEQLKRALM 299
Score = 58.5 bits (140), Expect = 1e-08
Identities = 51/185 (27%), Positives = 78/185 (41%), Gaps = 26/185 (14%)
Query: 24 YPLDVVQTRLQVHDGRL----FSQYPRYNNTTHAIFTIARKEGLKGLYAGFPAGLLGATI 79
+PLD + RLQ+ + +Y I T+ R EG + LY G AGL
Sbjct: 32 FPLDTAKVRLQIQGESQGLARTAASAQYRGVLGTILTMVRTEGPRSLYNGLVAGLQRQMS 91
Query: 80 SWSLLVFYYGIVKDRHARSKVEK-----------SGALMTVCLCANPAFVVKTRLQLQTP 128
S+ + Y VK + + +GAL A P VVK R Q Q
Sbjct: 92 FASVRIGLYDSVKQFYTKGSEHAGIGSRLLAGSTTGALAVAV--AQPTDVVKVRFQAQAR 149
Query: 129 LHHARPYSGLYDAFRTIKREEGFSAFYRGIVPGFFLISQAAI----QFIVYEQLRKTVV- 183
R Y +A++TI REEG ++G P +++ AI + + Y+ ++ T++
Sbjct: 150 AGGGRRYQSTVEAYKTIAREEGIRGLWKGTSPN---VARNAIVNCTELVTYDLIKDTLLK 206
Query: 184 -NLKT 187
NL T
Sbjct: 207 ANLMT 211
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.143 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,093,257
Number of Sequences: 164201
Number of extensions: 1169342
Number of successful extensions: 3626
Number of sequences better than 10.0: 157
Number of HSP's better than 10.0 without gapping: 77
Number of HSP's successfully gapped in prelim test: 80
Number of HSP's that attempted gapping in prelim test: 2949
Number of HSP's gapped (non-prelim): 395
length of query: 250
length of database: 59,974,054
effective HSP length: 108
effective length of query: 142
effective length of database: 42,240,346
effective search space: 5998129132
effective search space used: 5998129132
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC125480.1


