

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124970.4 + phase: 0 /pseudo
(204 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
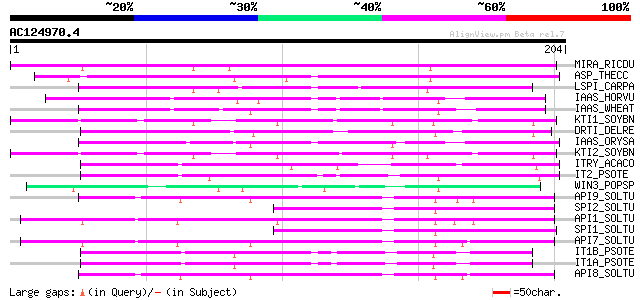
Score E
Sequences producing significant alignments: (bits) Value
MIRA_RICDU (P13087) Miraculin precursor (MIR) 152 5e-37
ASP_THECC (P32765) 21 kDa seed protein precursor 107 2e-23
LSPI_CARPA (P80691) Latex serine proteinase inhibitor 71 2e-12
IAAS_HORVU (P07596) Alpha-amylase/subtilisin inhibitor precursor... 65 1e-10
IAAS_WHEAT (P16347) Endogenous alpha-amylase/subtilisin inhibito... 64 2e-10
KTI1_SOYBN (P25272) Kunitz-type trypsin inhibitor KTI1 precursor 61 2e-09
DRTI_DELRE (P83667) Kunitz-type serine protease inhibitor DrTI 60 3e-09
IAAS_ORYSA (P29421) Alpha-amylase/subtilisin inhibitor (RASI) 58 1e-08
KTI2_SOYBN (P25273) Kunitz-type trypsin inhibitor KTI2 precursor 58 2e-08
ITRY_ACACO (P24924) Trypsin inhibitor precursor 55 9e-08
IT2_PSOTE (P25700) Trypsin inhibitor 2 (WTI-2) 55 1e-07
WIN3_POPSP (P16335) Wound responsive GWIN3 protein precursor 53 6e-07
API9_SOLTU (P58521) Aspartic protease inhibitor 9 (Novel inhibit... 52 7e-07
SPI2_SOLTU (P58515) Serine protease inhibitor 2 (PSPI-21) (PSPI-... 51 2e-06
API1_SOLTU (Q41480) Aspartic protease inhibitor 1 precursor (pA1... 51 2e-06
SPI1_SOLTU (P58514) Serine protease inhibitor 1 (PSPI-21) (PSPI-... 51 2e-06
API7_SOLTU (Q41448) Aspartic protease inhibitor 7 precursor (Cat... 51 2e-06
IT1B_PSOTE (P32877) Trypsin inhibitor 1B (WTI-1B) 50 3e-06
IT1A_PSOTE (P10821) Trypsin inhibitor 1A (WTI-1A) 50 3e-06
API8_SOLTU (P17979) Aspartic protease inhibitor 8 precursor (pi8... 50 5e-06
>MIRA_RICDU (P13087) Miraculin precursor (MIR)
Length = 220
Score = 152 bits (384), Expect = 5e-37
Identities = 87/210 (41%), Positives = 116/210 (54%), Gaps = 9/210 (4%)
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQ-PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMA 59
+ L F + LL L A+ P V D G +R NY+I+P G ++
Sbjct: 7 LSLSFFFVSALLAAAANPLLSAADSAPNPVLDIDGEKLRTGTNYYIVPVLRDHGGGLTVS 66
Query: 60 LLNTNKT--CPLDVVEEEEAMQ----FSFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSV 113
N T CP VV+ + + +F P N K+ V+RVSTDLN+ S C +S
Sbjct: 67 ATTPNGTFVCPPRVVQTRKEVDHDRPLAFFPENPKEDVVRVSTDLNINFSAFMPCRWTSS 126
Query: 114 TVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERF--ESGYKLVFCPTVCRECEV 171
TVW++DK D +T Q FVT GGV+GNPG ET+ +WFKIE F YKLVFCPTVC C+V
Sbjct: 127 TVWRLDKYDESTGQYFVTIGGVKGNPGPETISSWFKIEEFCGSGFYKLVFCPTVCGSCKV 186
Query: 172 VCKDIGIFLDENRNTRFVLSDFPFGVKFQR 201
C D+GI++D+ R LSD PF +F +
Sbjct: 187 KCGDVGIYIDQKGRRRLALSDKPFAFEFNK 216
>ASP_THECC (P32765) 21 kDa seed protein precursor
Length = 221
Score = 107 bits (266), Expect = 2e-23
Identities = 73/205 (35%), Positives = 105/205 (50%), Gaps = 16/205 (7%)
Query: 10 LLLTLFTTKPL-----QGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTN 64
LLL FT+K A P V DT G+ ++ + Y++L S G T
Sbjct: 9 LLLFAFTSKSYFFGVANAANSP--VLDTDGDELQTGVQYYVLSSISGAGGGGLALGRATG 66
Query: 65 KTCPLDVVEEEEAMQFS----FVPFNFKKGVIRVSTDLNV--IHSFPTNCSTSSVTVWKV 118
++CP VV+ + F + K V+RVSTD+N+ + CSTS TVW++
Sbjct: 67 QSCPEIVVQRRSDLDNGTPVIFSNADSKDDVVRVSTDVNIEFVPIRDRLCSTS--TVWRL 124
Query: 119 DKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERF-ESGYKLVFCPTVCRECEVVCKDIG 177
D D + + +VTT GV+G PG T+ +WFKIE+ GYK FCP+VC C +C DIG
Sbjct: 125 DNYDNSAGKWWVTTDGVKGEPGPNTLCSWFKIEKAGVLGYKFRFCPSVCDSCTTLCSDIG 184
Query: 178 IFLDENRNTRFVLSDFPFGVKFQRA 202
D++ R LSD + F++A
Sbjct: 185 RHSDDDGQIRLALSDNEWAWMFKKA 209
>LSPI_CARPA (P80691) Latex serine proteinase inhibitor
Length = 184
Score = 71.2 bits (173), Expect = 2e-12
Identities = 54/174 (31%), Positives = 84/174 (48%), Gaps = 11/174 (6%)
Query: 26 PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKT-CPLDVVEEE----EAMQF 80
P+ + D G V ++YF++ + G NK CPL VV++ E + F
Sbjct: 3 PKPIVDIDGKPVLYGVDYFVVSAIWGAGGGGLTVYGPGNKKKCPLSVVQDPFDNGEPIIF 62
Query: 81 SFVPFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPG 140
S + N K ++ S DLNV + NC+ + T WKVD+ VT GG +G G
Sbjct: 63 SAIK-NVKDNIVFESVDLNVKFNITINCNET--TAWKVDRFPGVIGWT-VTLGGEKGYHG 118
Query: 141 RETVDNWFKIER--FESGYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSD 192
E+ + FKI++ YK FCP+ R + C ++ IF D+ R R +L++
Sbjct: 119 FESTHSMFKIKKAGLPFSYKFHFCPSYPRTRLIPCNNVDIFFDKYRIRRLILTN 172
>IAAS_HORVU (P07596) Alpha-amylase/subtilisin inhibitor precursor
(BASI)
Length = 203
Score = 65.1 bits (157), Expect = 1e-10
Identities = 56/194 (28%), Positives = 87/194 (43%), Gaps = 22/194 (11%)
Query: 14 LFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVE 73
LF P A P V DT G+ +R NY++L ++ G MA + CPL V +
Sbjct: 13 LFWPAPPSRAADPPPVHDTDGHELRADANYYVLSANRAHGGGLTMA-PGHGRHCPLFVSQ 71
Query: 74 EEEAMQFSF----VPFNFKKG--VIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQ 127
+ F P+ +IR+STD+ + T C S T W +D ++A +
Sbjct: 72 DPNGQHDGFPVRITPYGVAPSDKIIRLSTDVRISFRAYTTCLQS--TEWHIDS-ELAAGR 128
Query: 128 RFVTTGGVQGNPGRETVDNWFKIERFESG----YKLVFCPTVCRECEVVCKDIGIFLDEN 183
R V TG V+ +P +N F+IE++ YKL+ C C+D+G+F D
Sbjct: 129 RHVITGPVK-DPSPSGRENAFRIEKYSGAEVHEYKLM-------SCGDWCQDLGVFRDLK 180
Query: 184 RNTRFVLSDFPFGV 197
F+ + P+ V
Sbjct: 181 GGAWFLGATEPYHV 194
>IAAS_WHEAT (P16347) Endogenous alpha-amylase/subtilisin inhibitor
(WASI)
Length = 180
Score = 63.9 bits (154), Expect = 2e-10
Identities = 54/183 (29%), Positives = 85/183 (45%), Gaps = 24/183 (13%)
Query: 26 PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVEEEEAMQFSFVPF 85
P V DT GN +R NY++LP++ G MA + CPL V +E + Q +P
Sbjct: 2 PPPVHDTDGNELRADANYYVLPANRAHGGGLTMA-PGHGRRCPLFVSQEADG-QRDGLPV 59
Query: 86 NF-------KKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGN 138
+IR+STD+ + T C S T W +D ++ + +R V TG V+ +
Sbjct: 60 RIAPHGGAPSDKIIRLSTDVRISFRAYTTCVQS--TEWHIDS-ELVSGRRHVITGPVR-D 115
Query: 139 PGRETVDNWFKIERFESG----YKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSDFP 194
P +N F+IE++ YKL+ C C+D+G+F D F+ + P
Sbjct: 116 PSPSGRENAFRIEKYSGAEVHEYKLMACGD-------SCQDLGVFRDLKGGAWFLGATEP 168
Query: 195 FGV 197
+ V
Sbjct: 169 YHV 171
>KTI1_SOYBN (P25272) Kunitz-type trypsin inhibitor KTI1 precursor
Length = 203
Score = 60.8 bits (146), Expect = 2e-09
Identities = 65/212 (30%), Positives = 95/212 (44%), Gaps = 26/212 (12%)
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
MK L+ FT L A + V DT + ++N Y++LP + G + +
Sbjct: 1 MKSTIFFALFLVCAFTISYLPSATA-QFVLDTDDDPLQNGGTYYMLP--VMRGKGGGIEV 57
Query: 61 LNTNKT-CPLDVVEEEEAMQFSFVPFNFKKGVIRVSTD----LNVIHSFPTNCSTSSVTV 115
+T K CPL VV+ P KG+ V T L + +P + S V
Sbjct: 58 DSTGKEICPLTVVQS---------PNELDKGIGLVFTSPLHALFIAERYPLSIKFGSFAV 108
Query: 116 WKVDKVDVATSQRFVTTGGVQGNP--GRETVDNWFKIERFE---SGYKLVFCPTVCRECE 170
+ + T V G+Q R+TVD WF IER + YKLVFCP + +
Sbjct: 109 ITLC-AGMPTEWAIVEREGLQAVKLAARDTVDGWFNIERVSREYNDYKLVFCPQQAEDNK 167
Query: 171 VVCKDIGIFLDENRNTRFVLS-DFPFGVKFQR 201
C+DIGI +D++ R VLS + P V+FQ+
Sbjct: 168 --CEDIGIQIDDDGIRRLVLSKNKPLVVQFQK 197
>DRTI_DELRE (P83667) Kunitz-type serine protease inhibitor DrTI
Length = 185
Score = 60.1 bits (144), Expect = 3e-09
Identities = 54/180 (30%), Positives = 79/180 (43%), Gaps = 16/180 (8%)
Query: 27 EEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVEEEEAMQFSFVPFN 86
E+V D G V Y+I+ + I G CP+ +++E+ +Q +P
Sbjct: 4 EKVYDIEGYPVFLGSEYYIVSAIIGAGGGGVRPGRTRGSMCPMSIIQEQSDLQMG-LPVR 62
Query: 87 FK-----KGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGR 141
F +G I T+L + +C+ SS V D + + V GG + +P
Sbjct: 63 FSSPEESQGKIYTDTELEIEFVEKPDCAESSKWVIVKD-----SGEARVAIGGSEDHPQG 117
Query: 142 ETVDNWFKIERFES-GYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLS-DFPFGVKF 199
E V +FKIE+ S YKLVFCP + C DIGI + R+ S D PF V F
Sbjct: 118 ELVRGFFKIEKLGSLAYKLVFCP---KSSSGSCSDIGINYEGRRSLVLKSSDDSPFRVVF 174
>IAAS_ORYSA (P29421) Alpha-amylase/subtilisin inhibitor (RASI)
Length = 176
Score = 58.2 bits (139), Expect = 1e-08
Identities = 56/185 (30%), Positives = 83/185 (44%), Gaps = 22/185 (11%)
Query: 26 PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVEEEEAMQFSFVPF 85
P V DT G+ + +Y++LP+S G MA CPL V +E + + F P
Sbjct: 2 PPPVYDTEGHELSADGSYYVLPASPGHGGGLTMAPRRVIP-CPLLVAQETDERRKGF-PV 59
Query: 86 NF-------KKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGN 138
F +RVSTD+ + + T C S T W V + ++R VT + +
Sbjct: 60 RFTPWGGAASPRTLRVSTDVRIRFNAATLCVQS--TEWHVGDEPLTGARRVVTGPLIGPS 117
Query: 139 P-GRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSDFPFGV 197
P GRE N F++E++ GYKLV C C+D+G+ D S P V
Sbjct: 118 PSGRE---NAFRVEKYGGGYKLV-------SCRDSCQDLGVSRDGAGAAWIGASQPPHVV 167
Query: 198 KFQRA 202
F++A
Sbjct: 168 VFKKA 172
>KTI2_SOYBN (P25273) Kunitz-type trypsin inhibitor KTI2 precursor
Length = 204
Score = 57.8 bits (138), Expect = 2e-08
Identities = 65/213 (30%), Positives = 93/213 (43%), Gaps = 27/213 (12%)
Query: 1 MKLHFPLLFLLLTLFTTKPLQGAEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL 60
MK L+ FT L A + V DT + ++N Y++LP + G +
Sbjct: 1 MKSTIFFALFLVCAFTISYLPSATA-QFVLDTDDDPLQNGGTYYMLP--VMRGKSGGIEG 57
Query: 61 LNTNKT-CPLDVVEEEEAMQFSFVPFNFKKGVIRVSTD----LNVIHSFPTNCSTSSVTV 115
+T K CPL VV+ P KG+ V L + +P + S V
Sbjct: 58 NSTGKEICPLTVVQS---------PNKHNKGIGLVFKSPLHALFIAERYPLSIKFDSFAV 108
Query: 116 WKVDKVDVATSQRFVTTGGVQGNP--GRETVDNWFKIER----FESGYKLVFCPTVCREC 169
+ V + T V G+Q R+TVD WF IER + YKLVFCP +
Sbjct: 109 IPLCGV-MPTKWAIVEREGLQAVTLAARDTVDGWFNIERVSREYNDYYKLVFCPQEAEDN 167
Query: 170 EVVCKDIGIFLDENRNTRFVLS-DFPFGVKFQR 201
+ C+DIGI +D + R VLS + P V+FQ+
Sbjct: 168 K--CEDIGIQIDNDGIRRLVLSKNKPLVVEFQK 198
>ITRY_ACACO (P24924) Trypsin inhibitor precursor
Length = 176
Score = 55.5 bits (132), Expect = 9e-08
Identities = 47/187 (25%), Positives = 83/187 (44%), Gaps = 26/187 (13%)
Query: 27 EEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVEEEEAMQFSFVPFN 86
+E+ D G+++RN Y+ILP+ G +A +++CPL VV+ + +
Sbjct: 1 KELLDADGDILRNGGAYYILPALRGKGGGLTLAKTG-DESCPLTVVQAQSETKRGLPAVI 59
Query: 87 FKKGVIRVSTDLNVIH------SFPTNCSTSSVTVWKVD----KVDVATSQRFVTTGGVQ 136
+ I + T ++ P S WKV+ +V +A ++
Sbjct: 60 WTPPKIAILTPGFYLNFEFQPRDLPACLQKYSTLPWKVEGESQEVKIAPKEK-------- 111
Query: 137 GNPGRETVDNWFKIERFESGYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSD-FPF 195
+ + FKI+ + YKLV+C + CKD+GI +D+ N R V+ D P
Sbjct: 112 ----EQFLVGSFKIKPYRDDYKLVYCEG--NSDDESCKDLGISIDDENNRRLVVKDGHPL 165
Query: 196 GVKFQRA 202
V+F++A
Sbjct: 166 AVRFEKA 172
>IT2_PSOTE (P25700) Trypsin inhibitor 2 (WTI-2)
Length = 182
Score = 54.7 bits (130), Expect = 1e-07
Identities = 52/185 (28%), Positives = 85/185 (45%), Gaps = 17/185 (9%)
Query: 27 EEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVV---EEEEAMQFSFV 83
ZE+ D G VRN Y+++P G E A + N+ CPL VV +E + +
Sbjct: 1 ZELVDVEGKTVRNGGTYYLVPQLRPGGGGMEAAKVG-NEDCPLTVVKSLDENSNGEPIRI 59
Query: 84 PFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRET 143
+ I + +N+ + P C+ S W V K D + + G + +
Sbjct: 60 ASRLRSTFIPEYSLVNLGFADPPKCAPSPF--WTVVK-DQSERLPSIKLGEYKDSE---- 112
Query: 144 VDNWFKIERFESG-----YKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSDF-PFGV 197
+D FK ER + YKL++C + E E++CKDIG++ D+ R V+S P V
Sbjct: 113 LDYPFKFERVYAASKMYAYKLLYCGSEDEEEEMMCKDIGVYRDQEGYQRLVVSKHNPLVV 172
Query: 198 KFQRA 202
F++A
Sbjct: 173 GFKKA 177
>WIN3_POPSP (P16335) Wound responsive GWIN3 protein precursor
Length = 200
Score = 52.8 bits (125), Expect = 6e-07
Identities = 61/198 (30%), Positives = 78/198 (38%), Gaps = 24/198 (12%)
Query: 7 LLFLLLTLFTTKPLQG--AEQPEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTN 64
L FLL T G AE P V D G V+ +Y I + + N
Sbjct: 9 LSFLLFAFAATSFPDGVHAEDPAAVLDFYGREVQAGASYLIDQEDFR------VVNATIN 62
Query: 65 KTCPLDVVEEE--EAMQFSFVP-FNFKKGVIRVSTDLNVIHSFPTNCSTSSVT-VWKVDK 120
C DV+ E + +F P N GVIR T + V T C + VT +WK+
Sbjct: 63 PICNSDVILSTGIEGLPVTFSPVINSTDGVIREGTLITVSFDAST-CGMAGVTPMWKIGF 121
Query: 121 VDVATSQRFVTTGGVQGNPGRETVDNWFKIERFESG---YKLVFCPTVCRECEVVCKDIG 177
A VTTGGV N FKI +FES Y+L +CP CE C +G
Sbjct: 122 NSTAKGY-IVTTGGVDRL-------NLFKITKFESDSSFYQLSYCPNSEPFCECPCVPVG 173
Query: 178 IFLDENRNTRFVLSDFPF 195
D+ +DF F
Sbjct: 174 ANSDKYLAPNVSYADFRF 191
>API9_SOLTU (P58521) Aspartic protease inhibitor 9 (Novel inhibitor
of cathepsin D) (NID)
Length = 187
Score = 52.4 bits (124), Expect = 7e-07
Identities = 49/187 (26%), Positives = 81/187 (43%), Gaps = 18/187 (9%)
Query: 26 PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALL----NTNKTCPLDVVEEEEAMQFS 81
P+ V DT+G + + +Y I+ SI G L N++ CP V + S
Sbjct: 5 PKPVLDTNGKELNPNSSYRII--SIGAGALGGDVYLGKSPNSDAPCPDGVFRYNSDVGPS 62
Query: 82 FVPFNF--KKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNP 139
P F G I LN+ + PT S T+WKV ++ + TGG G
Sbjct: 63 GTPVRFIPLSGGIFEDQLLNIQFNIPTVKLCVSYTIWKVGNLNAYFRTMLLETGGTIG-- 120
Query: 140 GRETVDNWFKIERFES-GYKLVFCP---TVCREC--EVVCKDIGIFLDENRNTRFVLSDF 193
+ +++FKI + + GY L+ CP +C C + C +G+ + + ++++
Sbjct: 121 --QADNSYFKIVKLSNFGYNLLSCPFTSIICLRCPEDQFCAKVGVVIQNGKRRLALVNEN 178
Query: 194 PFGVKFQ 200
P V FQ
Sbjct: 179 PLDVLFQ 185
>SPI2_SOLTU (P58515) Serine protease inhibitor 2 (PSPI-21)
(PSPI-21-5.2)
Length = 186
Score = 51.2 bits (121), Expect = 2e-06
Identities = 28/104 (26%), Positives = 47/104 (44%), Gaps = 5/104 (4%)
Query: 98 LNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFES-G 156
LN+ + T+ S T+WKV D + + TGG G + +WFKI + G
Sbjct: 85 LNIQFAISTSKLCVSYTIWKVGDYDASLGTMLLETGGTIG----QADSSWFKIVKSSQLG 140
Query: 157 YKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSDFPFGVKFQ 200
Y L++CP + C +G+ + ++++ P V FQ
Sbjct: 141 YNLLYCPVTSSSDDQFCSKVGVVHQNGKRRLALVNENPLDVLFQ 184
>API1_SOLTU (Q41480) Aspartic protease inhibitor 1 precursor (pA1)
(gCDI-A1) (STPIA) (STPID)
Length = 221
Score = 51.2 bits (121), Expect = 2e-06
Identities = 52/212 (24%), Positives = 90/212 (41%), Gaps = 21/212 (9%)
Query: 5 FPLLFLLLTLFTTKPLQGAEQ---PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL- 60
FP+L T + P+ + P+ V DT+G + + +Y I+ S+ ++ L
Sbjct: 13 FPILVFSSTFTSQNPINLPSESPVPKPVLDTNGKKLNPNSSYRII-STFWGALGGDVYLG 71
Query: 61 --LNTNKTCPLDVVEEEEAMQFSFVPFNFKKGVIRVSTD--LNVIHSFPTNCSTSSVTVW 116
N++ CP V + S P F + D LN+ + PT S T+W
Sbjct: 72 KSPNSDAPCPDGVFRYNSDVGPSGTPVRFIPLSTNIFEDQLLNIQFNIPTVKLCVSYTIW 131
Query: 117 KVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFES-GYKLVFC-----PTVCREC- 169
KV ++ + TGG G + ++FKI + GY L++C P VC C
Sbjct: 132 KVGNLNTHLWTMLLETGGTIG----KADSSYFKIVKSSKFGYNLLYCPITRPPIVCPFCR 187
Query: 170 -EVVCKDIGIFLDENRNTRFVLSDFPFGVKFQ 200
+ C +G+ + + ++++ P V FQ
Sbjct: 188 DDDFCAKVGVVIQNGKRRLALVNENPLDVLFQ 219
>SPI1_SOLTU (P58514) Serine protease inhibitor 1 (PSPI-21)
(PSPI-21-6.3)
Length = 187
Score = 50.8 bits (120), Expect = 2e-06
Identities = 27/105 (25%), Positives = 47/105 (44%), Gaps = 5/105 (4%)
Query: 98 LNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFES-G 156
LN+ + T+ S T+WKV D + + TGG G + +WFKI + G
Sbjct: 85 LNIQFAISTSKLCVSYTIWKVGDYDASLGTMLLETGGTIG----QADSSWFKIVKSSQLG 140
Query: 157 YKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSDFPFGVKFQR 201
Y L++CP + C +G+ + ++ D P + F++
Sbjct: 141 YNLLYCPVTSSSDDQFCLKVGVVHQNGKRRLALVKDNPLDISFKQ 185
>API7_SOLTU (Q41448) Aspartic protease inhibitor 7 precursor
(Cathepsin D inhibitor p749)
Length = 221
Score = 50.8 bits (120), Expect = 2e-06
Identities = 51/213 (23%), Positives = 89/213 (40%), Gaps = 23/213 (10%)
Query: 5 FPLLFLLLTLFTTKPLQGAEQ---PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMAL- 60
FP+L T + P+ + P+ V DT+G + + +Y I+ S+ ++ L
Sbjct: 13 FPILVFSSTFTSQNPINLPSESPVPKRVLDTNGKKLNPNSSYRII-STFWGALGGDVYLG 71
Query: 61 --LNTNKTCPLDVVEEEEAMQFSFVPFNF---KKGVIRVSTDLNVIHSFPTNCSTSSVTV 115
N++ CP V + S P F I LN+ + PT S T+
Sbjct: 72 KSPNSDAPCPDGVFRYNSDVGPSGTPVRFIPLSGANIFEDQLLNIQFNIPTVKLCGSYTI 131
Query: 116 WKVDKVDVATSQRFVTTGGVQGNPGRETVDNWFKIERFES-GYKLVFCPTV-------CR 167
WKV ++ + TGG G + ++FKI + GY L++CP CR
Sbjct: 132 WKVGNINAHLRTMLLETGGTIG----QADSSYFKIVKSSKFGYNLLYCPLTRHFLCPFCR 187
Query: 168 ECEVVCKDIGIFLDENRNTRFVLSDFPFGVKFQ 200
+ + C +G+ + + ++++ P V FQ
Sbjct: 188 D-DNFCAKVGVVIQNGKRRLALVNENPLDVLFQ 219
>IT1B_PSOTE (P32877) Trypsin inhibitor 1B (WTI-1B)
Length = 172
Score = 50.4 bits (119), Expect = 3e-06
Identities = 49/172 (28%), Positives = 71/172 (40%), Gaps = 20/172 (11%)
Query: 27 EEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVEEEEAMQFS---FV 83
E + D+ G LVRN Y++LP G E A T +TCPL VV + +
Sbjct: 1 EPLLDSEGELVRNGGTYYLLPDRWALGGGIEAAATGT-ETCPLTVVRSPNEVSVGEPLRI 59
Query: 84 PFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRET 143
+ G I + + + + P C+ S W V V+ Q V ++
Sbjct: 60 SSQLRSGFIPDYSLVRIGFANPPKCAPS--PWWTV--VEDQPQQPSVKLSELKST----K 111
Query: 144 VDNWFKIERFE---SGYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSD 192
D FK E+ S YKL +C CKDIGI+ D+ R V++D
Sbjct: 112 FDYLFKFEKVTSKFSSYKLKYCAK-----RDTCKDIGIYRDQKGYARLVVTD 158
>IT1A_PSOTE (P10821) Trypsin inhibitor 1A (WTI-1A)
Length = 172
Score = 50.4 bits (119), Expect = 3e-06
Identities = 49/172 (28%), Positives = 71/172 (40%), Gaps = 20/172 (11%)
Query: 27 EEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALLNTNKTCPLDVVEEEEAMQFS---FV 83
E + D+ G LVRN Y++LP G E A T +TCPL VV + +
Sbjct: 1 EPLLDSEGELVRNGGTYYLLPDRWALGGGIEAAATGT-ETCPLTVVRSPNEVSVGEPLRI 59
Query: 84 PFNFKKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNPGRET 143
+ G I + + + + P C+ S W V V+ Q V ++
Sbjct: 60 SSQLRSGFIPDYSVVRIGFANPPKCAPS--PWWTV--VEDQPQQPSVKLSELKST----K 111
Query: 144 VDNWFKIERFE---SGYKLVFCPTVCRECEVVCKDIGIFLDENRNTRFVLSD 192
D FK E+ S YKL +C CKDIGI+ D+ R V++D
Sbjct: 112 FDYLFKFEKVTSKFSSYKLKYCAK-----RDTCKDIGIYRDQKGYERLVVTD 158
>API8_SOLTU (P17979) Aspartic protease inhibitor 8 precursor (pi8)
(PI-8) (API) (API-8) (Cathepsin D inhibitor)
Length = 220
Score = 49.7 bits (117), Expect = 5e-06
Identities = 49/189 (25%), Positives = 80/189 (41%), Gaps = 21/189 (11%)
Query: 26 PEEVRDTSGNLVRNSINYFILPSSIQCGTRCEMALL----NTNKTCPLDVVEEEEAMQFS 81
P+ V DT+G + +Y I+ SI G L N++ CP V + S
Sbjct: 37 PKPVLDTNGKELNPDSSYRII--SIGRGALGGDVYLGKSPNSDAPCPDGVFRYNSDVGPS 94
Query: 82 FVPFNF--KKGVIRVSTDLNVIHSFPTNCSTSSVTVWKVDKVDVATSQRFVTTGGVQGNP 139
P F G I LN+ + PT S T+WKV ++ + TGG G
Sbjct: 95 GTPVRFIPLSGGIFEDQLLNIQFNIPTVKLCVSYTIWKVGNLNAYFRTMLLETGGTIG-- 152
Query: 140 GRETVDNWFKIERFES-GYKLVFCPTV-------CRECEVVCKDIGIFLDENRNTRFVLS 191
+ ++FKI + + GY L++CP CR+ + C +G+ + + +++
Sbjct: 153 --QADSSYFKIVKLSNFGYNLLYCPITPPFLCPFCRD-DNFCAKVGVVIQNGKRRLALVN 209
Query: 192 DFPFGVKFQ 200
+ P V FQ
Sbjct: 210 ENPLDVLFQ 218
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.139 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,266,826
Number of Sequences: 164201
Number of extensions: 963073
Number of successful extensions: 2648
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 2602
Number of HSP's gapped (non-prelim): 56
length of query: 204
length of database: 59,974,054
effective HSP length: 105
effective length of query: 99
effective length of database: 42,732,949
effective search space: 4230561951
effective search space used: 4230561951
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC124970.4


