

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124972.17 + phase: 0 /pseudo/partial
(289 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
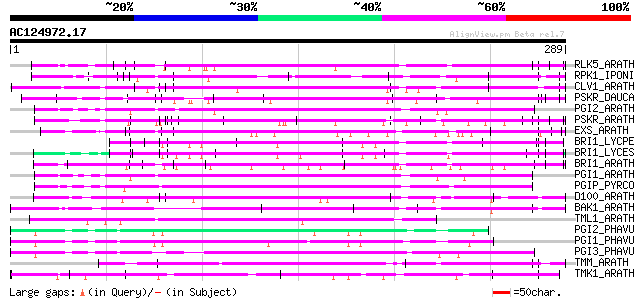
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 109 9e-24
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 102 1e-21
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 96 8e-20
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 96 1e-19
PGI2_ARATH (Q9M5J8) Polygalacturonase inhibitor 2 precursor (Pol... 96 1e-19
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 92 1e-18
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 92 1e-18
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 92 1e-18
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 91 4e-18
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 91 4e-18
PGI1_ARATH (Q9M5J9) Polygalacturonase inhibitor 1 precursor (Pol... 90 6e-18
PGIP_PYRCO (Q05091) Polygalacturonase inhibitor precursor (Polyg... 90 7e-18
D100_ARATH (Q00874) DNA-damage-repair/toleration protein DRT100 ... 87 4e-17
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 80 6e-15
TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor 79 2e-14
PGI2_PHAVU (P58822) Polygalacturonase inhibitor 2 precursor (Pol... 74 3e-13
PGI1_PHAVU (P35334) Polygalacturonase inhibitor 1 precursor (Pol... 74 5e-13
PGI3_PHAVU (P58823) Polygalacturonase inhibitor 3 precursor (Pol... 73 7e-13
TMM_ARATH (Q9SSD1) TOO MANY MOUTHS protein precursor (TMM) 72 1e-12
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 68 2e-11
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 109 bits (272), Expect = 9e-24
Identities = 98/330 (29%), Positives = 157/330 (46%), Gaps = 66/330 (20%)
Query: 12 NRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILC-SEGNVISITLDNAS 70
N+D L + K G+ +DP L+SW + + P W G+ C + NV+S+ L +
Sbjct: 22 NQDATILRQAKLGL-SDP-AQSLSSWSDNN---DVTPCKWLGVSCDATSNVVSVDLSSFM 76
Query: 71 LVGEFNFLAISNLPMLHNLSVVNNHFTGSM-------------LHISP------------ 105
LVG F + + +LP LH+LS+ NN GS+ L +S
Sbjct: 77 LVGPFPSI-LCHLPSLHSLSLYNNSINGSLSADDFDTCHNLISLDLSENLLVGSIPKSLP 135
Query: 106 --MKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFH 163
+ +LKFL++S N + ++P SF E R L LNL+ N SGT+P + L+ L
Sbjct: 136 FNLPNLKFLEISGNNLSDTIPSSFGEFRKLESLNLAGNFLSGTIPASLGNVTTLKELKLA 195
Query: 164 SNSFS-------------------------GDIMEIFYQMGSVLHVDLSNNKFSGALDLG 198
N FS G I ++ S++++DL+ N+ +G++
Sbjct: 196 YNLFSPSQIPSQLGNLTELQVLWLAGCNLVGPIPPSLSRLTSLVNLDLTFNQLTGSIP-- 253
Query: 199 LGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLR 258
++ L +++ + + +NS GEL + M + L+ FDAS N+L G IP +++L
Sbjct: 254 -SWITQLKTVEQIELFNNSFSGEL--PESMGNMTTLKRFDASMNKLTGKIPDNLNLLNLE 310
Query: 259 ILRLACNQLTGSLPETLLKESSMMLSELDL 288
L L N L G LPE++ + S LSEL L
Sbjct: 311 SLNLFENMLEGPLPESITR--SKTLSELKL 338
Score = 97.4 bits (241), Expect = 4e-20
Identities = 70/222 (31%), Positives = 113/222 (50%), Gaps = 13/222 (5%)
Query: 55 LCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVV---NNHFTGSMLH-ISPMKSLK 110
+C EG + + L + S GE + +NL +L+ V NN +G + H + L
Sbjct: 375 VCGEGKLEYLILIDNSFSGEIS----NNLGKCKSLTRVRLSNNKLSGQIPHGFWGLPRLS 430
Query: 111 FLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGD 170
L+LS N F GS+P + + ++L L +S N FSG++PN L+ + + N FSG+
Sbjct: 431 LLELSDNSFTGSIPKTIIGAKNLSNLRISKNRFSGSIPNEIGSLNGIIEISGAENDFSGE 490
Query: 171 IMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPY 230
I E ++ + +DLS N+ SG + L ++ LN+++N L GE+ G+
Sbjct: 491 IPESLVKLKQLSRLDLSKNQLSGEIPRELRGWK---NLNELNLANNHLSGEIPKEVGI-- 545
Query: 231 LDNLEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLP 272
L L D S+NQ G IP + L +L L+ N L+G +P
Sbjct: 546 LPVLNYLDLSSNQFSGEIPLELQNLKLNVLNLSYNHLSGKIP 587
Score = 81.3 bits (199), Expect = 3e-15
Identities = 64/225 (28%), Positives = 112/225 (49%), Gaps = 17/225 (7%)
Query: 66 LDNASLVGEFNFLA------ISNLPMLHNLSVVNNHFTGSML--HISPMKSLKFLDLSLN 117
L++ +L G NFL+ + N+ L L + N F+ S + + + L+ L L+
Sbjct: 165 LESLNLAG--NFLSGTIPASLGNVTTLKELKLAYNLFSPSQIPSQLGNLTELQVLWLAGC 222
Query: 118 KFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQ 177
G +PPS L SLV L+L+ N+ +G++P+ +L +E ++ +NSFSG++ E
Sbjct: 223 NLVGPIPPSLSRLTSLVNLDLTFNQLTGSIPSWITQLKTVEQIELFNNSFSGELPESMGN 282
Query: 178 MGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVF 237
M ++ D S NK +G + L L +++ LN+ N L G L + + L
Sbjct: 283 MTTLKRFDASMNKLTGKIPDNLN----LLNLESLNLFENMLEGPL--PESITRSKTLSEL 336
Query: 238 DASNNQLVGNIPSFTFVVS-LRILRLACNQLTGSLPETLLKESSM 281
NN+L G +PS S L+ + L+ N+ +G +P + E +
Sbjct: 337 KLFNNRLTGVLPSQLGANSPLQYVDLSYNRFSGEIPANVCGEGKL 381
Score = 78.6 bits (192), Expect = 2e-14
Identities = 72/211 (34%), Positives = 101/211 (47%), Gaps = 11/211 (5%)
Query: 82 NLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSL 140
NL L +L++ N G + I+ K+L L L N+ G LP L Y++LS
Sbjct: 305 NLLNLESLNLFENMLEGPLPESITRSKTLSELKLFNNRLTGVLPSQLGANSPLQYVDLSY 364
Query: 141 NEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLG 200
N FSG +P +LEYL NSFSG+I + S+ V LSNNK SG + G
Sbjct: 365 NRFSGEIPANVCGEGKLEYLILIDNSFSGEISNNLGKCKSLTRVRLSNNKLSGQIPHGFW 424
Query: 201 DVSFLFSIQHLNVSHNSLVGEL-FAHDGMPYLDNLEVFDASNNQLVGNIPS-FTFVVSLR 258
L + L +S NS G + G L NL + S N+ G+IP+ + +
Sbjct: 425 G---LPRLSLLELSDNSFTGSIPKTIIGAKNLSNLRI---SKNRFSGSIPNEIGSLNGII 478
Query: 259 ILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+ A N +G +PE+L+K LS LDLS
Sbjct: 479 EISGAENDFSGEIPESLVK--LKQLSRLDLS 507
Score = 78.2 bits (191), Expect = 2e-14
Identities = 62/216 (28%), Positives = 101/216 (46%), Gaps = 7/216 (3%)
Query: 61 VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFN 120
V I L N S GE ++ N+ L N TG + + +L+ L+L N
Sbjct: 262 VEQIELFNNSFSGELPE-SMGNMTTLKRFDASMNKLTGKIPDNLNLLNLESLNLFENMLE 320
Query: 121 GSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGS 180
G LP S ++L L L N +G +P+ L+Y+D N FSG+I G
Sbjct: 321 GPLPESITRSKTLSELKLFNNRLTGVLPSQLGANSPLQYVDLSYNRFSGEIPANVCGEGK 380
Query: 181 VLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDAS 240
+ ++ L +N FSG + LG S+ + +S+N L G++ G L L + + S
Sbjct: 381 LEYLILIDNSFSGEISNNLGKCK---SLTRVRLSNNKLSGQI--PHGFWGLPRLSLLELS 435
Query: 241 NNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
+N G+IP + +L LR++ N+ +GS+P +
Sbjct: 436 DNSFTGSIPKTIIGAKNLSNLRISKNRFSGSIPNEI 471
Score = 34.3 bits (77), Expect = 0.37
Identities = 23/69 (33%), Positives = 38/69 (54%), Gaps = 3/69 (4%)
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNK 118
N+ + L N L GE + LP+L+ L + +N F+G + L + +K L L+LS N
Sbjct: 524 NLNELNLANNHLSGEIP-KEVGILPVLNYLDLSSNQFSGEIPLELQNLK-LNVLNLSYNH 581
Query: 119 FNGSLPPSF 127
+G +PP +
Sbjct: 582 LSGKIPPLY 590
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 102 bits (253), Expect = 1e-21
Identities = 82/267 (30%), Positives = 129/267 (47%), Gaps = 14/267 (5%)
Query: 12 NRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVI-SITLDNAS 70
N D ALL L + + P + SW++ S+ P +W G+ C + ++ L +
Sbjct: 25 NSDGAALLSLTRHWTSIPSDIT-QSWNA----SDSTPCSWLGVECDRRQFVDTLNLSSYG 79
Query: 71 LVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVE 129
+ GEF IS+L L + + N F GS+ + L+ +DLS N F G++P +
Sbjct: 80 ISGEFG-PEISHLKHLKKVVLSGNGFFGSIPSQLGNCSLLEHIDLSSNSFTGNIPDTLGA 138
Query: 130 LRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNN 189
L++L L+L N G P + LE + F N +G I M + + L +N
Sbjct: 139 LQNLRNLSLFFNSLIGPFPESLLSIPHLETVYFTGNGLNGSIPSNIGNMSELTTLWLDDN 198
Query: 190 KFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP 249
+FSG + LG+++ ++Q L ++ N+LVG L + L+NL D NN LVG IP
Sbjct: 199 QFSGPVPSSLGNIT---TLQELYLNDNNLVGTLPV--TLNNLENLVYLDVRNNSLVGAIP 253
Query: 250 -SFTFVVSLRILRLACNQLTGSLPETL 275
F + + L+ NQ TG LP L
Sbjct: 254 LDFVSCKQIDTISLSNNQFTGGLPPGL 280
Score = 84.0 bits (206), Expect = 4e-16
Identities = 62/204 (30%), Positives = 97/204 (47%), Gaps = 7/204 (3%)
Query: 86 LHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSG 145
L L + N+ G + ++L F DLS N F G +PPS L+++ + LS N+ SG
Sbjct: 478 LERLILEENNLRGGLPDFVEKQNLLFFDLSGNNFTGPIPPSLGNLKNVTAIYLSSNQLSG 537
Query: 146 TVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFL 205
++P L +LE+L+ N G + + +D S+N +G++ LG L
Sbjct: 538 SIPPELGSLVKLEHLNLSHNILKGILPSELSNCHKLSELDASHNLLNGSIPSTLGS---L 594
Query: 206 FSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRILRLACN 265
+ L++ NS G + + + L N L G+IP + +LR L L+ N
Sbjct: 595 TELTKLSLGENSFSGGI--PTSLFQSNKLLNLQLGGNLLAGDIPPVGALQALRSLNLSSN 652
Query: 266 QLTGSLPETLLKESSMMLSELDLS 289
+L G LP L K ML ELD+S
Sbjct: 653 KLNGQLPIDLGK--LKMLEELDVS 674
Score = 80.5 bits (197), Expect = 4e-15
Identities = 71/236 (30%), Positives = 107/236 (45%), Gaps = 8/236 (3%)
Query: 42 LESNGCPQNWYGILCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM- 100
L SN N L + N+ +++L SL+G F +S +P L + N GS+
Sbjct: 123 LSSNSFTGNIPDTLGALQNLRNLSLFFNSLIGPFPESLLS-IPHLETVYFTGNGLNGSIP 181
Query: 101 LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYL 160
+I M L L L N+F+G +P S + +L L L+ N GT+P + L+ L YL
Sbjct: 182 SNIGNMSELTTLWLDDNQFSGPVPSSLGNITTLQELYLNDNNLVGTLPVTLNNLENLVYL 241
Query: 161 DFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVG 220
D +NS G I F + + LSNN+F+G L GLG+ + L + + + +
Sbjct: 242 DVRNNSLVGAIPLDFVSCKQIDTISLSNNQFTGGLPPGLGNCTSLREFGAFSCALSGPIP 301
Query: 221 ELFAHDGMPYLDNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACNQLTGSLPETL 275
F L L+ + N G I P S+ L+L NQL G +P L
Sbjct: 302 SCFGQ-----LTKLDTLYLAGNHFSGRIPPELGKCKSMIDLQLQQNQLEGEIPGEL 352
Score = 80.1 bits (196), Expect = 6e-15
Identities = 73/235 (31%), Positives = 108/235 (45%), Gaps = 17/235 (7%)
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNK 118
++I + L L GE + L L L + N+ +G + L I ++SL+ L L N
Sbjct: 333 SMIDLQLQQNQLEGEIPG-ELGMLSQLQYLHLYTNNLSGEVPLSIWKIQSLQSLQLYQNN 391
Query: 119 FNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM 178
+G LP EL+ LV L L N F+G +P LE LD N F+G I
Sbjct: 392 LSGELPVDMTELKQLVSLALYENHFTGVIPQDLGANSSLEVLDLTRNMFTGHIPPNLCSQ 451
Query: 179 GSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYL---DNLE 235
+ + L N G++ LG S +++ L + N+L G G+P NL
Sbjct: 452 KKLKRLLLGYNYLEGSVPSDLGGCS---TLERLILEENNLRG------GLPDFVEKQNLL 502
Query: 236 VFDASNNQLVGNI-PSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
FD S N G I PS + ++ + L+ NQL+GS+P L S + L L+LS
Sbjct: 503 FFDLSGNNFTGPIPPSLGNLKNVTAIYLSSNQLSGSIPPEL--GSLVKLEHLNLS 555
Score = 78.2 bits (191), Expect = 2e-14
Identities = 68/235 (28%), Positives = 112/235 (46%), Gaps = 11/235 (4%)
Query: 57 SEGNVISIT---LDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFL 112
S GN+ ++ L++ +LVG + ++NL L L V NN G++ L K + +
Sbjct: 207 SLGNITTLQELYLNDNNLVGTLP-VTLNNLENLVYLDVRNNSLVGAIPLDFVSCKQIDTI 265
Query: 113 DLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIM 172
LS N+F G LPP SL SG +P+ F +L +L+ L N FSG I
Sbjct: 266 SLSNNQFTGGLPPGLGNCTSLREFGAFSCALSGPIPSCFGQLTKLDTLYLAGNHFSGRIP 325
Query: 173 EIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLD 232
+ S++ + L N+ G + G++ L +Q+L++ N+L GE+ + +
Sbjct: 326 PELGKCKSMIDLQLQQNQLEGEIP---GELGMLSQLQYLHLYTNNLSGEVPL--SIWKIQ 380
Query: 233 NLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSEL 286
+L+ N L G +P T + L L L N TG +P+ L SS+ + +L
Sbjct: 381 SLQSLQLYQNNLSGELPVDMTELKQLVSLALYENHFTGVIPQDLGANSSLEVLDL 435
Score = 77.8 bits (190), Expect = 3e-14
Identities = 64/242 (26%), Positives = 106/242 (43%), Gaps = 32/242 (13%)
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH----------------- 102
N++ + + N SLVG +S + +S+ NN FTG +
Sbjct: 237 NLVYLDVRNNSLVGAIPLDFVS-CKQIDTISLSNNQFTGGLPPGLGNCTSLREFGAFSCA 295
Query: 103 --------ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKL 154
+ L L L+ N F+G +PP + +S++ L L N+ G +P L
Sbjct: 296 LSGPIPSCFGQLTKLDTLYLAGNHFSGRIPPELGKCKSMIDLQLQQNQLEGEIPGELGML 355
Query: 155 DQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVS 214
QL+YL ++N+ SG++ +++ S+ + L N SG L + + ++ L S L +
Sbjct: 356 SQLQYLHLYTNNLSGEVPLSIWKIQSLQSLQLYQNNLSGELPVDMTELKQLVS---LALY 412
Query: 215 HNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACNQLTGSLPE 273
N G + G +LEV D + N G+I P+ L+ L L N L GS+P
Sbjct: 413 ENHFTGVIPQDLGAN--SSLEVLDLTRNMFTGHIPPNLCSQKKLKRLLLGYNYLEGSVPS 470
Query: 274 TL 275
L
Sbjct: 471 DL 472
Score = 72.8 bits (177), Expect = 9e-13
Identities = 58/191 (30%), Positives = 95/191 (49%), Gaps = 9/191 (4%)
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNK 118
NV +I L + L G + +L L +L++ +N G + +S L LD S N
Sbjct: 524 NVTAIYLSSNQLSGSIP-PELGSLVKLEHLNLSHNILKGILPSELSNCHKLSELDASHNL 582
Query: 119 FNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM 178
NGS+P + L L L+L N FSG +P + ++L L N +GDI + +
Sbjct: 583 LNGSIPSTLGSLTELTKLSLGENSFSGGIPTSLFQSNKLLNLQLGGNLLAGDIPPVG-AL 641
Query: 179 GSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFD 238
++ ++LS+NK +G L + LG + L + L+VSHN+L G L + + +L +
Sbjct: 642 QALRSLNLSSNKLNGQLPIDLGKLKML---EELDVSHNNLSGTLRV---LSTIQSLTFIN 695
Query: 239 ASNNQLVGNIP 249
S+N G +P
Sbjct: 696 ISHNLFSGPVP 706
Score = 51.2 bits (121), Expect = 3e-06
Identities = 35/112 (31%), Positives = 56/112 (49%), Gaps = 6/112 (5%)
Query: 86 LHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSG 145
L NL + N G + + +++L+ L+LS NK NG LP +L+ L L++S N SG
Sbjct: 621 LLNLQLGGNLLAGDIPPVGALQALRSLNLSSNKLNGQLPIDLGKLKMLEELDVSHNNLSG 680
Query: 146 TVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
T+ V + L +++ N FSG + + ++ S FSG DL
Sbjct: 681 TL-RVLSTIQSLTFINISHNLFSGPVPPSLTKF-----LNSSPTSFSGNSDL 726
Score = 47.4 bits (111), Expect = 4e-05
Identities = 28/83 (33%), Positives = 48/83 (57%), Gaps = 6/83 (7%)
Query: 63 SITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGS 122
S+ L + L G+ + + L ML L V +N+ +G++ +S ++SL F+++S N F+G
Sbjct: 646 SLNLSSNKLNGQLP-IDLGKLKMLEELDVSHNNLSGTLRVLSTIQSLTFINISHNLFSGP 704
Query: 123 LPPSFVELRSLVYLNLSLNEFSG 145
+PPS + +LN S FSG
Sbjct: 705 VPPSLTK-----FLNSSPTSFSG 722
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 96.3 bits (238), Expect = 8e-20
Identities = 87/315 (27%), Positives = 142/315 (44%), Gaps = 39/315 (12%)
Query: 2 LLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSE-GN 60
L L + F D++ LL LK + P G L+ W S C ++ G+ C +
Sbjct: 15 LYLFFSPCFAYTDMEVLLNLKSSMIG-PKGHGLHDWIHSSSPDAHC--SFSGVSCDDDAR 71
Query: 61 VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLN-- 117
VIS+ + L G + I L L NL++ N+FTG + L + + SLK L++S N
Sbjct: 72 VISLNVSFTPLFGTIS-PEIGMLTHLVNLTLAANNFTGELPLEMKSLTSLKVLNISNNGN 130
Query: 118 ------------------------KFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHK 153
FNG LPP EL+ L YL+ N FSG +P +
Sbjct: 131 LTGTFPGEILKAMVDLEVLDTYNNNFNGKLPPEMSELKKLKYLSFGGNFFSGEIPESYGD 190
Query: 154 LDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLS-NNKFSGALDLGLGDVSFLFSIQHLN 212
+ LEYL + SG ++ ++ + + N ++G + G L ++ L+
Sbjct: 191 IQSLEYLGLNGAGLSGKSPAFLSRLKNLREMYIGYYNSYTGGVPPEFGG---LTKLEILD 247
Query: 213 VSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACNQLTGSL 271
++ +L GE+ + L +L N L G+I P + +VSL+ L L+ NQLTG +
Sbjct: 248 MASCTLTGEI--PTSLSNLKHLHTLFLHINNLTGHIPPELSGLVSLKSLDLSINQLTGEI 305
Query: 272 PETLLKESSMMLSEL 286
P++ + ++ L L
Sbjct: 306 PQSFINLGNITLINL 320
Score = 82.8 bits (203), Expect = 9e-16
Identities = 70/230 (30%), Positives = 118/230 (50%), Gaps = 17/230 (7%)
Query: 66 LDNAS--LVGEFNFLAISNLPMLHNLSVVNNHFTGSML-HISPMKSLKFLDLSLNKFNGS 122
LD AS L GE ++SNL LH L + N+ TG + +S + SLK LDLS+N+ G
Sbjct: 246 LDMASCTLTGEIP-TSLSNLKHLHTLFLHINNLTGHIPPELSGLVSLKSLDLSINQLTGE 304
Query: 123 LPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVL 182
+P SF+ L ++ +NL N G +P +L +LE + N+F+ + + G+++
Sbjct: 305 IPQSFINLGNITLINLFRNNLYGQIPEAIGELPKLEVFEVWENNFTLQLPANLGRNGNLI 364
Query: 183 HVDLSNNKFSGAL--DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDAS 240
+D+S+N +G + DL G+ ++ L +S+N G + + + +L
Sbjct: 365 KLDVSDNHLTGLIPKDLCRGE-----KLEMLILSNNFFFGPI--PEELGKCKSLTKIRIV 417
Query: 241 NNQLVGNIPSFTFVVSL-RILRLACNQLTGSLPETLLKESSMMLSELDLS 289
N L G +P+ F + L I+ L N +G LP T+ S +L ++ LS
Sbjct: 418 KNLLNGTVPAGLFNLPLVTIIELTDNFFSGELPVTM---SGDVLDQIYLS 464
Score = 78.2 bits (191), Expect = 2e-14
Identities = 48/168 (28%), Positives = 85/168 (50%), Gaps = 5/168 (2%)
Query: 82 NLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLN 141
NLP++ + + +N F+G + L + LS N F+G +PP+ +L L L N
Sbjct: 431 NLPLVTIIELTDNFFSGELPVTMSGDVLDQIYLSNNWFSGEIPPAIGNFPNLQTLFLDRN 490
Query: 142 EFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGD 201
F G +P +L L ++ +N+ +G I + + +++ VDLS N+ +G + G+ +
Sbjct: 491 RFRGNIPREIFELKHLSRINTSANNITGGIPDSISRCSTLISVDLSRNRINGEIPKGINN 550
Query: 202 VSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP 249
V ++ LN+S N L G + G+ + +L D S N L G +P
Sbjct: 551 VK---NLGTLNISGNQLTGSI--PTGIGNMTSLTTLDLSFNDLSGRVP 593
Score = 71.6 bits (174), Expect = 2e-12
Identities = 70/228 (30%), Positives = 105/228 (45%), Gaps = 30/228 (13%)
Query: 86 LHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFS 144
L L V +NH TG + + + L+ L LS N F G +P + +SL + + N +
Sbjct: 363 LIKLDVSDNHLTGLIPKDLCRGEKLEMLILSNNFFFGPIPEELGKCKSLTKIRIVKNLLN 422
Query: 145 GTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVL-HVDLSNNKFSGALDLGLGDVS 203
GTVP L + ++ N FSG++ G VL + LSNN FSG + +G+
Sbjct: 423 GTVPAGLFNLPLVTIIELTDNFFSGELPVTM--SGDVLDQIYLSNNWFSGEIPPAIGNFP 480
Query: 204 FL------------------FSIQHL---NVSHNSLVGELFAHDGMPYLDNLEVFDASNN 242
L F ++HL N S N++ G + D + L D S N
Sbjct: 481 NLQTLFLDRNRFRGNIPREIFELKHLSRINTSANNITGGI--PDSISRCSTLISVDLSRN 538
Query: 243 QLVGNIPS-FTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
++ G IP V +L L ++ NQLTGS+P + +S L+ LDLS
Sbjct: 539 RINGEIPKGINNVKNLGTLNISGNQLTGSIPTGIGNMTS--LTTLDLS 584
Score = 61.2 bits (147), Expect = 3e-09
Identities = 38/121 (31%), Positives = 58/121 (47%), Gaps = 1/121 (0%)
Query: 79 AISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
AI N P L L + N F G++ I +K L ++ S N G +P S +L+ ++
Sbjct: 475 AIGNFPNLQTLFLDRNRFRGNIPREIFELKHLSRINTSANNITGGIPDSISRCSTLISVD 534
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
LS N +G +P + + L L+ N +G I M S+ +DLS N SG + L
Sbjct: 535 LSRNRINGEIPKGINNVKNLGTLNISGNQLTGSIPTGIGNMTSLTTLDLSFNDLSGRVPL 594
Query: 198 G 198
G
Sbjct: 595 G 595
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 95.9 bits (237), Expect = 1e-19
Identities = 78/273 (28%), Positives = 136/273 (49%), Gaps = 24/273 (8%)
Query: 7 NTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISITL 66
N + D+ AL +G+++ G N +S S SN C +W GI C +S+ L
Sbjct: 26 NLTCNSNDLKALEGFMRGLESSIDGWKWN--ESSSFSSNCC--DWVGISCKSS--VSLGL 79
Query: 67 DNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPS 126
D+ + G L + + LS ++ + LK L+L+ N +GS+ S
Sbjct: 80 DDVNESGRVVELELGRRKLSGKLSE----------SVAKLDQLKVLNLTHNSLSGSIAAS 129
Query: 127 FVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI-MEIFYQMGSVLHVD 185
+ L +L L+LS N+FSG P++ + L L L+ + NSF G I + + + +D
Sbjct: 130 LLNLSNLEVLDLSSNDFSGLFPSLIN-LPSLRVLNVYENSFHGLIPASLCNNLPRIREID 188
Query: 186 LSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLV 245
L+ N F G++ +G+G+ S S+++L ++ N+L G + + L NL V NN+L
Sbjct: 189 LAMNYFDGSIPVGIGNCS---SVEYLGLASNNLSGSI--PQELFQLSNLSVLALQNNRLS 243
Query: 246 GNIPSFTFVVS-LRILRLACNQLTGSLPETLLK 277
G + S +S L L ++ N+ +G +P+ L+
Sbjct: 244 GALSSKLGKLSNLGRLDISSNKFSGKIPDVFLE 276
Score = 78.2 bits (191), Expect = 2e-14
Identities = 72/264 (27%), Positives = 122/264 (45%), Gaps = 40/264 (15%)
Query: 62 ISITLDNASLVGEFNFLAISNLP--------MLHNLSVV---NNHFTGSMLH-ISPMKSL 109
I + + N S V E+ LA +NL L NLSV+ NN +G++ + + +L
Sbjct: 198 IPVGIGNCSSV-EYLGLASNNLSGSIPQELFQLSNLSVLALQNNRLSGALSSKLGKLSNL 256
Query: 110 KFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSG 169
LD+S NKF+G +P F+EL L Y + N F+G +P + L +N+ SG
Sbjct: 257 GRLDISSNKFSGKIPDVFLELNKLWYFSAQSNLFNGEMPRSLSNSRSISLLSLRNNTLSG 316
Query: 170 DIMEIFYQMGSVLHVDLSNNKFSGAL-----------DLGLGDVSFLFSIQHLNVSHNSL 218
I M ++ +DL++N FSG++ + + F+ I + SL
Sbjct: 317 QIYLNCSAMTNLTSLDLASNSFSGSIPSNLPNCLRLKTINFAKIKFIAQIPESFKNFQSL 376
Query: 219 VGELFAHDGMPYLDN-LEVFDASNN------------QLVGNIPSFTFVVSLRILRLACN 265
F++ + + + LE+ N + + ++PS F +L++L +A
Sbjct: 377 TSLSFSNSSIQNISSALEILQHCQNLKTLVLTLNFQKEELPSVPSLQF-KNLKVLIIASC 435
Query: 266 QLTGSLPETLLKESSMMLSELDLS 289
QL G++P+ L S+ L LDLS
Sbjct: 436 QLRGTVPQWLSNSPSLQL--LDLS 457
Score = 77.8 bits (190), Expect = 3e-14
Identities = 58/203 (28%), Positives = 100/203 (48%), Gaps = 10/203 (4%)
Query: 77 FLAISNLPMLHNLSVVNNHFTGSMLH--ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLV 134
F ++ NLP L L+V N F G + + + ++ +DL++N F+GS+P S+
Sbjct: 150 FPSLINLPSLRVLNVYENSFHGLIPASLCNNLPRIREIDLAMNYFDGSIPVGIGNCSSVE 209
Query: 135 YLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGA 194
YL L+ N SG++P +L L L +N SG + ++ ++ +D+S+NKFSG
Sbjct: 210 YLGLASNNLSGSIPQELFQLSNLSVLALQNNRLSGALSSKLGKLSNLGRLDISSNKFSGK 269
Query: 195 LDLGLGDVSF-LFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNI-PSFT 252
+ DV L + + + N GE+ + ++ + NN L G I + +
Sbjct: 270 IP----DVFLELNKLWYFSAQSNLFNGEM--PRSLSNSRSISLLSLRNNTLSGQIYLNCS 323
Query: 253 FVVSLRILRLACNQLTGSLPETL 275
+ +L L LA N +GS+P L
Sbjct: 324 AMTNLTSLDLASNSFSGSIPSNL 346
Score = 71.2 bits (173), Expect = 3e-12
Identities = 55/183 (30%), Positives = 94/183 (51%), Gaps = 12/183 (6%)
Query: 107 KSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNS 166
K+LK L ++ + G++P SL L+LS N+ SGT+P L+ L YLD +N+
Sbjct: 425 KNLKVLIIASCQLRGTVPQWLSNSPSLQLLDLSWNQLSGTIPPWLGSLNSLFYLDLSNNT 484
Query: 167 FSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQH---------LNVSHNS 217
F G+I + S++ + + + S + +Q+ +++S+NS
Sbjct: 485 FIGEIPHSLTSLQSLVSKENAVEEPSPDFPFFKKKNTNAGGLQYNQPSSFPPMIDLSYNS 544
Query: 218 LVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLL 276
L G ++ G L L V + NN L GNIP + + + SL +L L+ N L+G++P +L+
Sbjct: 545 LNGSIWPEFG--DLRQLHVLNLKNNNLSGNIPANLSGMTSLEVLDLSHNNLSGNIPPSLV 602
Query: 277 KES 279
K S
Sbjct: 603 KLS 605
Score = 62.8 bits (151), Expect = 1e-09
Identities = 52/188 (27%), Positives = 79/188 (41%), Gaps = 20/188 (10%)
Query: 25 IQNDPFGLVLN-SWDSKSLESNGCPQNWYGILCSEGNVISITLDNASLVGEFNFLAISNL 83
+ N P +L+ SW+ S G W G L S + + L N + +GE S
Sbjct: 445 LSNSPSLQLLDLSWNQLS----GTIPPWLGSLNS---LFYLDLSNNTFIGEIPHSLTSLQ 497
Query: 84 PMLHNLSVVN------------NHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELR 131
++ + V N G + + P +DLS N NGS+ P F +LR
Sbjct: 498 SLVSKENAVEEPSPDFPFFKKKNTNAGGLQYNQPSSFPPMIDLSYNSLNGSIWPEFGDLR 557
Query: 132 SLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKF 191
L LNL N SG +P + LE LD N+ SG+I ++ + ++ NK
Sbjct: 558 QLHVLNLKNNNLSGNIPANLSGMTSLEVLDLSHNNLSGNIPPSLVKLSFLSTFSVAYNKL 617
Query: 192 SGALDLGL 199
SG + G+
Sbjct: 618 SGPIPTGV 625
Score = 59.7 bits (143), Expect = 8e-09
Identities = 50/155 (32%), Positives = 74/155 (47%), Gaps = 6/155 (3%)
Query: 133 LVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFS 192
+V L L + SG + KLDQL+ L+ NS SG I + ++ +DLS+N FS
Sbjct: 88 VVELELGRRKLSGKLSESVAKLDQLKVLNLTHNSLSGSIAASLLNLSNLEVLDLSSNDFS 147
Query: 193 GALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SF 251
G + L S++ LNV NS G L L + D + N G+IP
Sbjct: 148 GLFP----SLINLPSLRVLNVYENSFHG-LIPASLCNNLPRIREIDLAMNYFDGSIPVGI 202
Query: 252 TFVVSLRILRLACNQLTGSLPETLLKESSMMLSEL 286
S+ L LA N L+GS+P+ L + S++ + L
Sbjct: 203 GNCSSVEYLGLASNNLSGSIPQELFQLSNLSVLAL 237
Score = 57.0 bits (136), Expect = 5e-08
Identities = 51/180 (28%), Positives = 80/180 (44%), Gaps = 40/180 (22%)
Query: 80 ISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLV---- 134
+SN P L L + N +G++ + + SL +LDLS N F G +P S L+SLV
Sbjct: 445 LSNSPSLQLLDLSWNQLSGTIPPWLGSLNSLFYLDLSNNTFIGEIPHSLTSLQSLVSKEN 504
Query: 135 --------------------------------YLNLSLNEFSGTVPNVFHKLDQLEYLDF 162
++LS N +G++ F L QL L+
Sbjct: 505 AVEEPSPDFPFFKKKNTNAGGLQYNQPSSFPPMIDLSYNSLNGSIWPEFGDLRQLHVLNL 564
Query: 163 HSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGEL 222
+N+ SG+I M S+ +DLS+N SG + L +SFL + +V++N L G +
Sbjct: 565 KNNNLSGNIPANLSGMTSLEVLDLSHNNLSGNIPPSLVKLSFLST---FSVAYNKLSGPI 621
>PGI2_ARATH (Q9M5J8) Polygalacturonase inhibitor 2 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2)
Length = 330
Score = 95.5 bits (236), Expect = 1e-19
Identities = 81/290 (27%), Positives = 138/290 (46%), Gaps = 45/290 (15%)
Query: 14 DIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNV----ISITLDNA 69
D LL++KK + N+P+ L SWD K+ + C +WY + C + V S+ + +
Sbjct: 29 DKTTLLKIKKSL-NNPYHLA--SWDPKT---DCC--SWYCLECGDATVNHRVTSLIIQDG 80
Query: 70 SLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISP----MKSLKFLDLSLNKFNGSLPP 125
+ G+ + +LP L S++ T HI P +K+L FL LS G +P
Sbjct: 81 EISGQIP-PEVGDLPYL--TSLIFRKLTNLTGHIQPTIAKLKNLTFLRLSWTNLTGPVPE 137
Query: 126 SFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM-GSVLHV 184
+L++L Y++LS N+ SG++P+ L +LEYL+ N +G I E F G V +
Sbjct: 138 FLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRNKLTGPIPESFGTFSGKVPSL 197
Query: 185 DLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGE-------------------LFAH 225
LS+N+ SG + LG+ F +++S N L G+ +F
Sbjct: 198 FLSHNQLSGTIPKSLGNPDF----YRIDLSRNKLQGDASILFGAKKTTWIVDISRNMFQF 253
Query: 226 D--GMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPE 273
D + L D ++N + G+IP+ ++L ++ N+L G +P+
Sbjct: 254 DLSKVKLAKTLNNLDMNHNGITGSIPAEWSKAYFQLLNVSYNRLCGRIPK 303
Score = 39.3 bits (90), Expect = 0.011
Identities = 32/93 (34%), Positives = 49/93 (52%), Gaps = 7/93 (7%)
Query: 202 VSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS-FTFVVSLRIL 260
++ L ++ L +S +L G + + + L NLE D S N L G+IPS + + L L
Sbjct: 115 IAKLKNLTFLRLSWTNLTGPV--PEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYL 172
Query: 261 RLACNQLTGSLPETL----LKESSMMLSELDLS 289
L+ N+LTG +PE+ K S+ LS LS
Sbjct: 173 ELSRNKLTGPIPESFGTFSGKVPSLFLSHNQLS 205
Score = 32.3 bits (72), Expect = 1.4
Identities = 30/95 (31%), Positives = 48/95 (49%), Gaps = 6/95 (6%)
Query: 197 LGLGDVSFLFSIQHLNVSHNSLVGELFAHDG-MPYLDNLEVFDASNNQLVGNI-PSFTFV 254
L GD + + L + + G++ G +PYL +L +F N L G+I P+ +
Sbjct: 61 LECGDATVNHRVTSLIIQDGEISGQIPPEVGDLPYLTSL-IFRKLTN-LTGHIQPTIAKL 118
Query: 255 VSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+L LRL+ LTG +PE L + + L +DLS
Sbjct: 119 KNLTFLRLSWTNLTGPVPEFLSQLKN--LEYIDLS 151
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 92.4 bits (228), Expect = 1e-18
Identities = 90/304 (29%), Positives = 141/304 (45%), Gaps = 44/304 (14%)
Query: 14 DIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGN---VISITLDNAS 70
D++AL + ++ P G W + S ++ C NW GI C+ N VI + L N
Sbjct: 35 DLEALRDFIAHLEPKPDG-----WINSSSSTDCC--NWTGITCNSNNTGRVIRLELGNKK 87
Query: 71 LVGEF-----------------NF------LAISNLPMLHNLSVVNNHFTGSMLHISPMK 107
L G+ NF L+I NL L L + +N +G + +
Sbjct: 88 LSGKLSESLGKLDEIRVLNLSRNFIKDSIPLSIFNLKNLQTLDLSSNDLSGGIPTSINLP 147
Query: 108 SLKFLDLSLNKFNGSLPPSFVELRSLV-YLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNS 166
+L+ DLS NKFNGSLP + + + L++N F+G + F K LE+L N
Sbjct: 148 ALQSFDLSSNKFNGSLPSHICHNSTQIRVVKLAVNYFAGNFTSGFGKCVLLEHLCLGMND 207
Query: 167 FSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHD 226
+G+I E + + + + + N+ SG+L + ++S S+ L+VS N GE+ D
Sbjct: 208 LTGNIPEDLFHLKRLNLLGIQENRLSGSLSREIRNLS---SLVRLDVSWNLFSGEI--PD 262
Query: 227 GMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSMM-LS 284
L L+ F N +G IP S SL +L L N L+G L +L ++M+ L+
Sbjct: 263 VFDELPQLKFFLGQTNGFIGGIPKSLANSPSLNLLNLRNNSLSGRL---MLNCTAMIALN 319
Query: 285 ELDL 288
LDL
Sbjct: 320 SLDL 323
Score = 75.1 bits (183), Expect = 2e-13
Identities = 63/208 (30%), Positives = 105/208 (50%), Gaps = 10/208 (4%)
Query: 79 AISNLPMLHNLSVVNNHFTGS-MLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
+++N P L+ L++ NN +G ML+ + M +L LDL N+FNG LP + + + L +N
Sbjct: 287 SLANSPSLNLLNLRNNSLSGRLMLNCTAMIALNSLDLGTNRFNGRLPENLPDCKRLKNVN 346
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFS--GDIMEIFYQMGSVLHVDLSNNKFSGAL 195
L+ N F G VP F + L Y ++S + + I ++ + L+ N AL
Sbjct: 347 LARNTFHGQVPESFKNFESLSYFSLSNSSLANISSALGILQHCKNLTTLVLTLNFHGEAL 406
Query: 196 DLGLGDVSFLF-SIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFT-F 253
D S F ++ L V++ L G + + + L++ D S N+L G IPS+
Sbjct: 407 P---DDSSLHFEKLKVLVVANCRLTGSM--PRWLSSSNELQLLDLSWNRLTGAIPSWIGD 461
Query: 254 VVSLRILRLACNQLTGSLPETLLKESSM 281
+L L L+ N TG +P++L K S+
Sbjct: 462 FKALFYLDLSNNSFTGEIPKSLTKLESL 489
Score = 74.3 bits (181), Expect = 3e-13
Identities = 63/192 (32%), Positives = 92/192 (47%), Gaps = 25/192 (13%)
Query: 89 LSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTV 147
L V N TGSM +S L+ LDLS N+ G++P + ++L YL+LS N F+G +
Sbjct: 420 LVVANCRLTGSMPRWLSSSNELQLLDLSWNRLTGAIPSWIGDFKALFYLDLSNNSFTGEI 479
Query: 148 PNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSG---ALDLGLGDVSF 204
P KL+ L + N S D F+ + L N+ G ++LG
Sbjct: 480 PKSLTKLESLTSRNISVNEPSPDFP--FFMKRNESARALQYNQIFGFPPTIELG------ 531
Query: 205 LFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLA 263
HN+L G ++ G L L VFD N L G+IP S + + SL L L+
Sbjct: 532 ----------HNNLSGPIWEEFG--NLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLS 579
Query: 264 CNQLTGSLPETL 275
N+L+GS+P +L
Sbjct: 580 NNRLSGSIPVSL 591
Score = 70.1 bits (170), Expect = 6e-12
Identities = 66/237 (27%), Positives = 106/237 (43%), Gaps = 36/237 (15%)
Query: 82 NLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSL 140
+L L+ L + N +GS+ I + SL LD+S N F+G +P F EL L +
Sbjct: 218 HLKRLNLLGIQENRLSGSLSREIRNLSSLVRLDVSWNLFSGEIPDVFDELPQLKFFLGQT 277
Query: 141 NEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLG 200
N F G +P L L+ +NS SG +M M ++ +DL N+F+G L L
Sbjct: 278 NGFIGGIPKSLANSPSLNLLNLRNNSLSGRLMLNCTAMIALNSLDLGTNRFNGRLPENLP 337
Query: 201 DVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVG------------NI 248
D L +++N++ N+ G++ + ++L F SN+ L N+
Sbjct: 338 DCKRL---KNVNLARNTFHGQV--PESFKNFESLSYFSLSNSSLANISSALGILQHCKNL 392
Query: 249 PSFTFVVS----------------LRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+ ++ L++L +A +LTGS+P L SS L LDLS
Sbjct: 393 TTLVLTLNFHGEALPDDSSLHFEKLKVLVVANCRLTGSMPRWL--SSSNELQLLDLS 447
Score = 68.6 bits (166), Expect = 2e-11
Identities = 55/185 (29%), Positives = 92/185 (49%), Gaps = 7/185 (3%)
Query: 85 MLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEF 143
+L +L + N TG++ + +K L L + N+ +GSL L SLV L++S N F
Sbjct: 197 LLEHLCLGMNDLTGNIPEDLFHLKRLNLLGIQENRLSGSLSREIRNLSSLVRLDVSWNLF 256
Query: 144 SGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVS 203
SG +P+VF +L QL++ +N F G I + S+ ++L NN SG L L + +
Sbjct: 257 SGEIPDVFDELPQLKFFLGQTNGFIGGIPKSLANSPSLNLLNLRNNSLSGRLML---NCT 313
Query: 204 FLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRL 262
+ ++ L++ N G L + +P L+ + + N G +P SF SL L
Sbjct: 314 AMIALNSLDLGTNRFNGRL--PENLPDCKRLKNVNLARNTFHGQVPESFKNFESLSYFSL 371
Query: 263 ACNQL 267
+ + L
Sbjct: 372 SNSSL 376
Score = 64.7 bits (156), Expect = 3e-10
Identities = 58/191 (30%), Positives = 84/191 (43%), Gaps = 42/191 (21%)
Query: 64 ITLDNASLVGEF-NFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNG 121
+ + N L G +L+ SN L +LS N TG++ I K+L +LDLS N F G
Sbjct: 420 LVVANCRLTGSMPRWLSSSNELQLLDLSW--NRLTGAIPSWIGDFKALFYLDLSNNSFTG 477
Query: 122 SLPPSFVELRSLVYLNLSLNE------------------------------------FSG 145
+P S +L SL N+S+NE SG
Sbjct: 478 EIPKSLTKLESLTSRNISVNEPSPDFPFFMKRNESARALQYNQIFGFPPTIELGHNNLSG 537
Query: 146 TVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFL 205
+ F L +L D N+ SG I M S+ +DLSNN+ SG++ + L +SFL
Sbjct: 538 PIWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGSIPVSLQQLSFL 597
Query: 206 --FSIQHLNVS 214
FS+ + N+S
Sbjct: 598 SKFSVAYNNLS 608
Score = 62.4 bits (150), Expect = 1e-09
Identities = 51/192 (26%), Positives = 93/192 (47%), Gaps = 4/192 (2%)
Query: 83 LPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLN 141
LP L N F G + ++ SL L+L N +G L + + +L L+L N
Sbjct: 267 LPQLKFFLGQTNGFIGGIPKSLANSPSLNLLNLRNNSLSGRLMLNCTAMIALNSLDLGTN 326
Query: 142 EFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGD 201
F+G +P +L+ ++ N+F G + E F S+ + LSN+ + + LG
Sbjct: 327 RFNGRLPENLPDCKRLKNVNLARNTFHGQVPESFKNFESLSYFSLSNSSLAN-ISSALGI 385
Query: 202 VSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSF-TFVVSLRIL 260
+ ++ L ++ N GE D + + L+V +N +L G++P + + L++L
Sbjct: 386 LQHCKNLTTLVLTLN-FHGEALPDDSSLHFEKLKVLVVANCRLTGSMPRWLSSSNELQLL 444
Query: 261 RLACNQLTGSLP 272
L+ N+LTG++P
Sbjct: 445 DLSWNRLTGAIP 456
Score = 53.5 bits (127), Expect = 6e-07
Identities = 71/287 (24%), Positives = 115/287 (39%), Gaps = 65/287 (22%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISP-MKSLKFLDLSLNKFNGS 122
+ L N SL G L + + L++L + N F G + P K LK ++L+ N F+G
Sbjct: 297 LNLRNNSLSGRL-MLNCTAMIALNSLDLGTNRFNGRLPENLPDCKRLKNVNLARNTFHGQ 355
Query: 123 LPPSFVELRSLVYLNLS--------------------------LN--------------- 141
+P SF SL Y +LS LN
Sbjct: 356 VPESFKNFESLSYFSLSNSSLANISSALGILQHCKNLTTLVLTLNFHGEALPDDSSLHFE 415
Query: 142 ----------EFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKF 191
+G++P ++L+ LD N +G I ++ ++DLSNN F
Sbjct: 416 KLKVLVVANCRLTGSMPRWLSSSNELQLLDLSWNRLTGAIPSWIGDFKALFYLDLSNNSF 475
Query: 192 SGALDLGLGDVSFLFSIQHLNVSHNSLVGELFA--HDGMPYLDNLEVF------DASNNQ 243
+G + L + L S ++++V+ S F ++ L ++F + +N
Sbjct: 476 TGEIPKSLTKLESLTS-RNISVNEPSPDFPFFMKRNESARALQYNQIFGFPPTIELGHNN 534
Query: 244 LVGNI-PSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
L G I F + L + L N L+GS+P +L +S L LDLS
Sbjct: 535 LSGPIWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTS--LEALDLS 579
Score = 44.3 bits (103), Expect = 4e-04
Identities = 26/69 (37%), Positives = 40/69 (57%), Gaps = 1/69 (1%)
Query: 82 NLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSL 140
NL LH + N +GS+ +S M SL+ LDLS N+ +GS+P S +L L +++
Sbjct: 545 NLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGSIPVSLQQLSFLSKFSVAY 604
Query: 141 NEFSGTVPN 149
N SG +P+
Sbjct: 605 NNLSGVIPS 613
Score = 43.5 bits (101), Expect = 6e-04
Identities = 24/87 (27%), Positives = 40/87 (45%)
Query: 112 LDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDI 171
++L N +G + F L+ L +L N SG++P+ + LE LD +N SG I
Sbjct: 528 IELGHNNLSGPIWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGSI 587
Query: 172 MEIFYQMGSVLHVDLSNNKFSGALDLG 198
Q+ + ++ N SG + G
Sbjct: 588 PVSLQQLSFLSKFSVAYNNLSGVIPSG 614
Score = 41.2 bits (95), Expect = 0.003
Identities = 24/66 (36%), Positives = 33/66 (49%)
Query: 106 MKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSN 165
+K L DL N +GS+P S + SL L+LS N SG++P +L L N
Sbjct: 546 LKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGSIPVSLQQLSFLSKFSVAYN 605
Query: 166 SFSGDI 171
+ SG I
Sbjct: 606 NLSGVI 611
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells
protein) (EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 92.4 bits (228), Expect = 1e-18
Identities = 87/300 (29%), Positives = 141/300 (47%), Gaps = 60/300 (20%)
Query: 17 ALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASLVGEFN 76
+L+ K+ ++N +L+SW+ S S+ C +W G+ C G V S++L + SL G+
Sbjct: 29 SLISFKRSLENPS---LLSSWNVSSSASH-C--DWVGVTCLLGRVNSLSLPSLSLRGQ-- 80
Query: 77 FLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYL 136
+P IS +K+L+ L L+ N+F+G +PP L+ L L
Sbjct: 81 ------IPK----------------EISSLKNLRELCLAGNQFSGKIPPEIWNLKHLQTL 118
Query: 137 NLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFY-QMGSVLHVDLSNNKFSGAL 195
+LS N +G +P + +L QL YLD N FSG + F+ + ++ +D+SNN SG +
Sbjct: 119 DLSGNSLTGLLPRLLSELPQLLYLDLSDNHFSGSLPPSFFISLPALSSLDVSNNSLSGEI 178
Query: 196 DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDG----------------------MPYLDN 233
+G +S ++ +L + NS G++ + G + L +
Sbjct: 179 PPEIGKLS---NLSNLYMGLNSFSGQIPSEIGNISLLKNFAAPSCFFNGPLPKEISKLKH 235
Query: 234 LEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL---LKESSMMLSELDLS 289
L D S N L +IP SF + +L IL L +L G +P L S+MLS LS
Sbjct: 236 LAKLDLSYNPLKCSIPKSFGELHNLSILNLVSAELIGLIPPELGNCKSLKSLMLSFNSLS 295
Score = 89.0 bits (219), Expect = 1e-17
Identities = 65/192 (33%), Positives = 102/192 (52%), Gaps = 9/192 (4%)
Query: 61 VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKF 119
++ I+L N L GE ++S L L L + N TGS+ + L+ L+L+ N+
Sbjct: 606 LVEISLSNNHLSGEIP-ASLSRLTNLTILDLSGNALTGSIPKEMGNSLKLQGLNLANNQL 664
Query: 120 NGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMG 179
NG +P SF L SLV LNL+ N+ G VP L +L ++D N+ SG++ M
Sbjct: 665 NGHIPESFGLLGSLVKLNLTKNKLDGPVPASLGNLKELTHMDLSFNNLSGELSSELSTME 724
Query: 180 SVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAH-DGMPYLDNLEVFD 238
++ + + NKF+G + LG+ L +++L+VS N L GE+ G+P NLE +
Sbjct: 725 KLVGLYIEQNKFTGEIPSELGN---LTQLEYLDVSENLLSGEIPTKICGLP---NLEFLN 778
Query: 239 ASNNQLVGNIPS 250
+ N L G +PS
Sbjct: 779 LAKNNLRGEVPS 790
Score = 80.5 bits (197), Expect = 4e-15
Identities = 69/227 (30%), Positives = 113/227 (49%), Gaps = 10/227 (4%)
Query: 63 SITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNG 121
S+ L SL G L +S +P+L S N +GS+ + K L L L+ N+F+G
Sbjct: 286 SLMLSFNSLSGPLP-LELSEIPLL-TFSAERNQLSGSLPSWMGKWKVLDSLLLANNRFSG 343
Query: 122 SLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSV 181
+P + L +L+L+ N SG++P LE +D N SG I E+F S+
Sbjct: 344 EIPHEIEDCPMLKHLSLASNLLSGSIPRELCGSGSLEAIDLSGNLLSGTIEEVFDGCSSL 403
Query: 182 LHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASN 241
+ L+NN+ +G++ L + + L++ N+ GE+ + NL F AS
Sbjct: 404 GELLLTNNQINGSIPEDL----WKLPLMALDLDSNNFTGEI--PKSLWKSTNLMEFTASY 457
Query: 242 NQLVGNIPS-FTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELD 287
N+L G +P+ SL+ L L+ NQLTG +P + K +S+ + L+
Sbjct: 458 NRLEGYLPAEIGNAASLKRLVLSDNQLTGEIPREIGKLTSLSVLNLN 504
Score = 75.5 bits (184), Expect = 1e-13
Identities = 77/249 (30%), Positives = 111/249 (43%), Gaps = 49/249 (19%)
Query: 80 ISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNL 138
I N+ +L N + + F G + IS +K L LDLS N S+P SF EL +L LNL
Sbjct: 206 IGNISLLKNFAAPSCFFNGPLPKEISKLKHLAKLDLSYNPLKCSIPKSFGELHNLSILNL 265
Query: 139 SLNEFSGTVPNVFHKLDQLEYLDFHSNSFSG----DIMEIFY----------------QM 178
E G +P L+ L NS SG ++ EI M
Sbjct: 266 VSAELIGLIPPELGNCKSLKSLMLSFNSLSGPLPLELSEIPLLTFSAERNQLSGSLPSWM 325
Query: 179 GSVLHVD---LSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVG----ELFAHDGMPYL 231
G +D L+NN+FSG + + D L +HL+++ N L G EL + +
Sbjct: 326 GKWKVLDSLLLANNRFSGEIPHEIEDCPML---KHLSLASNLLSGSIPRELCGSGSLEAI 382
Query: 232 D---NL------EVFDA---------SNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPE 273
D NL EVFD +NNQ+ G+IP + + L L L N TG +P+
Sbjct: 383 DLSGNLLSGTIEEVFDGCSSLGELLLTNNQINGSIPEDLWKLPLMALDLDSNNFTGEIPK 442
Query: 274 TLLKESSMM 282
+L K +++M
Sbjct: 443 SLWKSTNLM 451
Score = 74.3 bits (181), Expect = 3e-13
Identities = 71/240 (29%), Positives = 112/240 (46%), Gaps = 16/240 (6%)
Query: 61 VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKF 119
++++ LD+ + GE ++ L + N G + I SLK L LS N+
Sbjct: 426 LMALDLDSNNFTGEIP-KSLWKSTNLMEFTASYNRLEGYLPAEIGNAASLKRLVLSDNQL 484
Query: 120 NGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMG 179
G +P +L SL LNL+ N F G +P L LD SN+ G I + +
Sbjct: 485 TGEIPREIGKLTSLSVLNLNANMFQGKIPVELGDCTSLTTLDLGSNNLQGQIPDKITALA 544
Query: 180 SVLHVDLSNNKFSGAL---------DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPY 230
+ + LS N SG++ + + D+SFL ++S+N L G + G
Sbjct: 545 QLQCLVLSYNNLSGSIPSKPSAYFHQIEMPDLSFLQHHGIFDLSYNRLSGPIPEELGECL 604
Query: 231 LDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+ L SNN L G IP S + + +L IL L+ N LTGS+P+ + +S+ L L+L+
Sbjct: 605 V--LVEISLSNNHLSGEIPASLSRLTNLTILDLSGNALTGSIPKEM--GNSLKLQGLNLA 660
Score = 58.2 bits (139), Expect = 2e-08
Identities = 56/204 (27%), Positives = 99/204 (48%), Gaps = 19/204 (9%)
Query: 86 LHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLP--PSF----VELRSLVYL-- 136
L L + +N+ G + I+ + L+ L LS N +GS+P PS +E+ L +L
Sbjct: 522 LTTLDLGSNNLQGQIPDKITALAQLQCLVLSYNNLSGSIPSKPSAYFHQIEMPDLSFLQH 581
Query: 137 ----NLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFS 192
+LS N SG +P + L + +N SG+I ++ ++ +DLS N +
Sbjct: 582 HGIFDLSYNRLSGPIPEELGECLVLVEISLSNNHLSGEIPASLSRLTNLTILDLSGNALT 641
Query: 193 GALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SF 251
G++ +G+ +Q LN+++N L G + G+ L +L + + N+L G +P S
Sbjct: 642 GSIPKEMGN---SLKLQGLNLANNQLNGHIPESFGL--LGSLVKLNLTKNKLDGPVPASL 696
Query: 252 TFVVSLRILRLACNQLTGSLPETL 275
+ L + L+ N L+G L L
Sbjct: 697 GNLKELTHMDLSFNNLSGELSSEL 720
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 92.0 bits (227), Expect = 1e-18
Identities = 68/205 (33%), Positives = 104/205 (50%), Gaps = 9/205 (4%)
Query: 79 AISNLPMLHNLSVVNNHFTG---SMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVY 135
+ SNLP L L + +N+ TG S + PM +LK L L N F G +P S LV
Sbjct: 396 SFSNLPKLETLDMSSNNLTGIIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVS 455
Query: 136 LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGAL 195
L+LS N +G++P+ L +L+ L N SG+I + + ++ ++ L N +G +
Sbjct: 456 LDLSFNYLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPI 515
Query: 196 DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS-FTFV 254
L + + + +++S+N L GE+ A G L NL + NN + GNIP+
Sbjct: 516 PASLSNCT---KLNWISLSNNQLSGEIPASLGR--LSNLAILKLGNNSISGNIPAELGNC 570
Query: 255 VSLRILRLACNQLTGSLPETLLKES 279
SL L L N L GS+P L K+S
Sbjct: 571 QSLIWLDLNTNFLNGSIPPPLFKQS 595
Score = 87.4 bits (215), Expect = 4e-17
Identities = 72/220 (32%), Positives = 110/220 (49%), Gaps = 11/220 (5%)
Query: 71 LVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVEL 130
L G L NL L +LS N+F+ +L+ LDLS NKF G + S
Sbjct: 224 LAGSIPELDFKNLSYL-DLSA--NNFSTVFPSFKDCSNLQHLDLSSNKFYGDIGSSLSSC 280
Query: 131 RSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM-GSVLHVDLSNN 189
L +LNL+ N+F G VP + + L+YL N F G + +V+ +DLS N
Sbjct: 281 GKLSFLNLTNNQFVGLVPKL--PSESLQYLYLRGNDFQGVYPNQLADLCKTVVELDLSYN 338
Query: 190 KFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP 249
FSG + LG+ S S++ +++S+N+ G+L D + L N++ S N+ VG +P
Sbjct: 339 NFSGMVPESLGECS---SLELVDISNNNFSGKL-PVDTLLKLSNIKTMVLSFNKFVGGLP 394
Query: 250 -SFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDL 288
SF+ + L L ++ N LTG +P + K+ L L L
Sbjct: 395 DSFSNLPKLETLDMSSNNLTGIIPSGICKDPMNNLKVLYL 434
Score = 68.6 bits (166), Expect = 2e-11
Identities = 67/237 (28%), Positives = 108/237 (45%), Gaps = 36/237 (15%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVV---NNHFTGSM-LHISPMKSLKFLDLSLNKF 119
I+L N L GE ++L L NL+++ NN +G++ + +SL +LDL+ N
Sbjct: 528 ISLSNNQLSGEIP----ASLGRLSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFL 583
Query: 120 NGSLPPSFVELRSLVYLNL---------------------SLNEFSGTVPNVFHKLDQLE 158
NGS+PP + + + L +L EF G ++
Sbjct: 584 NGSIPPPLFKQSGNIAVALLTGKRYVYIKNDGSKECHGAGNLLEFGGIRQEQLDRISTRH 643
Query: 159 YLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSL 218
+F + + G F GS++ +DLS NK G++ LG + +L LN+ HN L
Sbjct: 644 PCNF-TRVYRGITQPTFNHNGSMIFLDLSYNKLEGSIPKELGAMYYL---SILNLGHNDL 699
Query: 219 VGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPET 274
G + G L N+ + D S N+ G IP S T + L + L+ N L+G +PE+
Sbjct: 700 SGMIPQQLG--GLKNVAILDLSYNRFNGTIPNSLTSLTLLGEIDLSNNNLSGMIPES 754
Score = 58.2 bits (139), Expect = 2e-08
Identities = 58/216 (26%), Positives = 91/216 (41%), Gaps = 29/216 (13%)
Query: 83 LPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLN 141
L L NL + N TG + +S L ++ LS N+ +G +P S L +L L L N
Sbjct: 498 LQALENLILDFNDLTGPIPASLSNCTKLNWISLSNNQLSGEIPASLGRLSNLAILKLGNN 557
Query: 142 EFSGTVPNVFHKLDQLEYLDFHSNSFSGDIME-IFYQMGSVL--------HVDLSNN--- 189
SG +P L +LD ++N +G I +F Q G++ +V + N+
Sbjct: 558 SISGNIPAELGNCQSLIWLDLNTNFLNGSIPPPLFKQSGNIAVALLTGKRYVYIKNDGSK 617
Query: 190 ---------KFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDAS 240
+F G L +S V + + F H+G ++ D S
Sbjct: 618 ECHGAGNLLEFGGIRQEQLDRISTRHPCNFTRV-YRGITQPTFNHNG-----SMIFLDLS 671
Query: 241 NNQLVGNIPS-FTFVVSLRILRLACNQLTGSLPETL 275
N+L G+IP + L IL L N L+G +P+ L
Sbjct: 672 YNKLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQL 707
Score = 52.0 bits (123), Expect = 2e-06
Identities = 36/128 (28%), Positives = 65/128 (50%), Gaps = 6/128 (4%)
Query: 119 FNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM 178
+ G P+F S+++L+LS N+ G++P + L L+ N SG I + +
Sbjct: 651 YRGITQPTFNHNGSMIFLDLSYNKLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQLGGL 710
Query: 179 GSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFD 238
+V +DLS N+F+G + L ++ L I ++S+N+L G + + P+ D +
Sbjct: 711 KNVAILDLSYNRFNGTIPNSLTSLTLLGEI---DLSNNNLSGMI--PESAPF-DTFPDYR 764
Query: 239 ASNNQLVG 246
+NN L G
Sbjct: 765 FANNSLCG 772
Score = 48.9 bits (115), Expect = 1e-05
Identities = 40/131 (30%), Positives = 58/131 (43%), Gaps = 10/131 (7%)
Query: 53 GILCSEGNVISITLDNASLVGEFNFL----AISNLPMLHNLSVV-----NNHFTGSM-LH 102
G L G + LD S NF I+ HN S++ N GS+
Sbjct: 623 GNLLEFGGIRQEQLDRISTRHPCNFTRVYRGITQPTFNHNGSMIFLDLSYNKLEGSIPKE 682
Query: 103 ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDF 162
+ M L L+L N +G +P L+++ L+LS N F+GT+PN L L +D
Sbjct: 683 LGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTSLTLLGEIDL 742
Query: 163 HSNSFSGDIME 173
+N+ SG I E
Sbjct: 743 SNNNLSGMIPE 753
Score = 42.0 bits (97), Expect = 0.002
Identities = 68/282 (24%), Positives = 108/282 (38%), Gaps = 63/282 (22%)
Query: 13 RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASLV 72
+D LL K + P +L +W S + P ++ G+ C V SI L N L
Sbjct: 42 KDSQQLLSFKAALPPTP--TLLQNWLSST-----DPCSFTGVSCKNSRVSSIDLSNTFLS 94
Query: 73 GEFNFLAISNLPM--LHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVEL 130
+F+ + LP+ L +L + N + +GS+ + + LD
Sbjct: 95 VDFSLVTSYLLPLSNLESLVLKNANLSGSLTSAAKSQCGVTLDS---------------- 138
Query: 131 RSLVYLNLSLNEFSGTVPNV--FHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSN 188
++L+ N SG + ++ F L+ L+ N E+
Sbjct: 139 -----IDLAENTISGPISDISSFGVCSNLKSLNLSKNFLDPPGKEML------------- 180
Query: 189 NKFSGALDLGLGDVSFLFSIQHLNVSHNSLVG-ELFAHDGMPYLDNLEVFDASNNQLVGN 247
GA FS+Q L++S+N++ G LF LE F N+L G+
Sbjct: 181 ---KGAT----------FSLQVLDLSYNNISGFNLFPWVSSMGFVELEFFSIKGNKLAGS 227
Query: 248 IPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
IP F +L L L+ N + P K+ S L LDLS
Sbjct: 228 IPELDF-KNLSYLDLSANNFSTVFPS--FKDCS-NLQHLDLS 265
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor
(EC 2.7.1.37) (tBRI1) (Altered brassinolide sensitivity
1) (Systemin receptor SR160)
Length = 1207
Score = 90.5 bits (223), Expect = 4e-18
Identities = 73/226 (32%), Positives = 113/226 (49%), Gaps = 11/226 (4%)
Query: 65 TLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLP 124
+L L G L NL L +LS N+F+ +L+ LDLS NKF G +
Sbjct: 218 SLKGNKLAGSIPELDFKNLSYL-DLSA--NNFSTVFPSFKDCSNLQHLDLSSNKFYGDIG 274
Query: 125 PSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM-GSVLH 183
S L +LNL+ N+F G VP + + L+YL N F G + +V+
Sbjct: 275 SSLSSCGKLSFLNLTNNQFVGLVPKL--PSESLQYLYLRGNDFQGVYPNQLADLCKTVVE 332
Query: 184 VDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQ 243
+DLS N FSG + LG+ S S++ +++S+N+ G+L D + L N++ S N+
Sbjct: 333 LDLSYNNFSGMVPESLGECS---SLELVDISYNNFSGKL-PVDTLSKLSNIKTMVLSFNK 388
Query: 244 LVGNIP-SFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDL 288
VG +P SF+ ++ L L ++ N LTG +P + K+ L L L
Sbjct: 389 FVGGLPDSFSNLLKLETLDMSSNNLTGVIPSGICKDPMNNLKVLYL 434
Score = 88.2 bits (217), Expect = 2e-17
Identities = 67/205 (32%), Positives = 103/205 (49%), Gaps = 9/205 (4%)
Query: 79 AISNLPMLHNLSVVNNHFTG---SMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVY 135
+ SNL L L + +N+ TG S + PM +LK L L N F G +P S LV
Sbjct: 396 SFSNLLKLETLDMSSNNLTGVIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVS 455
Query: 136 LNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGAL 195
L+LS N +G++P+ L +L+ L N SG+I + + ++ ++ L N +G +
Sbjct: 456 LDLSFNYLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPI 515
Query: 196 DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS-FTFV 254
L + + + +++S+N L GE+ A G L NL + NN + GNIP+
Sbjct: 516 PASLSNCT---KLNWISLSNNQLSGEIPASLGR--LSNLAILKLGNNSISGNIPAELGNC 570
Query: 255 VSLRILRLACNQLTGSLPETLLKES 279
SL L L N L GS+P L K+S
Sbjct: 571 QSLIWLDLNTNFLNGSIPPPLFKQS 595
Score = 58.9 bits (141), Expect = 1e-08
Identities = 33/88 (37%), Positives = 44/88 (49%)
Query: 108 SLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSF 167
S+ FLDLS NK GS+P + L LNL N+ SG +P L + LD N F
Sbjct: 664 SMIFLDLSYNKLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRF 723
Query: 168 SGDIMEIFYQMGSVLHVDLSNNKFSGAL 195
+G I + + +DLSNN SG +
Sbjct: 724 NGTIPNSLTSLTLLGEIDLSNNNLSGMI 751
Score = 58.2 bits (139), Expect = 2e-08
Identities = 58/216 (26%), Positives = 91/216 (41%), Gaps = 29/216 (13%)
Query: 83 LPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLN 141
L L NL + N TG + +S L ++ LS N+ +G +P S L +L L L N
Sbjct: 498 LQALENLILDFNDLTGPIPASLSNCTKLNWISLSNNQLSGEIPASLGRLSNLAILKLGNN 557
Query: 142 EFSGTVPNVFHKLDQLEYLDFHSNSFSGDIME-IFYQMGSVL--------HVDLSNN--- 189
SG +P L +LD ++N +G I +F Q G++ +V + N+
Sbjct: 558 SISGNIPAELGNCQSLIWLDLNTNFLNGSIPPPLFKQSGNIAVALLTGKRYVYIKNDGSK 617
Query: 190 ---------KFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDAS 240
+F G L +S V + + F H+G ++ D S
Sbjct: 618 ECHGAGNLLEFGGIRQEQLDRISTRHPCNFTRV-YRGITQPTFNHNG-----SMIFLDLS 671
Query: 241 NNQLVGNIPS-FTFVVSLRILRLACNQLTGSLPETL 275
N+L G+IP + L IL L N L+G +P+ L
Sbjct: 672 YNKLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQL 707
Score = 53.1 bits (126), Expect = 8e-07
Identities = 65/253 (25%), Positives = 99/253 (38%), Gaps = 55/253 (21%)
Query: 64 ITLDNASLVGEFNFLAISNLPMLHNLSVV---NNHFTGSM-LHISPMKSLKFLDLSLNKF 119
I+L N L GE ++L L NL+++ NN +G++ + +SL +LDL+ N
Sbjct: 528 ISLSNNQLSGEIP----ASLGRLSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFL 583
Query: 120 NGSLPPSFVELRSLVYLNL---------------------SLNEFSGTVPNVFHKLDQLE 158
NGS+PP + + + L +L EF G ++
Sbjct: 584 NGSIPPPLFKQSGNIAVALLTGKRYVYIKNDGSKECHGAGNLLEFGGIRQEQLDRISTRH 643
Query: 159 YLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSL 218
+F + + G F GS++ +DLS NK G++ LG + +L LN+ HN L
Sbjct: 644 PCNF-TRVYRGITQPTFNHNGSMIFLDLSYNKLEGSIPKELGAMYYL---SILNLGHNDL 699
Query: 219 VGELFAHDG----------------------MPYLDNLEVFDASNNQLVGNIPSFTFVVS 256
G + G + L L D SNN L G IP +
Sbjct: 700 SGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTSLTLLGEIDLSNNNLSGMIPESAPFDT 759
Query: 257 LRILRLACNQLTG 269
R A N L G
Sbjct: 760 FPDYRFANNSLCG 772
Score = 48.9 bits (115), Expect = 1e-05
Identities = 40/131 (30%), Positives = 58/131 (43%), Gaps = 10/131 (7%)
Query: 53 GILCSEGNVISITLDNASLVGEFNFL----AISNLPMLHNLSVV-----NNHFTGSM-LH 102
G L G + LD S NF I+ HN S++ N GS+
Sbjct: 623 GNLLEFGGIRQEQLDRISTRHPCNFTRVYRGITQPTFNHNGSMIFLDLSYNKLEGSIPKE 682
Query: 103 ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDF 162
+ M L L+L N +G +P L+++ L+LS N F+GT+PN L L +D
Sbjct: 683 LGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTSLTLLGEIDL 742
Query: 163 HSNSFSGDIME 173
+N+ SG I E
Sbjct: 743 SNNNLSGMIPE 753
Score = 43.1 bits (100), Expect = 8e-04
Identities = 68/282 (24%), Positives = 108/282 (38%), Gaps = 63/282 (22%)
Query: 13 RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASLV 72
+D LL K + P +L +W L S G P ++ G+ C V SI L N L
Sbjct: 42 KDSQQLLSFKAALPPTP--TLLQNW----LSSTG-PCSFTGVSCKNSRVSSIDLSNTFLS 94
Query: 73 GEFNFLAISNLPM--LHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVEL 130
+F+ + LP+ L +L + N + +GS+ + + LD
Sbjct: 95 VDFSLVTSYLLPLSNLESLVLKNANLSGSLTSAAKSQCGVTLDS---------------- 138
Query: 131 RSLVYLNLSLNEFSGTVPNV--FHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSN 188
++L+ N SG + ++ F L+ L+ N E+
Sbjct: 139 -----IDLAENTISGPISDISSFGVCSNLKSLNLSKNFLDPPGKEMLK------------ 181
Query: 189 NKFSGALDLGLGDVSFLFSIQHLNVSHNSLVG-ELFAHDGMPYLDNLEVFDASNNQLVGN 247
+ FS+Q L++S+N++ G LF LE F N+L G+
Sbjct: 182 --------------AATFSLQVLDLSYNNISGFNLFPWVSSMGFVELEFFSLKGNKLAGS 227
Query: 248 IPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
IP F +L L L+ N + P K+ S L LDLS
Sbjct: 228 IPELDF-KNLSYLDLSANNFSTVFPS--FKDCS-NLQHLDLS 265
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 90.5 bits (223), Expect = 4e-18
Identities = 95/307 (30%), Positives = 142/307 (45%), Gaps = 46/307 (14%)
Query: 13 RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNVISITLDNASLV 72
R+I L+ K + P +L W S N P + G+ C + V SI L + L
Sbjct: 34 REIHQLISFKDVL---PDKNLLPDWSS-----NKNPCTFDGVTCRDDKVTSIDLSSKPLN 85
Query: 73 GEFNFLAISNLPM--LHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLP--PSFV 128
F+ ++ S L + L +L + N+H GS+ SL LDLS N +G + S
Sbjct: 86 VGFSAVSSSLLSLTGLESLFLSNSHINGSVSGFKCSASLTSLDLSRNSLSGPVTTLTSLG 145
Query: 129 ELRSLVYLNLSLN--EFSGTVPNVFHKLDQLEYLDFHSNSFSGDIM---EIFYQMGSVLH 183
L +LN+S N +F G V KL+ LE LD +NS SG + + G + H
Sbjct: 146 SCSGLKFLNVSSNTLDFPGKVSGGL-KLNSLEVLDLSANSISGANVVGWVLSDGCGELKH 204
Query: 184 VDLSNNKFSGALDLG---------------------LGDVSFLFSIQHLNVSHNSLVGEL 222
+ +S NK SG +D+ LGD S ++QHL++S N L G+
Sbjct: 205 LAISGNKISGDVDVSRCVNLEFLDVSSNNFSTGIPFLGDCS---ALQHLDISGNKLSGDF 261
Query: 223 FAHDGMPYLDNLEVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPETLLKESSMM 282
+ L++ + S+NQ VG IP + SL+ L LA N+ TG +P+ L +
Sbjct: 262 --SRAISTCTELKLLNISSNQFVGPIPPLP-LKSLQYLSLAENKFTGEIPD-FLSGACDT 317
Query: 283 LSELDLS 289
L+ LDLS
Sbjct: 318 LTGLDLS 324
Score = 83.2 bits (204), Expect = 7e-16
Identities = 76/264 (28%), Positives = 121/264 (45%), Gaps = 15/264 (5%)
Query: 31 GLVLNSWDSKSLESNGCP-QNWYGILCSEG--NVISITLDNASLVGEFNFLAISNLPMLH 87
GL LNS + L +N N G + S+G + + + + G+ + +S L
Sbjct: 169 GLKLNSLEVLDLSANSISGANVVGWVLSDGCGELKHLAISGNKISGDVD---VSRCVNLE 225
Query: 88 NLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTV 147
L V +N+F+ + + +L+ LD+S NK +G + L LN+S N+F G +
Sbjct: 226 FLDVSSNNFSTGIPFLGDCSALQHLDISGNKLSGDFSRAISTCTELKLLNISSNQFVGPI 285
Query: 148 PNVFHKLDQLEYLDFHSNSFSGDIMEIFY-QMGSVLHVDLSNNKFSGALDLGLGDVSFLF 206
P + L L+YL N F+G+I + ++ +DLS N F GA+ G S L
Sbjct: 286 PPL--PLKSLQYLSLAENKFTGEIPDFLSGACDTLTGLDLSGNHFYGAVPPFFGSCSLL- 342
Query: 207 SIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS--FTFVVSLRILRLAC 264
+ L +S N+ GEL D + + L+V D S N+ G +P SL L L+
Sbjct: 343 --ESLALSSNNFSGEL-PMDTLLKMRGLKVLDLSFNEFSGELPESLTNLSASLLTLDLSS 399
Query: 265 NQLTGSLPETLLKESSMMLSELDL 288
N +G + L + L EL L
Sbjct: 400 NNFSGPILPNLCQNPKNTLQELYL 423
Score = 81.6 bits (200), Expect = 2e-15
Identities = 71/259 (27%), Positives = 116/259 (44%), Gaps = 54/259 (20%)
Query: 63 SITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNG 121
++ LD L GE +SN L+ +S+ NN TG + I +++L L LS N F+G
Sbjct: 492 TLILDFNDLTGEIPS-GLSNCTNLNWISLSNNRLTGEIPKWIGRLENLAILKLSNNSFSG 550
Query: 122 SLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKL--------------------------- 154
++P + RSL++L+L+ N F+GT+P K
Sbjct: 551 NIPAELGDCRSLIWLDLNTNLFNGTIPAAMFKQSGKIAANFIAGKRYVYIKNDGMKKECH 610
Query: 155 -------------DQLEYL------DFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGAL 195
+QL L + S + G F GS++ +D+S N SG +
Sbjct: 611 GAGNLLEFQGIRSEQLNRLSTRNPCNITSRVYGGHTSPTFDNNGSMMFLDMSYNMLSGYI 670
Query: 196 DLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFV 254
+G + +LF LN+ HN + G + D + L L + D S+N+L G IP + + +
Sbjct: 671 PKEIGSMPYLFI---LNLGHNDISGSI--PDEVGDLRGLNILDLSSNKLDGRIPQAMSAL 725
Query: 255 VSLRILRLACNQLTGSLPE 273
L + L+ N L+G +PE
Sbjct: 726 TMLTEIDLSNNNLSGPIPE 744
Score = 81.3 bits (199), Expect = 3e-15
Identities = 61/198 (30%), Positives = 99/198 (49%), Gaps = 9/198 (4%)
Query: 86 LHNLSVVNNHFTGSMLHI---SPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNE 142
L L + +N+F+G +L +P +L+ L L N F G +PP+ LV L+LS N
Sbjct: 392 LLTLDLSSNNFSGPILPNLCQNPKNTLQELYLQNNGFTGKIPPTLSNCSELVSLHLSFNY 451
Query: 143 FSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDV 202
SGT+P+ L +L L N G+I + + ++ + L N +G + GL +
Sbjct: 452 LSGTIPSSLGSLSKLRDLKLWLNMLEGEIPQELMYVKTLETLILDFNDLTGEIPSGLSNC 511
Query: 203 SFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPS-FTFVVSLRILR 261
+ ++ +++S+N L GE+ G L+NL + SNN GNIP+ SL L
Sbjct: 512 T---NLNWISLSNNRLTGEIPKWIGR--LENLAILKLSNNSFSGNIPAELGDCRSLIWLD 566
Query: 262 LACNQLTGSLPETLLKES 279
L N G++P + K+S
Sbjct: 567 LNTNLFNGTIPAAMFKQS 584
Score = 60.1 bits (144), Expect = 6e-09
Identities = 68/293 (23%), Positives = 121/293 (41%), Gaps = 75/293 (25%)
Query: 70 SLVGEFNFLA------ISNLPMLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGS 122
SL FN+L+ + +L L +L + N G + + +K+L+ L L N G
Sbjct: 444 SLHLSFNYLSGTIPSSLGSLSKLRDLKLWLNMLEGEIPQELMYVKTLETLILDFNDLTGE 503
Query: 123 LPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVL 182
+P +L +++LS N +G +P +L+ L L +NSFSG+I S++
Sbjct: 504 IPSGLSNCTNLNWISLSNNRLTGEIPKWIGRLENLAILKLSNNSFSGNIPAELGDCRSLI 563
Query: 183 HVDLSNNKFSGALDLGL----GDV--SFLFSIQHLNVSHNSL------VGELFAHDGM-- 228
+DL+ N F+G + + G + +F+ +++ + ++ + G L G+
Sbjct: 564 WLDLNTNLFNGTIPAAMFKQSGKIAANFIAGKRYVYIKNDGMKKECHGAGNLLEFQGIRS 623
Query: 229 ------------------------PYLDN---LEVFDASNNQLVGNIPS------FTFVV 255
P DN + D S N L G IP + F++
Sbjct: 624 EQLNRLSTRNPCNITSRVYGGHTSPTFDNNGSMMFLDMSYNMLSGYIPKEIGSMPYLFIL 683
Query: 256 S-------------------LRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+ L IL L+ N+L G +P+ + + ML+E+DLS
Sbjct: 684 NLGHNDISGSIPDEVGDLRGLNILDLSSNKLDGRIPQAM--SALTMLTEIDLS 734
Score = 47.0 bits (110), Expect = 5e-05
Identities = 26/72 (36%), Positives = 37/72 (51%)
Query: 103 ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDF 162
I M L L+L N +GS+P +LR L L+LS N+ G +P L L +D
Sbjct: 674 IGSMPYLFILNLGHNDISGSIPDEVGDLRGLNILDLSSNKLDGRIPQAMSALTMLTEIDL 733
Query: 163 HSNSFSGDIMEI 174
+N+ SG I E+
Sbjct: 734 SNNNLSGPIPEM 745
>PGI1_ARATH (Q9M5J9) Polygalacturonase inhibitor 1 precursor
(Polygalacturonase-inhibiting protein) (PGIP-1)
Length = 330
Score = 90.1 bits (222), Expect = 6e-18
Identities = 76/287 (26%), Positives = 140/287 (48%), Gaps = 41/287 (14%)
Query: 14 DIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSEGNV----ISITLDNA 69
D + LL++KK + N+P+ L SWD +++ C +WY + C + V ++T+ +
Sbjct: 29 DKNTLLKIKKSL-NNPYHLA--SWDP---QTDCC--SWYCLECGDATVNHRVTALTIFSG 80
Query: 70 SLVGEFNFLAISNLPMLHNLSVVN-NHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSF 127
+ G+ + +LP L L ++ TG++ I+ +K+L+ L LS G +P
Sbjct: 81 QISGQIP-AEVGDLPYLETLVFRKLSNLTGTIQPTIAKLKNLRMLRLSWTNLTGPIPDFI 139
Query: 128 VELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQM-GSVLHVDL 186
+L++L +L LS N+ SG++P+ L ++ L+ N +G I E F G+V + L
Sbjct: 140 SQLKNLEFLELSFNDLSGSIPSSLSTLPKILALELSRNKLTGSIPESFGSFPGTVPDLRL 199
Query: 187 SNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGE---LFAHDGMPYLDNLE-------- 235
S+N+ SG + LG++ F +++S N L G+ LF + + +L
Sbjct: 200 SHNQLSGPIPKSLGNIDF----NRIDLSRNKLQGDASMLFGSNKTTWSIDLSRNMFQFDI 255
Query: 236 ----------VFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLP 272
+ D ++N + GNIP L+ ++ N+L G +P
Sbjct: 256 SKVDIPKTLGILDLNHNGITGNIPVQWTEAPLQFFNVSYNKLCGHIP 302
Score = 40.4 bits (93), Expect = 0.005
Identities = 35/111 (31%), Positives = 60/111 (53%), Gaps = 5/111 (4%)
Query: 167 FSGDIM-EIFYQMGSVLHVD-LSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFA 224
FSG I +I ++G + +++ L K S ++ L +++ L +S +L G +
Sbjct: 78 FSGQISGQIPAEVGDLPYLETLVFRKLSNLTGTIQPTIAKLKNLRMLRLSWTNLTGPI-- 135
Query: 225 HDGMPYLDNLEVFDASNNQLVGNIPS-FTFVVSLRILRLACNQLTGSLPET 274
D + L NLE + S N L G+IPS + + + L L+ N+LTGS+PE+
Sbjct: 136 PDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILALELSRNKLTGSIPES 186
Score = 38.1 bits (87), Expect = 0.025
Identities = 28/95 (29%), Positives = 48/95 (50%), Gaps = 4/95 (4%)
Query: 197 LGLGDVSFLFSIQHLNVSHNSLVGELFAHDG-MPYLDNLEVFDASNNQLVGNI-PSFTFV 254
L GD + + L + + G++ A G +PYL+ L SN L G I P+ +
Sbjct: 61 LECGDATVNHRVTALTIFSGQISGQIPAEVGDLPYLETLVFRKLSN--LTGTIQPTIAKL 118
Query: 255 VSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
+LR+LRL+ LTG +P+ + + ++ EL +
Sbjct: 119 KNLRMLRLSWTNLTGPIPDFISQLKNLEFLELSFN 153
Score = 35.8 bits (81), Expect = 0.13
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 5/107 (4%)
Query: 181 VLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDAS 240
V + + + + SG + +GD+ +L ++ +S+ L G + + L NL + S
Sbjct: 72 VTALTIFSGQISGQIPAEVGDLPYLETLVFRKLSN--LTGTI--QPTIAKLKNLRMLRLS 127
Query: 241 NNQLVGNIPSF-TFVVSLRILRLACNQLTGSLPETLLKESSMMLSEL 286
L G IP F + + +L L L+ N L+GS+P +L ++ EL
Sbjct: 128 WTNLTGPIPDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILALEL 174
>PGIP_PYRCO (Q05091) Polygalacturonase inhibitor precursor
(Polygalacturonase-inhibiting protein)
Length = 330
Score = 89.7 bits (221), Expect = 7e-18
Identities = 78/264 (29%), Positives = 128/264 (47%), Gaps = 20/264 (7%)
Query: 14 DIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCSE--GNVISITLDNASL 71
D LL++KK DP+ VL SW S +++ C +WY + C + S+T+ +
Sbjct: 31 DKKVLLQIKKAF-GDPY--VLASWKS---DTDCC--DWYCVTCDSTTNRINSLTIFAGQV 82
Query: 72 VGEFNFLAISNLPMLHNLSVVNN-HFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVE 129
G+ L + +LP L L + TG + I+ +K LK L LS +GS+P +
Sbjct: 83 SGQIPAL-VGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKSLRLSWTNLSGSVPDFLSQ 141
Query: 130 LRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQ-MGSVLHVDLSN 188
L++L +L+LS N +G +P+ +L L L N +G I F Q +G+V + LS+
Sbjct: 142 LKNLTFLDLSFNNLTGAIPSSLSELPNLGALRLDRNKLTGHIPISFGQFIGNVPDLYLSH 201
Query: 189 NKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNI 248
N+ SG + + F +++S N L G+ G+ ++ D S N L N+
Sbjct: 202 NQLSGNIPTSFAQMDF----TSIDLSRNKLEGDASVIFGLN--KTTQIVDLSRNLLEFNL 255
Query: 249 PSFTFVVSLRILRLACNQLTGSLP 272
F SL L + N++ GS+P
Sbjct: 256 SKVEFPTSLTSLDINHNKIYGSIP 279
Score = 32.7 bits (73), Expect = 1.1
Identities = 29/84 (34%), Positives = 45/84 (53%), Gaps = 6/84 (7%)
Query: 208 IQHLNVSHNSLVGELFAHDG-MPYLDNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACN 265
I L + + G++ A G +PYL+ LE N L G I P+ + L+ LRL+
Sbjct: 72 INSLTIFAGQVSGQIPALVGDLPYLETLEFHKQPN--LTGPIQPAIAKLKGLKSLRLSWT 129
Query: 266 QLTGSLPETLLKESSMMLSELDLS 289
L+GS+P+ L + + L+ LDLS
Sbjct: 130 NLSGSVPDFLSQLKN--LTFLDLS 151
>D100_ARATH (Q00874) DNA-damage-repair/toleration protein DRT100
precursor
Length = 372
Score = 87.4 bits (215), Expect = 4e-17
Identities = 93/325 (28%), Positives = 143/325 (43%), Gaps = 67/325 (20%)
Query: 13 RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCS--EGNVISITLDNAS 70
+D AL K + G + N+W E+ C + WYGI C G V I+L S
Sbjct: 30 KDQTALNAFKSSLSEPNLG-IFNTWS----ENTDCCKEWYGISCDPDSGRVTDISLRGES 84
Query: 71 ------LVGEFNFLAISNLPMLHNLSVVNN-------HFTGSMLH-ISPMKSLKFLDLSL 116
G +++ S P + +L+ + + TG + I+ + SL+ LDL+
Sbjct: 85 EDAIFQKAGRSGYMSGSIDPAVCDLTALTSLVLADWKGITGEIPPCITSLASLRILDLAG 144
Query: 117 NKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYL-------------DFH 163
NK G +P +L L LNL+ N+ SG +P L +L++L DF
Sbjct: 145 NKITGEIPAEIGKLSKLAVLNLAENQMSGEIPASLTSLIELKHLELTENGITGVIPADFG 204
Query: 164 S-----------NSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLN 212
S N +G I E M + +DLS N G + +G++ L LN
Sbjct: 205 SLKMLSRVLLGRNELTGSIPESISGMERLADLDLSKNHIEGPIPEWMGNMKVL---SLLN 261
Query: 213 VSHNSLV----GELFAHDGMPYLDNLEVFDASNNQLVGNIP----SFTFVVSLRILRLAC 264
+ NSL G L ++ G L+V + S N L G IP S T++VS L L+
Sbjct: 262 LDCNSLTGPIPGSLLSNSG------LDVANLSRNALEGTIPDVFGSKTYLVS---LDLSH 312
Query: 265 NQLTGSLPETLLKESSMMLSELDLS 289
N L+G +P++L S+ + LD+S
Sbjct: 313 NSLSGRIPDSL--SSAKFVGHLDIS 335
Score = 78.2 bits (191), Expect = 2e-14
Identities = 55/172 (31%), Positives = 83/172 (47%), Gaps = 7/172 (4%)
Query: 82 NLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSL 140
+L ML + + N TGS+ IS M+ L LDLS N G +P ++ L LNL
Sbjct: 205 SLKMLSRVLLGRNELTGSIPESISGMERLADLDLSKNHIEGPIPEWMGNMKVLSLLNLDC 264
Query: 141 NEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLG 200
N +G +P L+ + N+ G I ++F ++ +DLS+N SG + L
Sbjct: 265 NSLTGPIPGSLLSNSGLDVANLSRNALEGTIPDVFGSKTYLVSLDLSHNSLSGRIPDSLS 324
Query: 201 DVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSFT 252
F + HL++SHN L G + G P+ D+LE S+NQ + P T
Sbjct: 325 SAKF---VGHLDISHNKLCGRI--PTGFPF-DHLEATSFSDNQCLCGGPLTT 370
Score = 67.4 bits (163), Expect = 4e-11
Identities = 42/121 (34%), Positives = 58/121 (47%), Gaps = 1/121 (0%)
Query: 79 AISNLPMLHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
+IS + L +L + NH G + + MK L L+L N G +P S + L N
Sbjct: 226 SISGMERLADLDLSKNHIEGPIPEWMGNMKVLSLLNLDCNSLTGPIPGSLLSNSGLDVAN 285
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
LS N GT+P+VF L LD NS SG I + V H+D+S+NK G +
Sbjct: 286 LSRNALEGTIPDVFGSKTYLVSLDLSHNSLSGRIPDSLSSAKFVGHLDISHNKLCGRIPT 345
Query: 198 G 198
G
Sbjct: 346 G 346
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 80.1 bits (196), Expect = 6e-15
Identities = 61/213 (28%), Positives = 100/213 (46%), Gaps = 32/213 (15%)
Query: 1 MLLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILC-SEG 59
++L LV GN + DAL LK + DP VL SWD+ + P W+ + C S+
Sbjct: 15 LVLDLVLRVSGNAEGDALSALKNSLA-DP-NKVLQSWDATLVT----PCTWFHVTCNSDN 68
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKF 119
+V + L NA+L G+ ++ + + +L++L+L N
Sbjct: 69 SVTRVDLGNANLSGQL------------------------VMQLGQLPNLQYLELYSNNI 104
Query: 120 NGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMG 179
G++P L LV L+L LN SG +P+ +L +L +L ++NS SG+I +
Sbjct: 105 TGTIPEQLGNLTELVSLDLYLNNLSGPIPSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVL 164
Query: 180 SVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLN 212
++ +DLSNN +G + + G S I N
Sbjct: 165 TLQVLDLSNNPLTGDIPVN-GSFSLFTPISFAN 196
Score = 54.7 bits (130), Expect = 3e-07
Identities = 41/144 (28%), Positives = 69/144 (47%), Gaps = 8/144 (5%)
Query: 132 SLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKF 191
S+ ++L SG + +L L+YL+ +SN+ +G I E + ++ +DL N
Sbjct: 69 SVTRVDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNL 128
Query: 192 SGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-- 249
SG + LG + ++ L +++NSL GE+ + + L+V D SNN L G+IP
Sbjct: 129 SGPIPSTLGRLK---KLRFLRLNNNSLSGEI--PRSLTAVLTLQVLDLSNNPLTGDIPVN 183
Query: 250 -SFTFVVSLRILRLACNQLTGSLP 272
SF+ + L S P
Sbjct: 184 GSFSLFTPISFANTKLTPLPASPP 207
Score = 46.2 bits (108), Expect = 9e-05
Identities = 39/111 (35%), Positives = 56/111 (50%), Gaps = 8/111 (7%)
Query: 180 SVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDA 239
SV VDL N SG L + LG + ++Q+L + N++ G + + + L L D
Sbjct: 69 SVTRVDLGNANLSGQLVMQLGQLP---NLQYLELYSNNITGTI--PEQLGNLTELVSLDL 123
Query: 240 SNNQLVGNIPS-FTFVVSLRILRLACNQLTGSLPETLLKESSMMLSELDLS 289
N L G IPS + LR LRL N L+G +P +L + + L LDLS
Sbjct: 124 YLNNLSGPIPSTLGRLKKLRFLRLNNNSLSGEIPRSL--TAVLTLQVLDLS 172
>TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor
Length = 674
Score = 78.6 bits (192), Expect = 2e-14
Identities = 73/239 (30%), Positives = 109/239 (45%), Gaps = 32/239 (13%)
Query: 11 GNRDIDALL-ELKKGIQNDPFGLVLNSWDS----------KSLESNGCP--------QNW 51
G+ D+ LL ++K +Q + L+L+SW+S K + SNG P W
Sbjct: 29 GSSDVKLLLGKIKSSLQGNSESLLLSSWNSSVPVCQWRGVKWVFSNGSPLQCSDLSSPQW 88
Query: 52 YGILC---SEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSM-LHISPMK 107
S +++S+ L +A+L G I ML ++ + N +GS+ L +
Sbjct: 89 TNTSLFNDSSLHLLSLQLPSANLTGSLP-REIGEFSMLQSVFLNINSLSGSIPLELGYTS 147
Query: 108 SLKFLDLSLNKFNGSLPPSFVEL-RSLVYLNLSLNEFSGTVPNVF---HKLDQLEYLDFH 163
SL +DLS N G LPPS L LV + N SG +P L+ LD
Sbjct: 148 SLSDVDLSGNALAGVLPPSIWNLCDKLVSFKIHGNNLSGVLPEPALPNSTCGNLQVLDLG 207
Query: 164 SNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGEL 222
N FSG+ E + V +DLS+N F G + GLG + ++ LN+SHN+ G L
Sbjct: 208 GNKFSGEFPEFITRFKGVKSLDLSSNVFEGLVPEGLG----VLELESLNLSHNNFSGML 262
Score = 38.9 bits (89), Expect = 0.015
Identities = 31/85 (36%), Positives = 40/85 (46%), Gaps = 2/85 (2%)
Query: 86 LHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFS 144
L L + N F+G I+ K +K LDLS N F G +P L L LNLS N FS
Sbjct: 201 LQVLDLGGNKFSGEFPEFITRFKGVKSLDLSSNVFEGLVPEGLGVLE-LESLNLSHNNFS 259
Query: 145 GTVPNVFHKLDQLEYLDFHSNSFSG 169
G +P+ E + +S S G
Sbjct: 260 GMLPDFGESKFGAESFEGNSPSLCG 284
>PGI2_PHAVU (P58822) Polygalacturonase inhibitor 2 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2)
Length = 342
Score = 74.3 bits (181), Expect = 3e-13
Identities = 86/335 (25%), Positives = 136/335 (39%), Gaps = 102/335 (30%)
Query: 1 MLLLLVNTAFGN----RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILC 56
++L+ ++TA +D ALL++KK + N L+SW + + C + W G+LC
Sbjct: 19 VILVSLSTAHSELCNPQDKQALLQIKKDLGNPT---TLSSWLPTT---DCCNRTWLGVLC 72
Query: 57 SEGNVISITLDNASLVG-----------------EFNFL--------------AISNLPM 85
+ + + ++N L G NFL AI+ L
Sbjct: 73 -DTDTQTYRVNNLDLSGLNLPKPYPIPSSLANLPYLNFLYIGGINNLVGPIPPAIAKLTQ 131
Query: 86 LHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLNEFS 144
LH L + + + +G++ +S +K+L LD S N +G+LPPS L +LV + N S
Sbjct: 132 LHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSLPNLVGITFDGNRIS 191
Query: 145 GTVPNVFHKLDQL------------------------EYLDFHSNSFSGDIMEIF----- 175
G +P+ + +L ++D N GD +F
Sbjct: 192 GAIPDSYGSFSKLFTSMTISRNRLTGKIPPTFANLNLAFVDLSRNMLEGDASVLFGSDKN 251
Query: 176 ------------YQMGSV------LHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNS 217
+ +G V +DL NN+ G L GL + FL S LNVS N+
Sbjct: 252 TQKIHLAKNSLAFDLGKVGLSKNLNGLDLRNNRIYGTLPQGLTQLKFLHS---LNVSFNN 308
Query: 218 LVGELFAHDGMPYLDNLEVFDAS---NNQLVGNIP 249
L GE+ P NL+ FD S NN+ + P
Sbjct: 309 LCGEI------PQGGNLQRFDVSAYANNKCLCGSP 337
>PGI1_PHAVU (P35334) Polygalacturonase inhibitor 1 precursor
(Polygalacturonase-inhibiting protein) (PGIP-1)
Length = 342
Score = 73.6 bits (179), Expect = 5e-13
Identities = 90/341 (26%), Positives = 140/341 (40%), Gaps = 106/341 (31%)
Query: 1 MLLLLVNTAFGN----RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILC 56
++L+ + TA +D ALL++KK + N L+SW + + C + W G+LC
Sbjct: 19 VILVSLRTALSELCNPQDKQALLQIKKDLGNPT---TLSSWLPTT---DCCNRTWLGVLC 72
Query: 57 SEGNVISITLDNASLVGE-----------------FNFL--------------AISNLPM 85
+ + + ++N L G NFL AI+ L
Sbjct: 73 -DTDTQTYRVNNLDLSGHNLPKPYPIPSSLANLPYLNFLYIGGINNLVGPIPPAIAKLTQ 131
Query: 86 LHNLSVVNNHFTGSMLH-ISPMKSLKFLDLSLNKFNGSLPPSFVELRSL----------- 133
LH L + + + +G++ +S +K+L LD S N +G+LPPS L +L
Sbjct: 132 LHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSLPNLGGITFDGNRIS 191
Query: 134 --------------VYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIF---- 175
+ +S N +G +P F L+ L ++D N GD +F
Sbjct: 192 GAIPDSYGSFSKLFTAMTISRNRLTGKIPPTFANLN-LAFVDLSRNMLEGDASVLFGSDK 250
Query: 176 -------------YQMGSV------LHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHN 216
+ +G V +DL NN+ G L GL + FL Q LNVS N
Sbjct: 251 NTKKIHLAKNSLAFDLGKVGLSKNLNGLDLRNNRIYGTLPQGLTQLKFL---QSLNVSFN 307
Query: 217 SLVGELFAHDGMPYLDNLEVFD----ASNNQLVGN-IPSFT 252
+L GE+ P NL+ FD A+N L G+ +PS T
Sbjct: 308 NLCGEI------PQGGNLKRFDVSSYANNKCLCGSPLPSCT 342
>PGI3_PHAVU (P58823) Polygalacturonase inhibitor 3 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2) (PGIP-3)
Length = 342
Score = 73.2 bits (178), Expect = 7e-13
Identities = 74/323 (22%), Positives = 134/323 (40%), Gaps = 76/323 (23%)
Query: 1 MLLLLVNTAFGN----RDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILC 56
++L+ + TA +D ALL++KK + N L+SW + + C + W G+LC
Sbjct: 19 VILVSLRTALSELCNPQDKQALLQIKKDLGNPT---TLSSWLPTT---DCCNRTWLGVLC 72
Query: 57 SEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLS- 115
+ + + ++N L G NLP + + ++ + L FL +
Sbjct: 73 -DTDTQTYRVNNLDLSGH-------NLPKPYPIPS----------SLANLPYLNFLYIGG 114
Query: 116 LNKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIF 175
+N G +PP+ +L L YL ++ SG +P+ ++ L LDF N+ SG +
Sbjct: 115 INNLVGPIPPAIAKLTQLHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSI 174
Query: 176 YQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGEL---FAHDGMPYLD 232
+ +++ + N+ SGA+ G S LF+ + +S N L G++ FA+ + ++D
Sbjct: 175 SSLPNLVGITFDGNRISGAIPDSYGSFSKLFT--SMTISRNRLTGKIPPTFANLNLAFVD 232
Query: 233 -----------------------------------------NLEVFDASNNQLVGNIP-S 250
NL D NN++ G +P
Sbjct: 233 LSRNMLQGDASVLFGSDKNTQKIHLAKNSLDFDLEKVGLSKNLNGLDLRNNRIYGTLPQG 292
Query: 251 FTFVVSLRILRLACNQLTGSLPE 273
T + L L ++ N L G +P+
Sbjct: 293 LTQLKFLHSLNVSFNNLCGEIPQ 315
>TMM_ARATH (Q9SSD1) TOO MANY MOUTHS protein precursor (TMM)
Length = 496
Score = 72.4 bits (176), Expect = 1e-12
Identities = 57/196 (29%), Positives = 88/196 (44%), Gaps = 29/196 (14%)
Query: 78 LAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLN 137
L+ + L +L + N TGS+ + +L LDL+ N G +PP+ SL+ ++
Sbjct: 201 LSFNRFSGLRSLDLSGNRLTGSIPGFV-LPALSVLDLNQNLLTGPVPPTLTSCGSLIKID 259
Query: 138 LSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFYQMGSVLHVDLSNNKFSGALDL 197
LS N +G +P ++L+QL L DLS N+ SG
Sbjct: 260 LSRNRVTGPIPESINRLNQLVLL------------------------DLSYNRLSGPFPS 295
Query: 198 GLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVS 256
L ++ S+Q L + N+ + L NL + SN + G+IP S T + S
Sbjct: 296 SLQGLN---SLQALMLKGNTKFSTTIPENAFKGLKNLMILVLSNTNIQGSIPKSLTRLNS 352
Query: 257 LRILRLACNQLTGSLP 272
LR+L L N LTG +P
Sbjct: 353 LRVLHLEGNNLTGEIP 368
Score = 63.2 bits (152), Expect = 7e-10
Identities = 63/231 (27%), Positives = 103/231 (44%), Gaps = 16/231 (6%)
Query: 47 CPQNWYGILCSEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPM 106
C W+GI C DN V +F A+S+ ++ + S+ + +
Sbjct: 81 CRGRWHGIECMPDQ------DNVYHVVSLSFGALSDDTAFPTCDPQRSYVSESLTRLKHL 134
Query: 107 KSLKFLDLSLNKFNGSLPPSFVEL-RSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSN 165
K+L F L + +P L SL L L N F G +P+ L L+ LD H N
Sbjct: 135 KAL-FFYRCLGRAPQRIPAFLGRLGSSLQTLVLRENGFLGPIPDELGNLTNLKVLDLHKN 193
Query: 166 SFSGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAH 225
+G I F + + +DLS N+ +G++ G V L ++ L+++ N L G +
Sbjct: 194 HLNGSIPLSFNRFSGLRSLDLSGNRLTGSIP---GFV--LPALSVLDLNQNLLTGPV--P 246
Query: 226 DGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACNQLTGSLPETL 275
+ +L D S N++ G IP S + L +L L+ N+L+G P +L
Sbjct: 247 PTLTSCGSLIKIDLSRNRVTGPIPESINRLNQLVLLDLSYNRLSGPFPSSL 297
Score = 43.5 bits (101), Expect = 6e-04
Identities = 34/84 (40%), Positives = 46/84 (54%), Gaps = 7/84 (8%)
Query: 207 SIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIP-SFTFVVSLRILRLACN 265
S+Q L + N +G + D + L NL+V D N L G+IP SF LR L L+ N
Sbjct: 160 SLQTLVLRENGFLGPI--PDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGLRSLDLSGN 217
Query: 266 QLTGSLPETLLKESSMMLSELDLS 289
+LTGS+P +L LS LDL+
Sbjct: 218 RLTGSIPGFVLP----ALSVLDLN 237
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 68.2 bits (165), Expect = 2e-11
Identities = 77/292 (26%), Positives = 127/292 (43%), Gaps = 26/292 (8%)
Query: 1 MLLLLVNTAFGNRDIDALLELKKGIQNDPFGLVLNSWDSKSLESNGCPQNWYGILCS-EG 59
+LLL ++ A + D+ A+L LKK + N P W S+ P W I+C+
Sbjct: 15 LLLLSLSKADSDGDLSAMLSLKKSL-NPPSSF---GW------SDPDPCKWTHIVCTGTK 64
Query: 60 NVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKF 119
V I + ++ L G + + NL L L + N+ +G + +S + SL+ L LS N F
Sbjct: 65 RVTRIQIGHSGLQGTLS-PDLRNLSELERLELQWNNISGPVPSLSGLASLQVLMLSNNNF 123
Query: 120 NGSLPPSFVELRSLVYLNLSLNEF-SGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIF--- 175
+ F L SL + + N F S +P L+ +S + SG +
Sbjct: 124 DSIPSDVFQGLTSLQSVEIDNNPFKSWEIPESLRNASALQNFSANSANVSGSLPGFLGPD 183
Query: 176 -YQMGSVLHVDLSNNKFSGALDLGLGDVSFLFSIQHLNVSHNSLVGELFAHDGMPYLDNL 234
+ S+LH L+ N G L + L +Q L ++ L G++ M L +
Sbjct: 184 EFPGLSILH--LAFNNLEGELPMSLAG----SQVQSLWLNGQKLTGDITVLQNMTGLKEV 237
Query: 235 EVFDASNNQLVGNIPSFTFVVSLRILRLACNQLTGSLPETLLKESSMMLSEL 286
+ +N+ G +P F+ + L L L N TG +P +LL S+ + L
Sbjct: 238 WL---HSNKFSGPLPDFSGLKELESLSLRDNSFTGPVPASLLSLESLKVVNL 286
Score = 58.5 bits (140), Expect = 2e-08
Identities = 58/228 (25%), Positives = 94/228 (40%), Gaps = 46/228 (20%)
Query: 61 VISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSLNKFN 120
V S+ L+ L G+ L N+ L + + +N F+G + S +K L+ L L N F
Sbjct: 211 VQSLWLNGQKLTGDITVL--QNMTGLKEVWLHSNKFSGPLPDFSGLKELESLSLRDNSFT 268
Query: 121 GSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSF------------- 167
G +P S + L SL +NL+ N G VP VF ++ LD SNSF
Sbjct: 269 GPVPASLLSLESLKVVNLTNNHLQGPVP-VFKSSVSVD-LDKDSNSFCLSSPGECDPRVK 326
Query: 168 ------------------------SGDIMEIFYQMGSVLHVDLSNNKFSGALDLGLGDVS 203
+ + I G++ + L + +G + G +
Sbjct: 327 SLLLIASSFDYPPRLAESWKGNDPCTNWIGIACSNGNITVISLEKMELTGTISPEFGAIK 386
Query: 204 FLFSIQHLNVSHNSLVGELFAHDGMPYLDNLEVFDASNNQLVGNIPSF 251
S+Q + + N+L G + + L NL+ D S+N+L G +P F
Sbjct: 387 ---SLQRIILGINNLTGMI--PQELTTLPNLKTLDVSSNKLFGKVPGF 429
Score = 50.8 bits (120), Expect = 4e-06
Identities = 81/310 (26%), Positives = 122/310 (39%), Gaps = 54/310 (17%)
Query: 2 LLLLVNTAFGNRDIDALLELKK----GIQNDPFGLVLNSWD-SKSLESNGCPQNWYGILC 56
+L+L N F + D L I N+PF SW+ +SL + QN+
Sbjct: 115 VLMLSNNNFDSIPSDVFQGLTSLQSVEIDNNPF----KSWEIPESLRNASALQNFSA--- 167
Query: 57 SEGNVISITLDNASLVGEFNFLAISNLPMLHNLSVVNNHFTGSMLHISPMKSLKFLDLSL 116
+ NV + SL G FL P L L + N+ G + ++ L L+
Sbjct: 168 NSANV------SGSLPG---FLGPDEFPGLSILHLAFNNLEGELPMSLAGSQVQSLWLNG 218
Query: 117 NKFNGSLPPSFVELRSLVYLNLSLNEFSGTVPNVFHKLDQLEYLDFHSNSFSGDIMEIFY 176
K G + + L + L N+FSG +P+ F L +LE L NSF+G +
Sbjct: 219 QKLTGDITV-LQNMTGLKEVWLHSNKFSGPLPD-FSGLKELESLSLRDNSFTGPVPASLL 276
Query: 177 QMGSVLHVDLSNNKFSG---------ALDLGLGDVSF-LFSIQHLNVSHNSLVGELFAHD 226
+ S+ V+L+NN G ++DL SF L S + SL+ + D
Sbjct: 277 SLESLKVVNLTNNHLQGPVPVFKSSVSVDLDKDSNSFCLSSPGECDPRVKSLLLIASSFD 336
Query: 227 GMPYL--------------------DNLEVFDASNNQLVGNI-PSFTFVVSLRILRLACN 265
P L N+ V +L G I P F + SL+ + L N
Sbjct: 337 YPPRLAESWKGNDPCTNWIGIACSNGNITVISLEKMELTGTISPEFGAIKSLQRIILGIN 396
Query: 266 QLTGSLPETL 275
LTG +P+ L
Sbjct: 397 NLTGMIPQEL 406
Score = 46.6 bits (109), Expect = 7e-05
Identities = 38/118 (32%), Positives = 61/118 (51%), Gaps = 15/118 (12%)
Query: 32 LVLNSWD-----SKSLESNGCPQNWYGILCSEGNVISITLDNASLVGEFN--FLAISNLP 84
L+ +S+D ++S + N NW GI CS GN+ I+L+ L G + F AI +
Sbjct: 330 LIASSFDYPPRLAESWKGNDPCTNWIGIACSNGNITVISLEKMELTGTISPEFGAIKS-- 387
Query: 85 MLHNLSVVNNHFTGSM-LHISPMKSLKFLDLSLNKFNGSLPPSFVELRSLVYLNLSLN 141
L + + N+ TG + ++ + +LK LD+S NK G +P RS V +N + N
Sbjct: 388 -LQRIILGINNLTGMIPQELTTLPNLKTLDVSSNKLFGKVP----GFRSNVVVNTNGN 440
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.138 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,489,554
Number of Sequences: 164201
Number of extensions: 1431233
Number of successful extensions: 4321
Number of sequences better than 10.0: 241
Number of HSP's better than 10.0 without gapping: 120
Number of HSP's successfully gapped in prelim test: 122
Number of HSP's that attempted gapping in prelim test: 3179
Number of HSP's gapped (non-prelim): 700
length of query: 289
length of database: 59,974,054
effective HSP length: 109
effective length of query: 180
effective length of database: 42,076,145
effective search space: 7573706100
effective search space used: 7573706100
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC124972.17


