

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124972.13 - phase: 0
(935 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
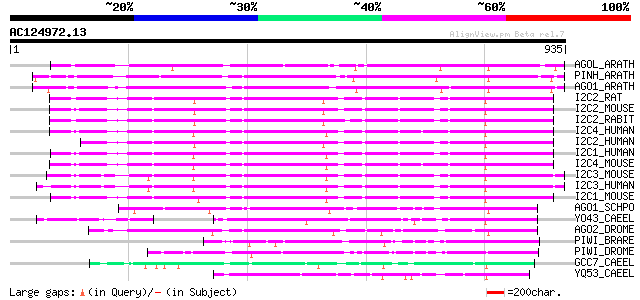
Score E
Sequences producing significant alignments: (bits) Value
AGOL_ARATH (Q9SJK3) Argonaute-like protein At2g27880 485 e-136
PINH_ARATH (Q9XGW1) PINHEAD protein (ZWILLE protein) 476 e-133
AGO1_ARATH (O04379) Argonaute protein 464 e-130
I2C2_RAT (Q9QZ81) Eukaryotic translation initiation factor 2C 2 ... 380 e-105
I2C2_MOUSE (Q8CJG0) Eukaryotic translation initiation factor 2C ... 379 e-104
I2C2_RABIT (O77503) Eukaryotic translation initiation factor 2C ... 378 e-104
I2C4_HUMAN (Q9HCK5) Eukaryotic translation initiation factor 2C ... 376 e-103
I2C2_HUMAN (Q9UKV8) Eukaryotic translation initiation factor 2C ... 370 e-102
I2C1_HUMAN (Q9UL18) Eukaryotic translation initiation factor 2C ... 365 e-100
I2C4_MOUSE (Q8CJF8) Eukaryotic translation initiation factor 2C ... 364 e-100
I2C3_MOUSE (Q8CJF9) Eukaryotic translation initiation factor 2C ... 363 1e-99
I2C3_HUMAN (Q9H9G7) Eukaryotic translation initiation factor 2C ... 362 3e-99
I2C1_MOUSE (Q8CJG1) Eukaryotic translation initiation factor 2C ... 357 1e-97
AGO1_SCHPO (O74957) Cell cycle control protein ago1 (RNA interfe... 318 3e-86
YO43_CAEEL (P34681) Hypothetical protein ZK757.3 in chromosome III 258 5e-68
AGO2_DROME (Q9VUQ5) Argonaute 2 protein 251 5e-66
PIWI_BRARE (Q8UVX0) Piwi protein 158 6e-38
PIWI_DROME (Q9VKM1) Piwi protein 140 2e-32
GCC7_CAEEL (Q21770) Germ cell expressed protein R06C7.1 94 1e-18
YQ53_CAEEL (Q09249) Hypothetical protein C16C10.3 in chromosome III 92 8e-18
>AGOL_ARATH (Q9SJK3) Argonaute-like protein At2g27880
Length = 997
Score = 485 bits (1249), Expect = e-136
Identities = 325/907 (35%), Positives = 475/907 (51%), Gaps = 102/907 (11%)
Query: 70 RRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQET 129
R G G+ G K+ + NHF V V + D Y + S+ V K R ++ + +
Sbjct: 150 RPGRGTLGKKVMVRANHFLVQVADRDLYHYDVSI------NPEVISKTVNRNVMKLLVKN 203
Query: 130 Y-GSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTD 188
Y S L GK AYDG K+L+T G L + EF V L + ++ ++G P
Sbjct: 204 YKDSHLGGKSPAYDGRKSLYTAGPLPFDSKEFVVNLAEKRADGSSGKDRP---------- 253
Query: 189 RKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGC 248
+KV + + L + L + E + I+VLD++LR +
Sbjct: 254 ------------FKVAVKNVTSTDLYQLQQFLDRKQREAPYDTIQVLDVVLRDKPSND-Y 300
Query: 249 LLVRQNFFHNDPKNFTDVGGGVLG-----CRGLHSSFRTTQSGLSLNIDVSTTMIVHPGP 303
+ V ++FFH G G LG RG S R TQ GLSLNIDVS P
Sbjct: 301 VSVGRSFFHTSLGKDARDGRGELGDGIEYWRGYFQSLRLTQMGLSLNIDVSARSFYEPIV 360
Query: 304 VVDFLIANQNVRD---PF-SLDWNKAKRTLKNLRITTSPTN--QEYKITGLSEMPCKDQL 357
V DF+ N+RD P D K K+ L+ L++ N + KI+G+S +P ++
Sbjct: 361 VTDFISKFLNIRDLNRPLRDSDRLKVKKVLRTLKVKLLHWNCTKSAKISGISSLPIRELR 420
Query: 358 FTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLV 417
FTL +D E TV YF + ++Y A LP I G RP ++P+ELC +
Sbjct: 421 FTL---------EDKSEKTVVQYFAEKYNYRVKYQA-LPAIQTGSDTRPVYLPMELCQID 470
Query: 418 SLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQ 477
QRYTK L+ Q ++L++ + Q+P +R + + + ++Y + + + G+S+T+
Sbjct: 471 EGQRYTKRLNEKQVTALLKATCQRPPDRENSIKNLVVKNNYNDD-LSKEFGMSVTTQLAS 529
Query: 478 VDGRVLQAPRLKF---GNGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLV 534
++ RVL P LK+ G + NPR G+WN +KK+V K+ W V+FS R D RGL
Sbjct: 530 IEARVLPPPMLKYHDSGKEKMVNPRLGQWNMIDKKMVNGAKVTSWTCVSFSTRID-RGLP 588
Query: 535 RDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKL-------PGAPKF 587
++ C + G+ C + +F+ P + + EH++ L PG +
Sbjct: 589 QEF--CKQLIGM-------CVSKGMEFKPQPAIPFISCPPEHIEEALLDIHKRAPGL-QL 638
Query: 588 LLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVND---QYLTNVLLKINAKLGGMN 644
L+ +L + S YG K+ E GIV+QC P +VN QY+ NV LKIN K GG N
Sbjct: 639 LIVILPDVTGS--YGKIKRICETELGIVSQCCQPRQVNKLNKQYMENVALKINVKTGGRN 696
Query: 645 SLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRT 704
++L +IP+++ PT+I+G DV+H PG+ PSIAAVV+S WP I+KYR V
Sbjct: 697 TVLNDAIRRNIPLITDRPTIIMGADVTHPQPGEDSSPSIAAVVASMDWPEINKYRGLVSA 756
Query: 705 QGAKVEMIDNLFKPVSDTE----DEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFN 760
Q + E+I +L+K V D + G+IRE I F ++G + P II +RDGVSE QF+
Sbjct: 757 QAHREEIIQDLYKLVQDPQRGLVHSGLIREHFIAFRRATG-QIPQRIIFYRDGVSEGQFS 815
Query: 761 QVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF--QPGSPD------NVPPGTV 812
QVL E++ I +AC L E + P+ ++ QK HHT+ F Q G+ D N+ PGTV
Sbjct: 816 QVLLHEMTAIRKACNSLQENYVPRVTFVIVQKRHHTRLFPEQHGNRDMTDKSGNIQPGTV 875
Query: 813 VDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTT 872
VD KICHP +DFY+ +HAG+ GTSRP HYHVLLDE GF+ D LQ L ++L Y Y R T
Sbjct: 876 VDTKICHPNEFDFYLNSHAGIQGTSRPAHYHVLLDENGFTADQLQMLTNNLCYTYARCTK 935
Query: 873 AISVVAPICYAHLAASQVGQFMKFEDKSETSSSHGGSGRDINASP-----IPQLPKLMDS 927
++S+V P YAHLAA + +M E+ S GGS R +++ I QLP + D+
Sbjct: 936 SVSIVPPAYYAHLAAFRARYYM------ESEMSDGGSSRSRSSTTGVGQVISQLPAIKDN 989
Query: 928 VCNSMFF 934
V MF+
Sbjct: 990 VKEVMFY 996
>PINH_ARATH (Q9XGW1) PINHEAD protein (ZWILLE protein)
Length = 988
Score = 476 bits (1224), Expect = e-133
Identities = 321/948 (33%), Positives = 485/948 (50%), Gaps = 104/948 (10%)
Query: 39 PPPP----------IVPADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFK 88
PPPP +I + + Q+ +K P R G G+ G K + NHF
Sbjct: 92 PPPPSQTTSSAVSVATAGEIVAVNHQMQMGVRKNSNFAP--RPGFGTLGTKCIVKANHFL 149
Query: 89 VNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETYGSELNGKDL-AYDGEKTL 147
++ D QY V + E V K R I+ + Y G+ L AYDG K+L
Sbjct: 150 ADLPTKD--LNQYDVTITPE----VSSKSVNRAIIAELVRLYKESDLGRRLPAYDGRKSL 203
Query: 148 FTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISF 207
+T G L EF+V + D NG R ++YKV I F
Sbjct: 204 YTAGELPFTWKEFSVKIVDEDDGIING--------------------PKRERSYKVAIKF 243
Query: 208 ASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVG 267
++ + + L G + QEA+++LDI+LR+ + K+ C + R +FF D K +G
Sbjct: 244 VARANMHHLGEFLAGKRADCPQEAVQILDIVLRELSVKRFCPVGR-SFFSPDIKTPQRLG 302
Query: 268 GGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDF---LIANQNVRDPFS-LDWN 323
G+ G + S R TQ GLSLNID+++ + P PV++F L+ + P S D
Sbjct: 303 EGLESWCGFYQSIRPTQMGLSLNIDMASAAFIEPLPVIEFVAQLLGKDVLSKPLSDSDRV 362
Query: 324 KAKRTLKNLRITTSP---TNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDY 380
K K+ L+ +++ + ++Y++ GL+ P ++ +F P +++ +V +Y
Sbjct: 363 KIKKGLRGVKVEVTHRANVRRKYRVAGLTTQPTRELMF--------PVDENCTMKSVIEY 414
Query: 381 FVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQ 440
F ++Q++ LPC+ VG K+ +++P+E C +V QRYTK L+ Q ++L++ + Q
Sbjct: 415 FQEMYGFTIQHT-HLPCLQVGNQKKASYLPMEACKIVEGQRYTKRLNEKQITALLKVTCQ 473
Query: 441 KPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFN 497
+P++R + ++ + Y +P + G++I+ V+ R+L AP LK+ G +D
Sbjct: 474 RPRDRENDILRTVQHNAYDQDPYAKEFGMNISEKLASVEARILPAPWLKYHENGKEKDCL 533
Query: 498 PRNGRWNFNNKKIVQPVKIEKWAVVNFSARCD---VRGLVRDLIKCGGMKGIHVEQPFDC 554
P+ G+WN NKK++ + + +WA VNFS RG +L + + G+
Sbjct: 534 PQVGQWNMMNKKMINGMTVSRWACVNFSRSVQENVARGFCNELGQMCEVSGMEFNP---- 589
Query: 555 FEENGQFRRAPPLVRVEKMFEHV----QSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLA 610
E A P +VEK +HV +K G LL + N LYG K+
Sbjct: 590 -EPVIPIYSARP-DQVEKALKHVYHTSMNKTKGKELELLLAILPDNNGSLYGDLKRICET 647
Query: 611 EFGIVTQCIAPT---RVNDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILG 667
E G+++QC +++ QYL NV LKIN K+GG N++L S IP+VS PT+I G
Sbjct: 648 ELGLISQCCLTKHVFKISKQYLANVSLKINVKMGGRNTVLVDAISCRIPLVSDIPTIIFG 707
Query: 668 MDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFK----PVSDTE 723
DV+H G+ PSIAAVV+S+ WP ++KY V Q + E+I +L+K PV T
Sbjct: 708 ADVTHPENGEESSPSIAAVVASQDWPEVTKYAGLVCAQAHRQELIQDLYKTWQDPVRGTV 767
Query: 724 DEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNP 783
G+IR+LLI F ++G +KP II +RDGVSE QF QVL EL I +AC L+ + P
Sbjct: 768 SGGMIRDLLISFRKATG-QKPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEPNYQP 826
Query: 784 KFLVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAGMIG 835
IV QK HHT+ F D N+ PGTVVD KICHP +DFY+C+HAG+ G
Sbjct: 827 PVTFIVVQKRHHTRLFANNHRDKNSTDRSGNILPGTVVDTKICHPTEFDFYLCSHAGIQG 886
Query: 836 TSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFMK 895
TSRP HYHVL DE F+ D +Q L ++L Y Y R T ++S+V P YAHLAA + +
Sbjct: 887 TSRPAHYHVLWDENNFTADGIQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRA----R 942
Query: 896 FEDKSETSSSHGGSGR---------DINASPIPQLPKLMDSVCNSMFF 934
F + E +G G+ D+ P LP L ++V MF+
Sbjct: 943 FYLEPEIMQDNGSPGKKNTKTTTVGDVGVKP---LPALKENVKRVMFY 987
>AGO1_ARATH (O04379) Argonaute protein
Length = 1048
Score = 464 bits (1193), Expect = e-130
Identities = 319/954 (33%), Positives = 482/954 (50%), Gaps = 108/954 (11%)
Query: 39 PPPPIVPADIEPIKIE---PQIVKKKLPT-----KVPMARRGLGSKGAKLPLLTNHFKVN 90
P P ++ E + +E P + +P+ K PM R G G G + + NHF
Sbjct: 144 PEPTVLAQQFEQLSVEQGAPSQAIQPIPSSSKAFKFPM-RPGKGQSGKRCIVKANHFFAE 202
Query: 91 VTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQETY-GSELNGKDLAYDGEKTLFT 149
+ + D Y V + E V +G R ++ ++ + Y S L + AYDG K+L+T
Sbjct: 203 LPDKD--LHHYDVTITPE----VTSRGVNRAVMKQLVDNYRDSHLGSRLPAYDGRKSLYT 256
Query: 150 IGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFAS 209
G L N EF + L D G G R + +KV I +
Sbjct: 257 AGPLPFNSKEFRINLLD----------EEVGAGG-----------QRREREFKVVIKLVA 295
Query: 210 KIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGG 269
+ L + L+G +++ QEA++VLDI+LR+ + + V ++F+ D +G G
Sbjct: 296 RADLHHLGMFLEGKQSDAPQEALQVLDIVLRELPTSR-YIPVGRSFYSPDIGKKQSLGDG 354
Query: 270 VLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFL--IANQNVRD-PFS-LDWNKA 325
+ RG + S R TQ GLSLNID+S+T + PV+ F+ + N+++ P S D K
Sbjct: 355 LESWRGFYQSIRPTQMGLSLNIDMSSTAFIEANPVIQFVCDLLNRDISSRPLSDADRVKI 414
Query: 326 KRTLKNLRITTSPTN---QEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFV 382
K+ L+ +++ + ++Y+I+GL+ + ++ F + +R + +V +YF
Sbjct: 415 KKALRGVKVEVTHRGNMRRKYRISGLTAVATRELTFPVDERNT--------QKSVVEYFH 466
Query: 383 NRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKP 442
+Q++ LPC+ VG RP ++P+E+C +V QRY+K L+ Q ++L++ + Q+P
Sbjct: 467 ETYGFRIQHT-QLPCLQVGNSNRPNYLPMEVCKIVEGQRYSKRLNERQITALLKVTCQRP 525
Query: 443 QERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKF---GNGEDFNPR 499
+R + + ++ +DY + + GI I++ V+ R+L P LK+ G P+
Sbjct: 526 IDREKDILQTVQLNDYAKDNYAQEFGIKISTSLASVEARILPPPWLKYHESGREGTCLPQ 585
Query: 500 NGRWNFNNKKIVQPVKIEKWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENG 559
G+WN NKK++ + W +NFS + L R + E C+
Sbjct: 586 VGQWNMMNKKMINGGTVNNWICINFSRQVQ-DNLARTFCQ---------ELAQMCYVSGM 635
Query: 560 QFRRAPPLVRVEKMFEHVQSKLP------------GAPKFLLCLLSERKNSDLYGPWKKK 607
F P L V E V+ L G LL ++ N LYG K+
Sbjct: 636 AFNPEPVLPPVSARPEQVEKVLKTRYHDATSKLSQGKEIDLLIVILPDNNGSLYGDLKRI 695
Query: 608 NLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTL 664
E GIV+QC V + QY+ NV LKIN K+GG N++L S IP+VS PT+
Sbjct: 696 CETELGIVSQCCLTKHVFKMSKQYMANVALKINVKVGGRNTVLVDALSRRIPLVSDRPTI 755
Query: 665 ILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTED 724
I G DV+H PG+ PSIAAVV+S+ WP I+KY V Q + E+I +LFK D +
Sbjct: 756 IFGADVTHPHPGEDSSPSIAAVVASQDWPEITKYAGLVCAQAHRQELIQDLFKEWKDPQK 815
Query: 725 E----GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEK 780
G+I+ELLI F S+G+ KP II +RDGVSE QF QVL EL I +AC L+
Sbjct: 816 GVVTGGMIKELLIAFRRSTGH-KPLRIIFYRDGVSEGQFYQVLLYELDAIRKACASLEAG 874
Query: 781 WNPKFLVIVAQKNHHTKFFQPGSPD--------NVPPGTVVDNKICHPRNYDFYMCAHAG 832
+ P +V QK HHT+ F D N+ PGTVVD+KICHP +DFY+C+HAG
Sbjct: 875 YQPPVTFVVVQKRHHTRLFAQNHNDRHSVDRSGNILPGTVVDSKICHPTEFDFYLCSHAG 934
Query: 833 MIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQ 892
+ GTSRP HYHVL DE F+ D LQ L ++L Y Y R T ++S+V P YAHLAA +
Sbjct: 935 IQGTSRPAHYHVLWDENNFTADGLQSLTNNLCYTYARCTRSVSIVPPAYYAHLAAFRARF 994
Query: 893 FMKFEDKSETSSSHGGSGR------------DINASPIPQLPKLMDSVCNSMFF 934
+M+ E S + G R ++NA+ P LP L ++V MF+
Sbjct: 995 YMEPETSDSGSMASGSMARGGGMAGRSTRGPNVNAAVRP-LPALKENVKRVMFY 1047
>I2C2_RAT (Q9QZ81) Eukaryotic translation initiation factor 2C 2
(eIF2C 2) (eIF-2C 2) (Golgi ER protein 95 kDa) (GERp95)
Length = 860
Score = 380 bits (977), Expect = e-105
Identities = 278/884 (31%), Positives = 442/884 (49%), Gaps = 102/884 (11%)
Query: 67 PMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVE-GKGAGRKILDR 125
P R G+ G + L N F++++ D Y ++ + +P + + R+I++
Sbjct: 26 PPPRPDFGTTGRTIKLQANFFEMDIPKIDIYHYELDI-------KPEKCPRRVNREIVEH 78
Query: 126 VQETYGSELNG-KDLAYDGEKTLFTIGSL--AQNKLEFTVVLEDVTSNRNNGNASPDGHG 182
+ + + +++ G + +DG K L+T L ++K+E V L G G
Sbjct: 79 MVQHFKTQIFGDRKPVFDGRKNLYTAMPLPIGRDKVELEVTLP--------------GEG 124
Query: 183 SPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQH 242
+ + +KV I + S + LQA+ +AL G E I+ LD+++R H
Sbjct: 125 --------------KDRIFKVSIKWVSCVSLQALHDALSGRLPSVPFETIQALDVVMR-H 169
Query: 243 AAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPG 302
V ++FF +GGG G H S R + + LNIDVS T
Sbjct: 170 LPSMRYTPVGRSFFTASEGCSNPLGGGREVWFGFHQSVRPSLWKMMLNIDVSATAFYKAQ 229
Query: 303 PVVDFLI------ANQNVRDPFSLDWNKAKRT--LKNLRITTSPTNQ---EYKITGLSEM 351
PV++F+ + + + P + D + K T +K L++ + Q +Y++ ++
Sbjct: 230 PVIEFVCEVLDFKSIEEQQKPLT-DSQRVKFTKEIKGLKVEITHCGQMKRKYRVCNVTRR 288
Query: 352 PCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPV 411
P Q F L++ T E TV YF +R K+ L+Y LPC+ VG+ ++ T++P+
Sbjct: 289 PASHQTFPLQQESG-----QTVECTVAQYFKDRHKLVLRYP-HLPCLQVGQEQKHTYLPL 342
Query: 412 ELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISI 471
E+C++V+ QR K L+ Q S+++ + + +R ++ ++++ + ++P +R GI +
Sbjct: 343 EVCNIVAGQRCIKKLTDNQTSTMIRATARSAPDRQEEISKLMRSASFNTDPYVREFGIMV 402
Query: 472 TSGFTQVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSAR-- 527
T V GRVLQ P + +G N P G W+ NK+ ++I+ WA+ F+ +
Sbjct: 403 KDEMTDVTGRVLQPPSILYGGRNKAIATPVQGVWDMRNKQFHTGIEIKVWAIACFAPQRQ 462
Query: 528 ---CDVRGLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPG 583
++ L K G+ ++ QP C + A VE MF H+++ G
Sbjct: 463 CTEVHLKSFTEQLRKISRDAGMPIQGQPCFC-------KYAQGADSVEPMFRHLKNTYAG 515
Query: 584 APKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPT---RVNDQYLTNVLLKINAKL 640
++ L + + +Y K+ G+ TQC+ R Q L+N+ LKIN KL
Sbjct: 516 LQLVVVILPGK---TPVYAEVKRVGDTVLGMATQCVQMKNVQRTTPQTLSNLCLKINVKL 572
Query: 641 GGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRA 700
GG+N++L + P V + P + LG DV+H G + PSIAAVV S ++Y A
Sbjct: 573 GGVNNILLPQGRPP---VFQQPVIFLGADVTHPPAGDGKKPSIAAVVGSMD-AHPNRYCA 628
Query: 701 CVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFN 760
VR Q + E+I +L ++RELLI FY S+ KP II +RDGVSE QF
Sbjct: 629 TVRVQQHRQEIIQDL---------AAMVRELLIQFYKST-RFKPTRIIFYRDGVSEGQFQ 678
Query: 761 QVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDNVPPGTVVD 814
QVL+ EL I EAC L++++ P IV QK HHT+ F + G N+P GT VD
Sbjct: 679 QVLHHELLAIREACIKLEKEYQPGITFIVVQKRHHTRLFCTDKNERVGKSGNIPAGTTVD 738
Query: 815 NKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAI 874
KI HP +DFY+C+HAG+ GTSRP+HYHVL D+ FS D+LQ L + L + Y R T ++
Sbjct: 739 TKITHPTEFDFYLCSHAGIQGTSRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSV 798
Query: 875 SVVAPICYAHLAASQVGQFM--KFEDKSETSSSHGGS-GRDINA 915
S+ AP YAHL A + + K D +E S + G S GRD A
Sbjct: 799 SIPAPAYYAHLVAFRARYHLVDKEHDSAEGSHTSGQSNGRDHQA 842
>I2C2_MOUSE (Q8CJG0) Eukaryotic translation initiation factor 2C 2
(eIF2C 2) (eIF-2C 2) (Piwi/argonaute family protain
meIF2C2)
Length = 860
Score = 379 bits (972), Expect = e-104
Identities = 278/883 (31%), Positives = 439/883 (49%), Gaps = 100/883 (11%)
Query: 67 PMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRV 126
P R G+ G + L N F++++ D Y ++ + + RP + R+I++ +
Sbjct: 26 PPPRPDFGTTGRTIKLQANFFEMDIPKIDIYHYELDIK---PEKRP---RRVNREIVEHM 79
Query: 127 QETYGSELNG-KDLAYDGEKTLFTIGSL--AQNKLEFTVVLEDVTSNRNNGNASPDGHGS 183
+ + +++ G + +DG K L+T L ++K+E V L G G
Sbjct: 80 VQHFKTQIFGDRKPVFDGRKNLYTAMPLPIGRDKVELEVTLP--------------GEG- 124
Query: 184 PNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHA 243
+ + KV I + S + LQA+ +AL G E I+ LD+++R H
Sbjct: 125 -------------KDRILKVSIKWVSCVSLQALHDALSGRLPSVPFETIQALDVVMR-HL 170
Query: 244 AKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGP 303
V ++FF +GGG G H S R + + LNIDVS T P
Sbjct: 171 PSMRYTPVGRSFFTASEGCSNPLGGGREVWFGFHQSVRPSLWKMMLNIDVSATAFYKAQP 230
Query: 304 VVDFLI------ANQNVRDPFSLDWNKAKRT--LKNLRITTSPTNQ---EYKITGLSEMP 352
V++F+ + + + P + D + K T +K L++ + Q +Y++ ++ P
Sbjct: 231 VIEFVCEVLDFKSIEEQQKPLT-DSQRVKFTKEIKGLKVEITHCGQMKRKYRVCNVTRRP 289
Query: 353 CKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVE 412
Q F L++ T E TV YF +R K+ L+Y LPC+ VG+ ++ T++P+E
Sbjct: 290 ASHQTFPLQQESG-----QTVECTVAQYFKDRHKLVLRYP-HLPCLQVGQEQKHTYLPLE 343
Query: 413 LCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISIT 472
+C++V+ QR K L+ Q S+++ + + +R ++ ++++ + ++P +R GI +
Sbjct: 344 VCNIVAGQRCIKKLTDDQTSTMIRATARSAPDRQEEISKLMRSASFNTDPYVREFGIMVK 403
Query: 473 SGFTQVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSAR--- 527
T V GRVLQ P + +G N P G W+ NK+ ++I+ WA+ F+ +
Sbjct: 404 DEMTDVTGRVLQPPSILYGGRNKAIATPVQGVWDMRNKQFHTGIEIKVWAIACFAPQRQC 463
Query: 528 --CDVRGLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGA 584
++ L K G+ ++ QP C + A VE MF H+++ G
Sbjct: 464 TEVHLKSFTEQLRKISRDAGMPIQGQPCFC-------KYAQGADSVEPMFRHLKNTYAGL 516
Query: 585 PKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPT---RVNDQYLTNVLLKINAKLG 641
++ L + + +Y K+ G+ TQC+ R Q L+N+ LKIN KLG
Sbjct: 517 QLVVVILPGK---TPVYAEVKRVGDTVLGMATQCVQMKNVQRTTPQTLSNLCLKINVKLG 573
Query: 642 GMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRAC 701
G+N++L + P V + P + LG DV+H G + PSIAAVV S ++Y A
Sbjct: 574 GVNNILLPQGRPP---VFQQPVIFLGADVTHPPAGDGKKPSIAAVVGSMD-AHPNRYCAT 629
Query: 702 VRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQ 761
VR Q + E+I +L ++RELLI FY S+ KP II +RDGVSE QF Q
Sbjct: 630 VRVQQHRQEIIQDL---------AAMVRELLIQFYKST-RFKPTRIIFYRDGVSEGQFQQ 679
Query: 762 VLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDNVPPGTVVDN 815
VL+ EL I EAC L++ + P IV QK HHT+ F + G N+P GT VD
Sbjct: 680 VLHHELLAIREACIKLEKDYQPGITFIVVQKRHHTRLFCTDKNERVGKSGNIPAGTTVDT 739
Query: 816 KICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAIS 875
KI HP +DFY+C+HAG+ GT RP+HYHVL D+ FS D+LQ L + L + Y R T ++S
Sbjct: 740 KITHPTEFDFYLCSHAGIQGTGRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSVS 799
Query: 876 VVAPICYAHLAASQVGQFM--KFEDKSETSSSHGGS-GRDINA 915
+ AP YAHL A + + K D +E S + G S GRD A
Sbjct: 800 IPAPAYYAHLVAFRARYHLVDKEHDSAEGSHTSGQSNGRDHQA 842
>I2C2_RABIT (O77503) Eukaryotic translation initiation factor 2C 2
(eIF2C 2) (eIF-2C 2) (Fragment)
Length = 840
Score = 378 bits (970), Expect = e-104
Identities = 277/884 (31%), Positives = 438/884 (49%), Gaps = 102/884 (11%)
Query: 67 PMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVE-GKGAGRKILDR 125
P R G+ G + L N F++++ D Y ++ + +P + + R+I++
Sbjct: 6 PPPRPDFGTSGRTIKLQANFFEMDIPKIDIYHYELDI-------KPEKCPRRVNREIVEH 58
Query: 126 VQETYGSELNG-KDLAYDGEKTLFTIGSL--AQNKLEFTVVLEDVTSNRNNGNASPDGHG 182
+ + + +++ G + +DG K L+T L + K+E V L G G
Sbjct: 59 MVQHFKAQIFGDRKPVFDGRKNLYTAMPLPIGREKVELEVTLP--------------GEG 104
Query: 183 SPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQH 242
+ + +KV I + S + LQA+ +AL G E I+ LD+++R H
Sbjct: 105 --------------KDRIFKVSIKWVSCVSLQALHDALSGRLPSVPFETIQALDVVMR-H 149
Query: 243 AAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPG 302
V ++FF +GGG G H S R + + LNIDVS T
Sbjct: 150 LPSMRYTPVGRSFFTASEGCSNPLGGGREVWFGFHQSVRPSLWKMMLNIDVSATAFYKAQ 209
Query: 303 PVVDFLI------ANQNVRDPFSLDWNKAKRT--LKNLRITTSPTNQ---EYKITGLSEM 351
PV++F+ + + + P + D + K T +K L++ + Q +Y++ ++
Sbjct: 210 PVIEFVCEVLDFKSIEEQQKPLT-DSQRVKFTKEIKGLKVEITHCGQMKRKYRVCNVTRR 268
Query: 352 PCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPV 411
P Q F L++ T E TV YF +R K+ L+Y LPC+ VG+ ++ T++P+
Sbjct: 269 PASHQTFPLQQESG-----QTVECTVAQYFKDRHKLVLRYP-HLPCLQVGQEQKHTYLPL 322
Query: 412 ELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISI 471
E+C++V+ QR K L+ Q S+++ + + +R ++ ++++ + ++P +R GI +
Sbjct: 323 EVCNIVAGQRCIKKLTDNQTSTMIRATARSAPDRQEEISKLMRSASFNTDPYVREFGIMV 382
Query: 472 TSGFTQVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSAR-- 527
T V GRVLQ P + +G N P G W+ NK+ ++I+ WA+ F+ +
Sbjct: 383 KDEMTDVTGRVLQPPSILYGGRNKAIATPVQGVWDMRNKQFHTGIEIKVWAIACFAPQRQ 442
Query: 528 ---CDVRGLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPG 583
++ L K G+ ++ QP C G P MF H+++ G
Sbjct: 443 CTEVHLKSFTEQLRKISRDAGMPIQGQPCFCKYAQGADSVGP-------MFRHLKNTYAG 495
Query: 584 APKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPT---RVNDQYLTNVLLKINAKL 640
++ L + + +Y K+ G+ TQC+ R Q L+N+ LKIN KL
Sbjct: 496 LQLVVVILPGK---TPVYAEVKRVGDTVLGMATQCVQMKNVQRTTPQTLSNLCLKINVKL 552
Query: 641 GGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRA 700
GG+N++L + P V + P + LG DV+H G + PSIAAVV S ++Y A
Sbjct: 553 GGVNNILLPQGRPP---VFQQPVIFLGADVTHPPAGDGKKPSIAAVVGSMD-AHPNRYCA 608
Query: 701 CVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFN 760
VR Q + E+I +L ++RELLI FY S+ KP II +RDGVSE QF
Sbjct: 609 TVRVQQHRQEIIQDL---------AAMVRELLIQFYKST-RFKPTRIIFYRDGVSEGQFQ 658
Query: 761 QVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDNVPPGTVVD 814
QVL+ EL I EAC L++ + P IV QK HHT+ F + G N+P GT VD
Sbjct: 659 QVLHHELLAIREACIKLEKDYQPGITFIVVQKRHHTRLFCTDKNERVGKSGNIPAGTTVD 718
Query: 815 NKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAI 874
KI HP +DFY+C+HAG+ GTSRP+HYHVL D+ FS D+LQ L + L + Y R T ++
Sbjct: 719 TKITHPTEFDFYLCSHAGIQGTSRPSHYHVLWDDNRFSSDELQILTYQLCHTYVRCTRSV 778
Query: 875 SVVAPICYAHLAASQVGQFM--KFEDKSETSSSHGGS-GRDINA 915
S+ AP YAHL A + + K D +E S + G S GRD A
Sbjct: 779 SIPAPAYYAHLVAFRARYHLVDKEHDSAEGSHTSGQSNGRDHQA 822
>I2C4_HUMAN (Q9HCK5) Eukaryotic translation initiation factor 2C 4
(eIF2C 4) (eIF-2C 4) (Argonaute 4)
Length = 861
Score = 376 bits (965), Expect = e-103
Identities = 275/885 (31%), Positives = 433/885 (48%), Gaps = 92/885 (10%)
Query: 67 PMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRV 126
P R GLG+ G + LL NHF+V + D Y + + + RP + R+++D +
Sbjct: 15 PPRRPGLGTVGKPIRLLANHFQVQIPKIDVYHYDVDIK---PEKRP---RRVNREVVDTM 68
Query: 127 QETYGSELNG-KDLAYDGEKTLFTIGSL--AQNKLEFTVVLEDVTSNRNNGNASPDGHGS 183
+ ++ G + YDG++ ++T L +++++ V L G G
Sbjct: 69 VRHFKMQIFGDRQPGYDGKRNMYTAHPLPIGRDRVDMEVTLP--------------GEG- 113
Query: 184 PNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHA 243
+ +T+KV + + S + LQ + AL GH E ++++ LD+I R H
Sbjct: 114 -------------KDQTFKVSVQWVSVVSLQLLLEALAGHLNEVPDDSVQALDVITR-HL 159
Query: 244 AKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGP 303
V ++FF + +GGG G H S R + LNIDVS T P
Sbjct: 160 PSMRYTPVGRSFFSPPEGYYHPLGGGREVWFGFHQSVRPAMWNMMLNIDVSATAFYRAQP 219
Query: 304 VVDFL-----IANQNVRDPFSLDWNKAKRT--LKNLRITTSPTNQ---EYKITGLSEMPC 353
+++F+ I N N + D + K T ++ L++ + Q +Y++ ++ P
Sbjct: 220 IIEFMCEVLDIQNINEQTKPLTDSQRVKFTKEIRGLKVEVTHCGQMKRKYRVCNVTRRPA 279
Query: 354 KDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVEL 413
Q F L+ E TV YF + + L+Y LPC+ VG+ ++ T++P+E+
Sbjct: 280 SHQTFPLQLENG-----QAMECTVAQYFKQKYSLQLKYP-HLPCLQVGQEQKHTYLPLEV 333
Query: 414 CSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDY--GSEPMLRNCGISI 471
C++V+ QR K L+ Q S++++ + + +R ++ +K++ G +P L+ GI +
Sbjct: 334 CNIVAGQRCIKKLTDNQTSTMIKATARSAPDRQEEISRLVKSNSMVGGPDPYLKEFGIVV 393
Query: 472 TSGFTQVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCD 529
+ T++ GRVL AP L++G N P G W+ K+ ++I+ WAV F+ +
Sbjct: 394 HNEMTELTGRVLPAPMLQYGGRNKTVATPNQGVWDMRGKQFYAGIEIKVWAVACFAPQKQ 453
Query: 530 VR-----GLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPG 583
R L K G+ ++ QP C + A VE MF+H++ G
Sbjct: 454 CREDLLKSFTDQLRKISKDAGMPIQGQPCFC-------KYAQGADSVEPMFKHLKMTYVG 506
Query: 584 APKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKL 640
++ L + + +Y K+ G+ TQC+ V + Q L+N+ LKINAKL
Sbjct: 507 LQLIVVILPGK---TPVYAEVKRVGDTLLGMATQCVQVKNVVKTSPQTLSNLCLKINAKL 563
Query: 641 GGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRA 700
GG+N++L PS V + P + LG DV+H G + PSIAAVV S S+Y A
Sbjct: 564 GGINNVLVPHQRPS---VFQQPVIFLGADVTHPPAGDGKKPSIAAVVGSMDGHP-SRYCA 619
Query: 701 CVRTQGAKVEMIDNLFKPVSDTED-EGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQF 759
VR Q ++ E+ L +D ++RELLI FY S+ KP II +R GVSE Q
Sbjct: 620 TVRVQTSRQEISQELLYSQEVIQDLTNMVRELLIQFYKST-RFKPTRIIYYRGGVSEGQM 678
Query: 760 NQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDNVPPGTVV 813
QV EL I +AC L+E + P IV QK HHT+ F + G NVP GT V
Sbjct: 679 KQVAWPELIAIRKACISLEEDYRPGITYIVVQKRHHTRLFCADKTERVGKSGNVPAGTTV 738
Query: 814 DNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTA 873
D+ I HP +DFY+C+HAG+ GTSRP+HY VL D+ F+ D+LQ L + L + Y R T +
Sbjct: 739 DSTITHPSEFDFYLCSHAGIQGTSRPSHYQVLWDDNCFTADELQLLTYQLCHTYVRCTRS 798
Query: 874 ISVVAPICYAHLAASQVGQFMKFEDKSETSSSH---GGSGRDINA 915
+S+ AP YA L A + + +D SH +GRD A
Sbjct: 799 VSIPAPAYYARLVAFRARYHLVDKDHDSAEGSHVSGQSNGRDPQA 843
>I2C2_HUMAN (Q9UKV8) Eukaryotic translation initiation factor 2C 2
(eIF2C 2) (eIF-2C 2)
Length = 851
Score = 370 bits (950), Expect = e-102
Identities = 268/830 (32%), Positives = 419/830 (50%), Gaps = 94/830 (11%)
Query: 120 RKILDRVQETYGSELNG-KDLAYDGEKTLFTIGSL--AQNKLEFTVVLEDVTSNRNNGNA 176
R+I++ + + + +++ G + +DG K L+T L ++K+E V L
Sbjct: 64 REIVEHMVQHFKTQIFGDRKPVFDGRKNLYTAMPLPIGRDKVELEVTLP----------- 112
Query: 177 SPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLD 236
G G + + +KV I + S + LQA+ +AL G E I+ LD
Sbjct: 113 ---GEG--------------KDRIFKVSIKWVSCVSLQALHDALSGRLPSVPFETIQALD 155
Query: 237 IILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTT 296
+++R H V ++FF +GGG G H S R + + LNIDVS T
Sbjct: 156 VVMR-HLPSMRYTPVGRSFFTASEGCSNPLGGGREVWFGFHQSVRPSLWKMMLNIDVSAT 214
Query: 297 MIVHPGPVVDFLI------ANQNVRDPFSLDWNKAKRT--LKNLRITTSPTNQ---EYKI 345
PV++F+ + + + P + D + K T +K L++ + Q +Y++
Sbjct: 215 AFYKAQPVIEFVCEVLDFKSIEEQQKPLT-DSQRVKFTKEIKGLKVEITHCGQMKRKYRV 273
Query: 346 TGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKR 405
++ P Q F L++ T E TV YF +R K+ L+Y LPC+ VG+ ++
Sbjct: 274 CNVTRRPASHQTFPLQQESG-----QTVECTVAQYFKDRHKLVLRYP-HLPCLQVGQEQK 327
Query: 406 PTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLR 465
T++P+E+C++V+ QR K L+ Q S+++ + + +R ++ ++++ + ++P +R
Sbjct: 328 HTYLPLEVCNIVAGQRCIKKLTDNQTSTMIRATARSAPDRQEEISKLMRSASFNTDPYVR 387
Query: 466 NCGISITSGFTQVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVN 523
GI + T V GRVLQ P + +G N P G W+ NK+ ++I+ WA+
Sbjct: 388 EFGIMVKDEMTDVTGRVLQPPSILYGGRNKAIATPVQGVWDMRNKQFHTGIEIKVWAIAC 447
Query: 524 FSAR-----CDVRGLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHV 577
F+ + ++ L K G+ ++ QP C + A VE MF H+
Sbjct: 448 FAPQRQCTEVHLKSFTEQLRKISRDAGMPIQGQPCFC-------KYAQGADSVEPMFRHL 500
Query: 578 QSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPT---RVNDQYLTNVLL 634
++ G ++ L + + +Y K+ G+ TQC+ R Q L+N+ L
Sbjct: 501 KNTYAGLQLVVVILPGK---TPVYAEVKRVGDTVLGMATQCVQMKNVQRTTPQTLSNLCL 557
Query: 635 KINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPL 694
KIN KLGG+N++L + P V + P + LG DV+H G + PSIAAVV S
Sbjct: 558 KINVKLGGVNNILLPQGRPP---VFQQPVIFLGADVTHPPAGDGKKPSIAAVVGSMD-AH 613
Query: 695 ISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGV 754
++Y A VR Q + E+I +L ++RELLI FY S+ KP II +RDGV
Sbjct: 614 PNRYCATVRVQQHRQEIIQDL---------AAMVRELLIQFYKST-RFKPTRIIFYRDGV 663
Query: 755 SESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDNVP 808
SE QF QVL+ EL I EAC L++ + P IV QK HHT+ F + G N+P
Sbjct: 664 SEGQFQQVLHHELLAIREACIKLEKDYQPGITFIVVQKRHHTRLFCTDKNERVGKSGNIP 723
Query: 809 PGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQ 868
GT VD KI HP +DFY+C+HAG+ GTSRP+HYHVL D+ FS D+LQ L + L + Y
Sbjct: 724 AGTTVDTKITHPTEFDFYLCSHAGIQGTSRPSHYHVLWDDNRFSSDELQILTYQLCHTYV 783
Query: 869 RSTTAISVVAPICYAHLAASQVGQFM--KFEDKSETSSSHGGS-GRDINA 915
R T ++S+ AP YAHL A + + K D +E S + G S GRD A
Sbjct: 784 RCTRSVSIPAPAYYAHLVAFRARYHLVDKEHDSAEGSHTSGQSNGRDHQA 833
>I2C1_HUMAN (Q9UL18) Eukaryotic translation initiation factor 2C 1
(eIF2C 1) (eIF-2C 1) (Putative RNA-binding protein Q99)
Length = 857
Score = 365 bits (936), Expect = e-100
Identities = 268/879 (30%), Positives = 433/879 (48%), Gaps = 98/879 (11%)
Query: 70 RRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQET 129
R G+G+ G + LL N+F+V++ D Y ++ + D P + R++++ + +
Sbjct: 26 RPGIGTVGKPIKLLANYFEVDIPKIDVYHYEVDIK---PDKCP---RRVNREVVEYMVQH 79
Query: 130 YGSELNG-KDLAYDGEKTLFTIGSL--AQNKLEFTVVLEDVTSNRNNGNASPDGHGSPND 186
+ ++ G + YDG+K ++T+ +L +++F V + G G
Sbjct: 80 FKPQIFGDRKPVYDGKKNIYTVTALPIGNERVDFEVTIP--------------GEG---- 121
Query: 187 TDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQ 246
+ + +KV I + + + + + AL + E+++ LD+ +R H A
Sbjct: 122 ----------KDRIFKVSIKWLAIVSWRMLHEALVSGQIPVPLESVQALDVAMR-HLASM 170
Query: 247 GCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVD 306
V ++FF + +GGG G H S R + LNIDVS T PV++
Sbjct: 171 RYTPVGRSFFSPPEGYYHPLGGGREVWFGFHQSVRPAMWKMMLNIDVSATAFYKAQPVIE 230
Query: 307 FL-----IANQNVRDPFSLDWNKAKRT--LKNLRITTSPTNQ---EYKITGLSEMPCKDQ 356
F+ I N N + D + + T +K L++ + Q +Y++ ++ P Q
Sbjct: 231 FMCEVLDIRNINEQPKPLTDSQRVRFTKEIKGLKVEVTHCGQMKRKYRVCNVTRRPASHQ 290
Query: 357 LFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSL 416
F L+ T E TV YF + + L+Y LPC+ VG+ ++ T++P+E+C++
Sbjct: 291 TFPLQLESG-----QTVECTVAQYFKQKYNLQLKYP-HLPCLQVGQEQKHTYLPLEVCNI 344
Query: 417 VSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFT 476
V+ QR K L+ Q S++++ + + +R ++ +K + Y +P ++ GI + T
Sbjct: 345 VAGQRCIKKLTDNQTSTMIKATARSAPDRQEEISRLMKNASYNLDPYIQEFGIKVKDDMT 404
Query: 477 QVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVR--- 531
+V GRVL AP L++G N P G W+ K+ ++I+ WA+ F+ + R
Sbjct: 405 EVTGRVLPAPILQYGGRNRAIATPNQGVWDMRGKQFYNGIEIKVWAIACFAPQKQCREEV 464
Query: 532 --GLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGAPKFL 588
L K G+ ++ QP C + A VE MF H+++ G +
Sbjct: 465 LKNFTDQLRKISKDAGMPIQGQPCFC-------KYAQGADSVEPMFRHLKNTYSGLQLII 517
Query: 589 LCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKLGGMNS 645
+ L + + +Y K+ G+ TQC+ V + Q L+N+ LKIN KLGG+N+
Sbjct: 518 VILPGK---TPVYAEVKRVGDTLLGMATQCVQVKNVVKTSPQTLSNLCLKINVKLGGINN 574
Query: 646 LLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQ 705
+L V H S V + P + LG DV+H G + PSI AVV S S+Y A VR Q
Sbjct: 575 IL-VPHQRSA--VFQQPVIFLGADVTHPPAGDGKKPSITAVVGSMD-AHPSRYCATVRVQ 630
Query: 706 GAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNI 765
+ E+I++L ++RELLI FY S+ KP II +RDGV E Q Q+L+
Sbjct: 631 RPRQEIIEDL---------SYMVRELLIQFYKST-RFKPTRIIFYRDGVPEGQLPQILHY 680
Query: 766 ELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDNVPPGTVVDNKICH 819
EL I +AC L++ + P IV QK HHT+ F + G N+P GT VD I H
Sbjct: 681 ELLAIRDACIKLEKDYQPGITYIVVQKRHHTRLFCADKNERIGKSGNIPAGTTVDTNITH 740
Query: 820 PRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAP 879
P +DFY+C+HAG+ GTSRP+HY+VL D+ F+ D+LQ L + L + Y R T ++S+ AP
Sbjct: 741 PFEFDFYLCSHAGIQGTSRPSHYYVLWDDNRFTADELQILTYQLCHTYVRCTRSVSIPAP 800
Query: 880 ICYAHLAASQVGQFM--KFEDKSETSSSHGGS-GRDINA 915
YA L A + + K D E S G S GRD A
Sbjct: 801 AYYARLVAFRARYHLVDKEHDSGEGSHISGQSNGRDPQA 839
>I2C4_MOUSE (Q8CJF8) Eukaryotic translation initiation factor 2C 4
(eIF2C 4) (eIF-2C 4) (Piwi/argonaute family protain
meIF2C4)
Length = 861
Score = 364 bits (934), Expect = e-100
Identities = 270/885 (30%), Positives = 429/885 (47%), Gaps = 92/885 (10%)
Query: 67 PMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRV 126
P R GLG+ G + LL NHF+V + D Y + + + RP + R+++D +
Sbjct: 15 PPRRPGLGTVGKPIRLLANHFQVQIPKIDVYHYDVDIK---PEKRP---RRVNREVVDTM 68
Query: 127 QETYGSELNG-KDLAYDGEKTLFTIGSL--AQNKLEFTVVLEDVTSNRNNGNASPDGHGS 183
+ ++ G + YDG++ ++T L +++++ V L G G
Sbjct: 69 VRHFKMQIFGDRQPGYDGKRNMYTAHPLPIGRDRIDMEVTLP--------------GEG- 113
Query: 184 PNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHA 243
+ +T+KV + + S LQ + AL GH E ++++ LD+I R H
Sbjct: 114 -------------KDQTFKVSVQWVSVASLQLLLEALAGHLNEVPDDSVQALDVITR-HL 159
Query: 244 AKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGP 303
V ++FF + +GGG G H S R + LNIDVS T P
Sbjct: 160 PSMRYTPVGRSFFSPPEGYYHPLGGGREVWFGFHQSVRPAMWNMMLNIDVSATAFYRAQP 219
Query: 304 VVDFL-----IANQNVRDPFSLDWNKAKRT--LKNLRITTSPTNQ---EYKITGLSEMPC 353
+++F+ I N N + D + K T ++ L++ + Q +Y++ ++ P
Sbjct: 220 IIEFMCEVLDIQNINEQTKPLTDSQRVKFTKEIRGLKVEVTHCGQMKRKYRVCNVTRRPA 279
Query: 354 KDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVEL 413
Q F L+ E TV YF + + L++ LPC+ VG+ ++ T++P+E+
Sbjct: 280 SHQTFPLQLENG-----QAMECTVAQYFKQKYSLQLKHP-HLPCLQVGQEQKHTYLPLEV 333
Query: 414 CSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDY--GSEPMLRNCGISI 471
C++V+ QR K L+ Q S++++ + + +R ++ +K++ G +P L+ GI +
Sbjct: 334 CNIVAGQRCIKKLTDNQTSTMIKATARSAPDRQEEISRLVKSNSMVGGPDPYLKEFGIVV 393
Query: 472 TSGFTQVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCD 529
+ T++ GRVL AP L++G N P G W+ K+ ++I+ WAV F+ +
Sbjct: 394 HNEMTELTGRVLPAPMLQYGGRNKTVATPSQGVWDMRGKQFYAGIEIKVWAVACFAPQKQ 453
Query: 530 VR-----GLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPG 583
R L K G+ ++ QP C + A VE MF+H++ G
Sbjct: 454 CREDLLKSFTDQLRKISKDAGMPIQGQPCFC-------KYAQGADSVEPMFKHLKMTYVG 506
Query: 584 APKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNVLLKINAKL 640
++ L + + +Y K+ G+ TQC+ V + Q L+N+ LK+NAKL
Sbjct: 507 LQLIVVILPGK---TPVYAEVKRVGDTLLGMATQCVQIKNVVKTSPQTLSNLCLKMNAKL 563
Query: 641 GGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRA 700
GG+N++ PS V + P + LG DV+H G + PSIAAVV S S+Y A
Sbjct: 564 GGINNVPVPHQRPS---VFQQPVIFLGADVTHPPAGDGKKPSIAAVVGSMDGHP-SRYCA 619
Query: 701 CVRTQGAKVEMIDNLFKPVSDTED-EGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQF 759
V Q ++ E+ L +D + RELLI FY S+ KP II +R GVSE Q
Sbjct: 620 TVWVQTSRQEIAQELLYSQEVVQDLTSMARELLIQFYKST-RFKPTRIIYYRGGVSEGQM 678
Query: 760 NQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDNVPPGTVV 813
QV EL I +AC L+E + P IV QK HHT+ F + G NVP GT V
Sbjct: 679 KQVAWPELIAIRKACISLEEDYRPGITYIVVQKRHHTRLFCADKMERVGKSGNVPAGTTV 738
Query: 814 DNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTA 873
D+ + HP +DFY+C+HAG+ GTSRP+HY VL D+ F+ D+LQ L + L + Y R T +
Sbjct: 739 DSTVTHPSEFDFYLCSHAGIQGTSRPSHYQVLWDDNCFTADELQLLTYQLCHTYVRCTRS 798
Query: 874 ISVVAPICYAHLAASQVGQFMKFEDKSETSSSH---GGSGRDINA 915
+S+ AP YA L A + + +D SH +GRD A
Sbjct: 799 VSIPAPAYYARLVAFRARYHLVDKDHDSAEGSHVSGQSNGRDPQA 843
>I2C3_MOUSE (Q8CJF9) Eukaryotic translation initiation factor 2C 3
(eIF2C 3) (eIF-2C 3) (Piwi/argonaute family protain
meIF2C3)
Length = 860
Score = 363 bits (931), Expect = 1e-99
Identities = 280/911 (30%), Positives = 436/911 (47%), Gaps = 103/911 (11%)
Query: 63 PTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKI 122
P + R G G+ G + LL N F+V + D Y ++ + D P + R++
Sbjct: 13 PLFIVPRRPGYGTMGKPIKLLANCFQVEIPKIDVYLYEVDIK---PDKCP---RRVNREV 66
Query: 123 LD-RVQETYGSELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGH 181
+D +VQ + + YDG+++L+T L + T V DVT G P
Sbjct: 67 VDSKVQHFKVTIFGDRRPVYDGKRSLYTANPLP---VATTGVDLDVTLPGEGGKDRP--- 120
Query: 182 GSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEA--------IR 233
+KV + F S++ + AL G E +
Sbjct: 121 -------------------FKVSVKFVSRVSWHLLHEALAGGTLPEPLELDKPVSTNPVH 161
Query: 234 VLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDV 293
+D++LR H V ++FF +GGG G H S R + LNIDV
Sbjct: 162 AVDVVLR-HLPSMKYTPVGRSFFSAPEGYDHPLGGGREVWFGFHQSVRPAMWKMMLNIDV 220
Query: 294 STTMIVHPGPVVDFL-----IANQNVRDPFSLDWNKAKRT--LKNLRITTS---PTNQEY 343
S T PV+ F+ I N + + D ++ K T +K L++ + ++Y
Sbjct: 221 SATAFYKAQPVIQFMCEVLDIHNIDEQPRPLTDSHRVKFTKEIKGLKVEVTHCGTMRRKY 280
Query: 344 KITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKP 403
++ ++ P Q F L+ T E TV YF + + L+Y LPC+ VG+
Sbjct: 281 RVCNVTRRPASHQTFPLQLENG-----QTVERTVAQYFREKYTLQLKYP-HLPCLQVGQE 334
Query: 404 KRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPM 463
++ T++P+E+C++V+ QR K L+ Q S++++ + + +R ++ +++++Y ++P
Sbjct: 335 QKHTYLPLEVCNIVAGQRCIKKLTDNQTSTMIKATARSAPDRQEEISRLVRSANYETDPF 394
Query: 464 LRNCGISITSGFTQVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAV 521
++ + + V GRVL AP L++G N P +G W+ K+ V+I+ WA+
Sbjct: 395 VQEFQLKVRDEMAHVTGRVLPAPMLQYGGRNRTVATPSHGVWDMRGKQFHTGVEIKMWAI 454
Query: 522 VNFSARCDVR-----GLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFE 575
F+ + R G L K G+ ++ QP C + A VE MF
Sbjct: 455 ACFATQRQCREEILKGFTDQLRKISKDAGMPIQGQPCFC-------KYAQGADSVEPMFR 507
Query: 576 HVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNV 632
H+++ G ++ L + + +Y K+ G+ TQC+ V + Q L+N+
Sbjct: 508 HLKNTYSGLQLIIVILPGK---TPVYAEVKRVGDTLLGMATQCVQVKNVIKTSPQTLSNL 564
Query: 633 LLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQW 692
LKIN KLGG+N++L PS V + P + LG DV+H G + PSIAAVV S
Sbjct: 565 CLKINVKLGGINNILVPHQRPS---VFQQPVIFLGADVTHPPAGDGKKPSIAAVVGSMD- 620
Query: 693 PLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRD 752
S+Y A VR Q + E+I +L ++RELLI FY S+ KP II +RD
Sbjct: 621 AHPSRYCATVRVQRPRQEIIQDL---------ASMVRELLIQFYKST-RFKPTRIIFYRD 670
Query: 753 GVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDN 806
GVSE QF QVL EL I EAC L++ + P IV QK HHT+ F + G N
Sbjct: 671 GVSEGQFRQVLYYELLAIREACISLEKDYQPGITYIVVQKRHHTRLFCADRTERVGRSGN 730
Query: 807 VPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYV 866
+P GT VD I HP +DFY+C+HAG+ GTSRP+HYHVL D+ F+ D+LQ L + L +
Sbjct: 731 IPAGTTVDTDITHPYEFDFYLCSHAGIQGTSRPSHYHVLWDDNFFTADELQLLTYQLCHT 790
Query: 867 YQRSTTAISVVAPICYAHLAASQVGQFM--KFEDKSETSSSHGGS-GRDINASPIPQLPK 923
Y R T ++S+ AP YAHL A + + K D +E S G S GRD A + + +
Sbjct: 791 YVRCTRSVSIPAPAYYAHLVAFRARYHLVDKEHDSAEGSHVSGQSNGRDPQA--LAKAVQ 848
Query: 924 LMDSVCNSMFF 934
+ +M+F
Sbjct: 849 IHQDTLRTMYF 859
>I2C3_HUMAN (Q9H9G7) Eukaryotic translation initiation factor 2C 3
(eIF2C 3) (eIF-2C 3) (Argonaute 3)
Length = 860
Score = 362 bits (928), Expect = 3e-99
Identities = 282/929 (30%), Positives = 441/929 (47%), Gaps = 116/929 (12%)
Query: 45 PADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVA 104
PA +P+ + P+ R G G+ G + LL N F+V + D Y ++ +
Sbjct: 8 PAGAQPLLMVPR-------------RPGYGAMGKPIKLLANCFQVEIPKIDVYLYEVDIK 54
Query: 105 LFYEDGRPVEGKGAGRKILDRVQETYGSELNG-KDLAYDGEKTLFTIGSLAQNKLEFTVV 163
D P + R+++D + + + + G + YDG+++L+T L + T V
Sbjct: 55 ---PDKCP---RRVNREVVDSMVQHFKVTIFGDRRPVYDGKRSLYTANPLP---VATTGV 105
Query: 164 LEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGH 223
DVT G P +KV I F S++ + L G
Sbjct: 106 DLDVTLPGEGGKDRP----------------------FKVSIKFVSRVSWHLLHEVLTGR 143
Query: 224 ETENYQEA--------IRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRG 275
E + +D++LR H V ++FF +GGG G
Sbjct: 144 TLPEPLELDKPISTNPVHAVDVVLR-HLPSMKYTPVGRSFFSAPEGYDHPLGGGREVWFG 202
Query: 276 LHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFL-----IANQNVRDPFSLDWNKAKRT-- 328
H S R + LNIDVS T PV+ F+ I N + + D ++ K T
Sbjct: 203 FHQSVRPAMWKMMLNIDVSATAFYKAQPVIQFMCEVLDIHNIDEQPRPLTDSHRVKFTKE 262
Query: 329 LKNLRITTS---PTNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRR 385
+K L++ + ++Y++ ++ P Q F L+ T E TV YF +
Sbjct: 263 IKGLKVEVTHCGTMRRKYRVCNVTRRPASHQTFPLQLENG-----QTVERTVAQYFREKY 317
Query: 386 KISLQYSADLPCINVGKPKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQER 445
+ L+Y LPC+ VG+ ++ T++P+E+C++V+ QR K L+ Q S++++ + + +R
Sbjct: 318 TLQLKYP-HLPCLQVGQEQKHTYLPLEVCNIVAGQRCIKKLTDNQTSTMIKATARSAPDR 376
Query: 446 MRVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRLKFG--NGEDFNPRNGRW 503
++ +++++Y ++P ++ + V GRVL AP L++G N P +G W
Sbjct: 377 QEEISRLVRSANYETDPFVQEFQFKVRDEMAHVTGRVLPAPMLQYGGRNRTVATPSHGVW 436
Query: 504 NFNNKKIVQPVKIEKWAVVNFSARCDVR-----GLVRDLIKCGGMKGIHVE-QPFDCFEE 557
+ K+ V+I+ WA+ F+ + R G L K G+ ++ QP C
Sbjct: 437 DMRGKQFHTGVEIKMWAIACFATQRQCREEILKGFTDQLRKISKDAGMPIQGQPCFC--- 493
Query: 558 NGQFRRAPPLVRVEKMFEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQ 617
+ A VE MF H+++ G ++ L + + +Y K+ G+ TQ
Sbjct: 494 ----KYAQGADSVEPMFRHLKNTYSGLQLIIVILPGK---TPVYAEVKRVGDTLLGMATQ 546
Query: 618 CIAPTRV---NDQYLTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGS 674
C+ V + Q L+N+ LKIN KLGG+N++L PS V + P + LG DV+H
Sbjct: 547 CVQVKNVIKTSPQTLSNLCLKINVKLGGINNILVPHQRPS---VFQQPVIFLGADVTHPP 603
Query: 675 PGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLID 734
G + PSIAAVV S S+Y A VR Q + E+I +L ++RELLI
Sbjct: 604 AGDGKKPSIAAVVGSMD-AHPSRYCATVRVQRPRQEIIQDL---------ASMVRELLIQ 653
Query: 735 FYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNH 794
FY S+ KP II +RDGVSE QF QVL EL I EAC L++ + P IV QK H
Sbjct: 654 FYKST-RFKPTRIIFYRDGVSEGQFRQVLYYELLAIREACISLEKDYQPGITYIVVQKRH 712
Query: 795 HTKFF------QPGSPDNVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDE 848
HT+ F + G N+P GT VD I HP +DFY+C+HAG+ GTSRP+HYHVL D+
Sbjct: 713 HTRLFCADRTERVGRSGNIPAGTTVDTDITHPYEFDFYLCSHAGIQGTSRPSHYHVLWDD 772
Query: 849 IGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQFM--KFEDKSETSSSH 906
F+ D+LQ L + L + Y R T ++S+ AP YAHL A + + K D +E S
Sbjct: 773 NCFTADELQLLTYQLCHTYVRCTRSVSIPAPAYYAHLVAFRARYHLVDKEHDSAEGSHVS 832
Query: 907 GGS-GRDINASPIPQLPKLMDSVCNSMFF 934
G S GRD A + + ++ +M+F
Sbjct: 833 GQSNGRDPQA--LAKAVQIHQDTLRTMYF 859
>I2C1_MOUSE (Q8CJG1) Eukaryotic translation initiation factor 2C 1
(eIF2C 1) (eIF-2C 1) (Piwi/argonaute family protain
meIF2C1)
Length = 857
Score = 357 bits (915), Expect = 1e-97
Identities = 263/879 (29%), Positives = 433/879 (48%), Gaps = 98/879 (11%)
Query: 70 RRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVALFYEDGRPVEGKGAGRKILDRVQET 129
R G+G+ G + LL N+F+V++ D Y ++ + D P + R++++ + +
Sbjct: 26 RPGIGTVGKPIKLLANYFEVDIPKIDVYHYEVDIK---PDKCP---RRVNREVVEYMVQH 79
Query: 130 YGSELNG-KDLAYDGEKTLFTIGSL--AQNKLEFTVVLEDVTSNRNNGNASPDGHGSPND 186
+ ++ G + YDG++ ++T+ +L +++F V + G G
Sbjct: 80 FKPQIFGDRKPVYDGKENIYTVTALPIGNERVDFEVTIP--------------GEG---- 121
Query: 187 TDRKRLKKSHRSKTYKVEISFASKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQ 246
+ + +KV I + + + + + AL + E+++ LD+ +R H A
Sbjct: 122 ----------KDRIFKVSIKWLAIVSWRMLHEALVSGQIPVPLESVQALDVAMR-HLASM 170
Query: 247 GCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVD 306
V ++FF + +GGG G H S R + LNIDVS T PV++
Sbjct: 171 RYTPVGRSFFSPPEGYYHPLGGGREVWFGFHQSVRPAMWKMMLNIDVSATAFYKAQPVIE 230
Query: 307 FLIANQNVRD----PFSLDWNKAKR---TLKNLRITTSPTNQ---EYKITGLSEMPCKDQ 356
F+ ++R+ P L ++ R +K L++ + Q +Y++ ++ P Q
Sbjct: 231 FMCEVLDIRNIDEQPKPLTDSQRVRFTKEIKGLKVEVTHCGQMKRKYRVCNVTRRPASHQ 290
Query: 357 LFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSL 416
F L+ T E TV +F + + L+Y LPC+ VG+ ++ T++P+E+C++
Sbjct: 291 TFPLQLESG-----QTVECTVAQHFKQKYNLQLKYP-HLPCLQVGQEQKHTYLPLEVCNI 344
Query: 417 VSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFT 476
V+ QR K L+ Q S++++ + + +R ++ +K + +P ++ GI + T
Sbjct: 345 VAGQRCIKKLTDNQTSTMIKATARSAPDRQEEISRLMKNASCNLDPYIQEFGIKVKDDMT 404
Query: 477 QVDGRVLQAPRLKFG--NGEDFNPRNGRWNFNNKKIVQPVKIEKWAVVNFSARCDVR--- 531
+V GRVL AP L++G N P G W+ K+ ++I+ WA+ F+ + R
Sbjct: 405 EVTGRVLPAPILQYGGRNRAIATPNQGVWDMRGKQFYNGIEIKVWAIACFAPQKQCREEV 464
Query: 532 --GLVRDLIKCGGMKGIHVE-QPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGAPKFL 588
L K G+ ++ QP C + A VE MF H+++ G +
Sbjct: 465 LKNFTDQLRKISKDAGMPIQGQPCFC-------KYAQGADSVEPMFRHLKNTYSGLQLII 517
Query: 589 LCLLSERKNSDLYGPWKKKNLAEFGIVTQCIA---PTRVNDQYLTNVLLKINAKLGGMNS 645
+ L + + +Y K+ G+ TQC+ + + Q L+N+ LKIN KLGG+N+
Sbjct: 518 VILPGK---TPVYAEVKRVGDTLLGMATQCVQVKNAVKTSPQTLSNLCLKINVKLGGINN 574
Query: 646 LLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQ 705
+L V H S V + P + LG DV+H G + PSI AVV S S+Y A VR Q
Sbjct: 575 IL-VPHQRSA--VFQQPVIFLGADVTHPPAGDGKKPSITAVVGSMD-AHPSRYCATVRVQ 630
Query: 706 GAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNI 765
+ E+I++L ++RELLI FY S+ KP II +RDGV E Q Q+L+
Sbjct: 631 RPRQEIIEDL---------SYMVRELLIQFYKST-RFKPTRIIFYRDGVPEGQLPQILHY 680
Query: 766 ELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPGSPDNVPPGTVVDNKICH 819
EL I +AC L++ + P IV QK HHT+ F + G N+P GT VD I H
Sbjct: 681 ELLAIRDACIKLEKDYQPGITYIVVQKRHHTRLFCADKNERIGKSGNIPAGTTVDTNITH 740
Query: 820 PRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAP 879
P +DFY+C+HAG+ GTSRP+HY+VL D+ F+ D+LQ L + L + Y R T ++S+ AP
Sbjct: 741 PFEFDFYLCSHAGIQGTSRPSHYYVLWDDNRFTADELQILTYQLCHTYVRCTRSVSIPAP 800
Query: 880 ICYAHLAASQVGQFM--KFEDKSETSSSHGGS-GRDINA 915
YA L A + + K D E S G S GRD A
Sbjct: 801 AYYARLVAFRARYHLVDKEHDSGEGSHISGQSNGRDPQA 839
>AGO1_SCHPO (O74957) Cell cycle control protein ago1 (RNA
interference pathway protein ago1)
Length = 834
Score = 318 bits (816), Expect = 3e-86
Identities = 232/741 (31%), Positives = 367/741 (49%), Gaps = 70/741 (9%)
Query: 183 SPNDTDRKRLKKSHRSKTYKVEISFA----SKIPLQAIANALKGHETENYQ--EAIRVLD 236
S D +K + S+++ EI F+ SKI L ++ + + + Q +I LD
Sbjct: 87 SKGDIADGTIKVNIGSESHPREIEFSIQKSSKINLHTLSQFVNSKYSSDPQVLSSIMFLD 146
Query: 237 IILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTT 296
++L++ ++ +FF + N +GGGV +G + S R Q +S+N+D+S++
Sbjct: 147 LLLKKKPSET-LFGFMHSFFTGE--NGVSLGGGVEAWKGFYQSIRPNQGFMSVNVDISSS 203
Query: 297 MIVHPGPVVDFLIAN---QNVRDPFSLDWNKAKRTLKNLRIT--------TSPTNQEYKI 345
++ L+ NVRD D + R + L++T T N+ Y I
Sbjct: 204 AFWRNDSLLQILMEYTDCSNVRDLTRFDLKRLSRKFRFLKVTCQHRNNVGTDLANRVYSI 263
Query: 346 TGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKR 405
G S D F + G + ++I+V +YF+ + LQY +LPCI V K
Sbjct: 264 EGFSSKSASDSFFVRRLNG------EEQKISVAEYFLENHNVRLQYP-NLPCILV---KN 313
Query: 406 PTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLR 465
+P+E C +V QRYT L++ Q ++++ + Q+P ER++ + D + D+ ++P L
Sbjct: 314 GAMLPIEFCFVVKGQRYTAKLNSDQTANMIRFAVQRPFERVQQIDDFVHQMDWDTDPYLT 373
Query: 466 NCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQPVK--IEKWAVVN 523
G+ I +V RVL+ P +++G P +GRWN K+ + P + I WAV+
Sbjct: 374 QYGMKIQKKMLEVPARVLETPSIRYGGDCIERPVSGRWNLRGKRFLDPPRAPIRSWAVMC 433
Query: 524 FSA--RCDVRGLVRDL------IKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFE 575
F++ R +RG+ L + G+ + + P + G + + K E
Sbjct: 434 FTSTRRLPMRGIENFLQTYVQTLTSLGINFVMKKPPVLYADIRGSVEEL--CITLYKKAE 491
Query: 576 HVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV---NDQYLTNV 632
V + AP L + ++ + + YG K+ G+ +QC + QY N+
Sbjct: 492 QVGN----APPDYLFFILDKNSPEPYGSIKRVCNTMLGVPSQCAISKHILQSKPQYCANL 547
Query: 633 LLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQW 692
+KIN K+GG+N L + +P + PTLILG DV H G T + SIA++V+S
Sbjct: 548 GMKINVKVGGINCSLIPKSNP----LGNVPTLILGGDVYHPGVGATGV-SIASIVASVDL 602
Query: 693 PLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRD 752
KY A R+Q E+I+ + + I+ LL F + ++P II FRD
Sbjct: 603 NGC-KYTAVSRSQPRHQEVIEGM---------KDIVVYLLQGF-RAMTKQQPQRIIYFRD 651
Query: 753 GVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSPD-----NV 807
G SE QF V+N ELSQI EAC L K+NPK LV QK HH +FF D N
Sbjct: 652 GTSEGQFLSVINDELSQIKEACHSLSPKYNPKILVCTTQKRHHARFFIKNKSDGDRNGNP 711
Query: 808 PPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVY 867
PGT+++ + HP YDFY+ +H + G S P HY VL DEI PD Q L ++L YVY
Sbjct: 712 LPGTIIEKHVTHPYQYDFYLISHPSLQGVSVPVHYTVLHDEIQMPPDQFQTLCYNLCYVY 771
Query: 868 QRSTTAISVVAPICYAHLAAS 888
R+T+A+S+V P+ YAHL ++
Sbjct: 772 ARATSAVSLVPPVYYAHLVSN 792
>YO43_CAEEL (P34681) Hypothetical protein ZK757.3 in chromosome III
Length = 1040
Score = 258 bits (659), Expect = 5e-68
Identities = 175/567 (30%), Positives = 290/567 (50%), Gaps = 45/567 (7%)
Query: 343 YKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGK 402
YK+ L ++P +F + E +V DYF + + L+Y LPC++VG
Sbjct: 419 YKVNSL-QLPADKLMFQ-----GIDEEGRQVVCSVADYF-SEKYGPLKYPK-LPCLHVGP 470
Query: 403 PKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEP 462
P R F+P+E C + S Q+Y K +S Q S++++ + +R + + +G++P
Sbjct: 471 PTRNIFLPMEHCLIDSPQKYNKKMSEKQTSAIIKAAAVDATQREDRIKQLAAQASFGTDP 530
Query: 463 MLRNCGISITSGFTQVDGRVLQAPRLKFG-NGEDFNP----RNGRWNFNNKKIVQPVKIE 517
L+ G++++S Q RV+Q P + FG N NP ++G W +N+ + P
Sbjct: 531 FLKEFGVAVSSQMIQTTARVIQPPPIMFGGNNRSVNPVVFPKDGSWTMDNQTLYMPATCR 590
Query: 518 KWAVVNFSARCDVRGLVRDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHV 577
++++ D L + + MK + F + + ++ R+ V +F +
Sbjct: 591 SYSMIALVDPRDQTSL-QTFCQSLTMKATAMGMNFPRWPDLVKYGRSKEDVCT--LFTEI 647
Query: 578 QSKLPGAPKFLLCLLS--ERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVN---DQYLTNV 632
+ C++ + KNSD+Y K+++ GI++QC+ V+ N+
Sbjct: 648 ADEYRVTNTVCDCIIVVLQSKNSDIYMTVKEQSDIVHGIMSQCVLMKNVSRPTPATCANI 707
Query: 633 LLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEI----PSIAAVVS 688
+LK+N K+GG+NS + + + +V + PT+++G+DV+H P Q E+ PS+AA+V+
Sbjct: 708 VLKLNMKMGGINSRIVADKITNKYLVDQ-PTMVVGIDVTH--PTQAEMRMNMPSVAAIVA 764
Query: 689 SRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNII 748
+ L Y A V+ Q E + L IRE +I FY + +KP II
Sbjct: 765 NVDL-LPQSYGANVKVQKKCRESVVYLLDA---------IRERIITFYRHT-KQKPARII 813
Query: 749 IFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFF------QPG 802
++RDGVSE QF++VL E+ I AC + E + P IV QK HH + F G
Sbjct: 814 VYRDGVSEGQFSEVLREEIQSIRTACLAIAEDFRPPITYIVVQKRHHARIFCKYQNDMVG 873
Query: 803 SPDNVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHS 862
NVPPGT VD I P +DFY+C+H G+ GTSRP YHVLLDE F+ D++Q + +
Sbjct: 874 KAKNVPPGTTVDTGIVSPEGFDFYLCSHYGVQGTSRPARYHVLLDECKFTADEIQSITYG 933
Query: 863 LSYVYQRSTTAISVVAPICYAHLAASQ 889
+ + Y R T ++S+ P+ YA L A++
Sbjct: 934 MCHTYGRCTRSVSIPTPVYYADLVATR 960
Score = 45.8 bits (107), Expect = 5e-04
Identities = 52/199 (26%), Positives = 84/199 (42%), Gaps = 29/199 (14%)
Query: 45 PADIEPIKIEPQIVKKKLPTKVPMARRGLGSKGAKLPLLTNHFKVNVTNTDGYFFQYSVA 104
P D+ P+ + L + +AR GLG+ G ++P+ +N F +++ N QY V
Sbjct: 67 PRDLSPLLLSELAC---LNMREVVARPGLGTIGRQIPVKSNFFAMDLKNPKMVVIQYHVE 123
Query: 105 LFYEDGRPVEGKGAGRKILDRVQETYGSELNGK-DLAYDGEKTLFTIGSLAQNKLEFTVV 163
+ + R ++ K R I + + + + K LAYDG L+T+ +LEF
Sbjct: 124 IHHPGCRKLD-KDEMRIIFWKAVSDHPNIFHNKFALAYDGAHQLYTVA-----RLEF--- 174
Query: 164 LEDVTSNRNNGNASPDGHGSPNDTDRKRLKKSHRSKTYKVEISFASKIP-LQAIANALKG 222
PD GS L K +R +T + IS + P L +
Sbjct: 175 --------------PDDQGSVRLDCEASLPKDNRDRT-RCAISIQNVGPVLLEMQRTRTN 219
Query: 223 HETENYQEAIRVLDIILRQ 241
+ E I++LDII RQ
Sbjct: 220 NLDERVLTPIQILDIICRQ 238
>AGO2_DROME (Q9VUQ5) Argonaute 2 protein
Length = 1214
Score = 251 bits (642), Expect = 5e-66
Identities = 204/786 (25%), Positives = 355/786 (44%), Gaps = 95/786 (12%)
Query: 133 ELNGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRL 192
+L G LAYDG+ + +++ L N +VT NG
Sbjct: 459 QLGGAVLAYDGKASCYSVDKLPLNSQN-----PEVTVTDRNG------------------ 495
Query: 193 KKSHRSKTYKVEISFA--SKIPLQAIANALKGHETENYQEAIRVLDIILRQHAAKQGCLL 250
R+ Y +EI S I L+++ + + A++ ++++L + +
Sbjct: 496 ----RTLRYTIEIKETGDSTIDLKSLTTYMNDRIFDKPMRAMQCVEVVLASPCHNKAIRV 551
Query: 251 VRQNFFHNDPKNFTDVGGGVLGCRGLHSSFRTTQSGLSLNIDVSTTMIVHPGPVVDFL-- 308
R F +DP N ++ G GL+ +F LN+D+S P++++L
Sbjct: 552 GRSFFKMSDPNNRHELDDGYEALVGLYQAFMLGDRPF-LNVDISHKSFPISMPMIEYLER 610
Query: 309 -IANQNVRDPFSLDWNKA--KRTLKNLRITTSPTN------QEYKITGLSEMPCKDQLFT 359
+ + +LD+++ + L+ + + +P + Y++ GLS P + F
Sbjct: 611 FSLKAKINNTTNLDYSRRFLEPFLRGINVVYTPPQSFQSAPRVYRVNGLSRAPASSETF- 669
Query: 360 LKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKRPTFVPVELCSLVSL 419
E D +++T+ YF + R L++ L C+NVG + +P+ELCS+
Sbjct: 670 ---------EHDGKKVTIASYF-HSRNYPLKFP-QLHCLNVGSSIKSILLPIELCSIEEG 718
Query: 420 QRYTKALSTLQRSSLVEKSRQKPQERMRVLTDALKTSDYGSEPMLRNCGISITSGFTQVD 479
Q + Q +++++ + R R + + L+ + +P + GI I + F V
Sbjct: 719 QALNRKDGATQVANMIKYAATSTNVRKRKIMNLLQYFQHNLDPTISRFGIRIANDFIVVS 778
Query: 480 GRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQPV-KIEKWAVVNFSARCDVRGLVRDLI 538
RVL P++++ + +NG W + K ++P K K AV+ R + L
Sbjct: 779 TRVLSPPQVEYHSKRFTMVKNGSWRMDGMKFLEPKPKAHKCAVLYCDPRSGRKMNYTQLN 838
Query: 539 KCGGM-----KGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGAPKFLLCLLS 593
G + K +++ D P E+ + + + L + L ++
Sbjct: 839 DFGNLIISQGKAVNISLDSDVTYR--------PFTDDERSLDTIFADLKRSQHDLAIVII 890
Query: 594 ERKNSDLYGPWKKKNLAEFGIVTQCIAPTRV----NDQYLTNVLLKINAKLGGMNSLLGV 649
+ Y K+K + GI+TQCI V N+Q + N+LLKIN+KL G+N +
Sbjct: 891 PQFRIS-YDTIKQKAELQHGILTQCIKQFTVERKCNNQTIGNILLKINSKLNGINHK--I 947
Query: 650 EHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKV 709
+ P +P++ T+ +G DV+H SP Q EIPS+ V +S P + Y R Q +
Sbjct: 948 KDDPRLPMMKN--TMYIGADVTHPSPDQREIPSVVGVAASHD-PYGASYNMQYRLQRGAL 1004
Query: 710 EMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQ 769
E I+++F + + Y N PD+II +RDGVS+ QF ++ N EL
Sbjct: 1005 EEIEDMFSITLEH----------LRVYKEYRNAYPDHIIYYRDGVSDGQFPKIKNEELRC 1054
Query: 770 IIEACKFLDEKWNPKFLVIVAQKNHHTKFFQPGSP------DNVPPGTVVDNKICHPRNY 823
I +AC + K PK ++ K HHT+FF G +NV PGTVVD I HP
Sbjct: 1055 IKQACDKVGCK--PKICCVIVVKRHHTRFFPSGDVTTSNKFNNVDPGTVVDRTIVHPNEM 1112
Query: 824 DFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYA 883
F+M +H + GT++PT Y+V+ + D LQ+L ++L +++ R ++S AP A
Sbjct: 1113 QFFMVSHQAIQGTAKPTRYNVIENTGNLDIDLLQQLTYNLCHMFPRCNRSVSYPAPAYLA 1172
Query: 884 HLAASQ 889
HL A++
Sbjct: 1173 HLVAAR 1178
>PIWI_BRARE (Q8UVX0) Piwi protein
Length = 858
Score = 158 bits (400), Expect = 6e-38
Identities = 150/589 (25%), Positives = 261/589 (43%), Gaps = 69/589 (11%)
Query: 327 RTLKNLRITTSPTNQEYKITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRK 386
+ L L I T N+ Y+I ++ + F K+G DTE I+ +YF ++
Sbjct: 299 KELVGLIILTKYNNKTYRIDDIAWDHTPNNTF---KKG------DTE-ISFKNYFKSQYG 348
Query: 387 ISLQYSADLPCINVGK--------PKRPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKS 438
+ + + ++ K P P + E C L L +A + + L +
Sbjct: 349 LDITDGNQVLLVSHVKRLGPSGRPPPGPAMLVPEFCYLTGLTDKMRADFNIMKD-LASHT 407
Query: 439 RQKPQERM----RVLTDALKTSDYGSEPMLRNCGISITSGFTQVDGRVLQAPRL-KFGNG 493
R P++R R++++ + D +E L G+S + ++GRVL + R+ + G
Sbjct: 408 RLSPEQREGRINRLISNINRNGDVQNE--LTTWGLSFENKLLSLNGRVLPSERIIQGGRA 465
Query: 494 EDFNPRNGRWN--FNNKKIVQPVKIEKWAVVNFSARCDV-RGLVRDLIKCGGMKGIHVEQ 550
++NP W+ ++ + ++ W + DV + L++ L K G GI +++
Sbjct: 466 FEYNPWTADWSKEMRGLPLISCMSLDNWLMFYTRRNADVAQSLLQTLNKVSGPMGIRMQR 525
Query: 551 PFDCFEENGQFRRAPPLVRVEKMFEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLA 610
E+ Q E + +Q + + ++ +L + D Y KK
Sbjct: 526 AVMIEYEDRQ----------ESLLRALQQNVARETQMVVVILPTNRK-DKYDCVKKYLCV 574
Query: 611 EFGIVTQCIAPTRVNDQYL-----TNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLI 665
+ +QC+ ++ T + L++N K+GG L VE IP+ +I
Sbjct: 575 DCPTPSQCVVSRTISKPQALMTVATKIALQMNCKMGG--ELWSVE----IPL---RQLMI 625
Query: 666 LGMDVSHGSPGQTEIPSIAAVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDE 725
+G+D H + SI A+V+S + + CV Q E+ID L +
Sbjct: 626 VGIDCYHDTAAGKR--SIGAMVASLNQGMSRWFSKCV-LQNRGQEIIDAL---------K 673
Query: 726 GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKF 785
G ++ L Y N P II++RDGV + V++ E+ QI+++ K + + + PK
Sbjct: 674 GSLQGAL-KAYLKYNNSLPSRIIVYRDGVGDGMLQSVVDYEVPQIMQSIKTMGQDYEPKL 732
Query: 786 LVIVAQKNHHTKFFQ--PGSPDNVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYH 843
V+V +K ++FF G N PPGTV+D ++ P YDF++ + A G PTHY+
Sbjct: 733 SVVVVKKRISSRFFARIDGKIANPPPGTVIDTEVTRPEWYDFFIVSQAVRFGCVAPTHYN 792
Query: 844 VLLDEIGFSPDDLQELVHSLSYVYQRSTTAISVVAPICYAHLAASQVGQ 892
V+ D G PD +Q L + L ++Y + V AP YAH A VGQ
Sbjct: 793 VVFDNSGLKPDHMQRLTYKLCHMYYNWQGIVRVPAPCQYAHKLAFLVGQ 841
>PIWI_DROME (Q9VKM1) Piwi protein
Length = 843
Score = 140 bits (352), Expect = 2e-32
Identities = 156/682 (22%), Positives = 300/682 (43%), Gaps = 72/682 (10%)
Query: 232 IRVLDIILRQHAAKQGCLLVRQNFFHNDPKNFTDVGGGVLGC-RGLHSSFRTTQSGLSLN 290
++VL++ILR+ LV +N F DP+ ++ + G +S R + + L
Sbjct: 194 LQVLNLILRRSMKGLNLELVGRNLF--DPRAKIEIREFKMELWPGYETSIRQHEKDILLG 251
Query: 291 IDVSTTMIVHPGPVVDFLIANQNVRDPFSLDWNKAKRTLKNLRITTSPTNQEYKITGLSE 350
++ T ++ + D I + +P + ++ + + +L + T N+ Y+I +
Sbjct: 252 TEI-THKVMRTETIYD--IMRRCSHNP-ARHQDEVRVNVLDLIVLTDYNNRTYRINDVDF 307
Query: 351 MPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKPKR----- 405
F+ K R +I+ +Y++ + I ++ I+ + K
Sbjct: 308 GQTPKSTFSCKGR----------DISFVEYYLTKYNIRIRDHNQPLLISKNRDKALKTNA 357
Query: 406 ---PTFVPVELCSLVSLQRYTKALSTLQR--SSLVEKSRQKPQERMRVLTDALKTSDYGS 460
+P ELC + L ++ L R SS + ++ +R+R L+ + S
Sbjct: 358 SELVVLIP-ELCRVTGLNAEMRSNFQLMRAMSSYTRMNPKQRTDRLRAFNHRLQNTPE-S 415
Query: 461 EPMLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRW--NFNNKKIVQPVK--I 516
+LR+ + + T+V GR++ + F NG+ N W +F +++++ +
Sbjct: 416 VKVLRDWNMELDKNVTEVQGRIIGQQNIVFHNGKVPAGENADWQRHFRDQRMLTTPSDGL 475
Query: 517 EKWAVVNFSARC-DVRGLVRDLIKCGGMKGIHVEQPFDCF---EENGQFRRAPPLVRVEK 572
++WAV+ ++R L+ L + G+ + P + + G + RA
Sbjct: 476 DRWAVIAPQRNSHELRTLLDSLYRAASGMGLRIRSPQEFIIYDDRTGTYVRA-------- 527
Query: 573 MFEHVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVNDQYLTNV 632
M + V+S PK +LCL+ N++ Y KK+ + + TQ + ++ L ++
Sbjct: 528 MDDCVRSD----PKLILCLVPN-DNAERYSSIKKRGYVDRAVPTQVVTLKTTKNRSLMSI 582
Query: 633 LLKINAKLGGMNSLLGVEHSPSIPIVSKAPTLILGMDVSHGSPGQTEIPSIAAVVSSRQW 692
KI +L N LG ++P + + + + +G D++ + + + A+++S
Sbjct: 583 ATKIAIQL---NCKLG--YTPWMIELPLSGLMTIGFDIAKSTRDRKR--AYGALIASMDL 635
Query: 693 PLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELLIDFYNSSGNRKPDNIIIFRD 752
S Y + V T+ + +++ N P+ I + L Y + P I+ +RD
Sbjct: 636 QQNSTYFSTV-TECSAFDVLANTLWPM--------IAKALRQ-YQHEHRKLPSRIVFYRD 685
Query: 753 GVSESQFNQVLNIELSQIIEACKFLDEKWN---PKFLVIVAQKNHHTKFFQPGSPDNVPP 809
GVS Q+ E+ IIE K + P+ IV ++ +T+FF G N PP
Sbjct: 686 GVSSGSLKQLFEFEVKDIIEKLKTEYARVQLSPPQLAYIVVTRSMNTRFFLNGQ--NPPP 743
Query: 810 GTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDDLQELVHSLSYVYQR 869
GT+VD+ I P YDFY+ + GT PT Y+VL +G SP+ +Q+L + + ++Y
Sbjct: 744 GTIVDDVITLPERYDFYLVSQQVRQGTVSPTSYNVLYSSMGLSPEKMQKLTYKMCHLYYN 803
Query: 870 STTAISVVAPICYAHLAASQVG 891
+ V A YA A+ VG
Sbjct: 804 WSGTTRVPAVCQYAKKLATLVG 825
>GCC7_CAEEL (Q21770) Germ cell expressed protein R06C7.1
Length = 945
Score = 94.4 bits (233), Expect = 1e-18
Identities = 183/809 (22%), Positives = 313/809 (38%), Gaps = 120/809 (14%)
Query: 135 NGKDLAYDGEKTLFTIGSLAQNKLEFTVVLEDVTSNRNNGNASPDGHGSPNDTDRKRLKK 194
+G + YDG+ TL+T +L ++ +N + D K L
Sbjct: 148 DGNQIVYDGQSTLYTTVNL----------FSELDANGTKSKVFQINGADTGNDDLKTLP- 196
Query: 195 SHRSKTYKVEISFASKIPLQAIANALKGHETE------NYQEAIRVLDIILRQHAAKQ-- 246
+EI +A + +++ G T N +E + L++ L QH ++
Sbjct: 197 -----CISLEI-YAPRDNSITLSSENLGKRTADQNIEVNNREYTQFLELALNQHCVRETN 250
Query: 247 --GCLLVRQNFFHN------DPKNFTDVGGGVLGCRGLHSSFRTT-------QSGLSLNI 291
GC + +F N D ++ DVG G GL + + Q+ SL I
Sbjct: 251 RFGCFEHGKVYFLNATEEGFDQRDCVDVGDGKQLYPGLKKTIQFIEGPYGRGQNNPSLVI 310
Query: 292 DVSTTMIVHPGPVVD--FLIANQNVRDPFS-LDWNKAKRTLKNLRITTSPTNQE--YKIT 346
D V+ F I Q+ + + + KA +K L ++ TN++ +I
Sbjct: 311 DGMKAAFHKEQTVIQKLFDITGQDPSNGLNNMTREKAAAVIKGLDCYSTYTNRKRHLRIE 370
Query: 347 GLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQY-SADLP-CINVGKPK 404
G+ F L D + ++ +Y+ ++ KISLQY +A+L C + G
Sbjct: 371 GIFHESATKTRFELP---------DGKTCSIAEYYADKYKISLQYPNANLVVCKDRGNNN 421
Query: 405 RPTFVPVELCSLVSLQRYTKALSTLQRSSLVEKS-RQKPQERMRVLTDALKTSDYGSE-P 462
+ P EL ++ QR T T +S K P R R++ + E
Sbjct: 422 ---YFPAELMTVSRNQRVTIPQQTGNQSQKTTKECAVLPDVRQRMIITGKNAVNITLENE 478
Query: 463 MLRNCGISITSGFTQVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNKKIVQPVKIEK---- 518
+L GI + S V R L L + + G+W V+P +
Sbjct: 479 LLVALGIKVYSEPLMVQARELDGKELVYQRSVMSDM--GKWRAPPGWFVKPATVPDLWAA 536
Query: 519 WAVVNFSARC---DVRGLVRDLIKCGGMKGIHVEQPFDCFEENGQFRRAPPLVRVEKMFE 575
+AV N R DV LV I KG+ ++ P E G L EK+
Sbjct: 537 YAVGNPGCRFSIGDVNQLVGMFIDSCKKKGMVIKPPC----ETG-------LYSTEKIMT 585
Query: 576 HVQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVNDQY------- 628
++ K++L ++++ L+ +K IV Q + ++ N
Sbjct: 586 QLEKVAASKCKYVL-MITDDAIVHLHKQYKALEQRTMMIV-QDMKISKANAVVKDGKRLT 643
Query: 629 LTNVLLKINAKLGGMNSLLGVEHSPSIPIVSKAPT---LILGMDVSHGSPGQTEIPSIAA 685
L N++ K N KLGG+N ++ K+ T LI+G+ VS +P P+
Sbjct: 644 LENIINKTNVKLGGLNY--------TVSDAKKSMTDEQLIIGVGVS--AP-----PAGTK 688
Query: 686 VVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDEGIIRELL---IDFYNSSGNR 742
+ + L + A E + + S + I ++L ID + +
Sbjct: 689 YMMDNKGHLNPQIIGFASNAVANHEFVGDFVLAPSGQDTMASIEDVLQNSIDLFEKNRKA 748
Query: 743 KPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKFLVIVAQKNHHTKFFQP- 801
P III+R G SE +L E+ ++ K + IV K H +FF+
Sbjct: 749 LPKRIIIYRSGASEGSHASILAYEIPLARAIIHGYSKEI--KLIFIVVTKEHSYRFFRDQ 806
Query: 802 ------GSPDNVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRPTHYHVLLDEIGFSPDD 855
+ N+PPG V+DN + +P F++ H + GT++ Y VL D+ D
Sbjct: 807 LRSGGKATEMNIPPGIVLDNAVTNPACKQFFLNGHTTLQGTAKTPLYTVLADDCKAPMDR 866
Query: 856 LQELVHSLSYVYQRSTTAISVVAPICYAH 884
L+EL +L + +Q + + S+ P+ A+
Sbjct: 867 LEELTFTLCHHHQIVSLSTSIPTPLYVAN 895
>YQ53_CAEEL (Q09249) Hypothetical protein C16C10.3 in chromosome III
Length = 1032
Score = 91.7 bits (226), Expect = 8e-18
Identities = 121/577 (20%), Positives = 239/577 (40%), Gaps = 76/577 (13%)
Query: 344 KITGLSEMPCKDQLFTLKKRGAVPGEDDTEEITVYDYFVNRRKISLQYSADLPCINVGKP 403
+++ ++E ++ F +K + E+TV +YF+ + I L+Y LP + +
Sbjct: 414 RVSSIAENNAENTSFMMKD------DKGEREVTVAEYFLLQYNIKLKYPR-LPLVVSKRF 466
Query: 404 KRPTFVPVELCSLVSLQRY-TKALSTLQRSSLVEKSRQKPQERMRVLTDALKTS-DYGSE 461
K +F P+EL + QR +S +S++ ++ PQ ++++ D L+ +
Sbjct: 467 KHESFFPMELLRIAPGQRIKVNKMSPTVQSAMTGRNASMPQHHVKLVQDILRDNLKLEQN 526
Query: 462 PMLRNCGISITSGFT-QVDGRVLQAPRLKFGNGEDFNPRNGRWNFNNK-KIVQPVKIEKW 519
+ GI + S Q+ ++L ++KF G+ + P R F + K V+P +I K
Sbjct: 527 KYMDAFGIKLMSTEPIQMTAKLLPPAQIKF-KGQTYMPDMSRPAFRTQDKFVEPARIRKI 585
Query: 520 AVVNFSARCDVRGLVRDLIKCGGM---KGIHVEQPFDCFEENGQFRRAPPLVRVEKMFEH 576
+V F +R K GI VE+ D + + + + V ++ + +
Sbjct: 586 GIVVFDNCIQMRQAEDFCDKLSNFCRDNGITVEK--DSRDWSIRELNSSDSVAIQNLMK- 642
Query: 577 VQSKLPGAPKFLLCLLSERKNSDLYGPWKKKNLAEFGIVTQCIAPTRVND--------QY 628
K +L ++ K D++ K G+ T + V+ Q
Sbjct: 643 ---KWLDDRVDILVGIAREKKPDVHDILKYFE-ESIGLQTIQLCQQTVDKMMGGQGGRQT 698
Query: 629 LTNVLLKINAKLGGMNSLLGVEHSP-SIPIVSKAPTL---------ILGMDVSHGSP--- 675
+ NV+ K N K GG N + + ++ + S TL +G ++SHG+
Sbjct: 699 IDNVMRKFNLKCGGTNFFVEIPNAVRGKAVCSNNETLRKKLLEHVQFIGFEISHGASRTL 758
Query: 676 ---------GQTEIPSIA-AVVSSRQWPLISKYRACVRTQGAKVEMIDNLFKPVSDTEDE 725
G+ + ++ ++ +S Q + + + K++ +D F
Sbjct: 759 FDRSRSQMDGEPSVVGVSYSLTNSTQ---LGGFTYLQTQKEYKLQKLDEFFPKC------ 809
Query: 726 GIIRELLIDFYNSSGNRKPDNIIIFRDGVSESQFNQVLNIELSQIIEACKFLDEKWNPKF 785
+ Y P I+I+R G E FN+V E+ ++ + + P
Sbjct: 810 -------VRSYKEHSKTLPTRIVIYRVGAGEGNFNRVKE-EVEEMRRTFDKIQPGYRPHL 861
Query: 786 LVIVAQKNHHTKFFQP------GSPDNVPPGTVVDNKICHPRNYDFYMCAHAGMIGTSRP 839
+VI+AQ+ H + F + N+P GT V+N + +F + + +IGT RP
Sbjct: 862 VVIIAQRASHARVFPSCISGNRATDQNIPSGTCVENVLTSYGYDEFILSSQTPLIGTVRP 921
Query: 840 THYHVLLDEIGFSPDDLQELVHSLSYVYQRSTTAISV 876
Y +L+++ +S ++L L + ++ +Q S SV
Sbjct: 922 CKYTILVNDAKWSKNELMHLTYFRAFGHQVSYQPPSV 958
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.137 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 115,402,976
Number of Sequences: 164201
Number of extensions: 5229364
Number of successful extensions: 13220
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 13036
Number of HSP's gapped (non-prelim): 47
length of query: 935
length of database: 59,974,054
effective HSP length: 120
effective length of query: 815
effective length of database: 40,269,934
effective search space: 32819996210
effective search space used: 32819996210
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC124972.13


