

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124963.1 + phase: 1 /pseudo
(826 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
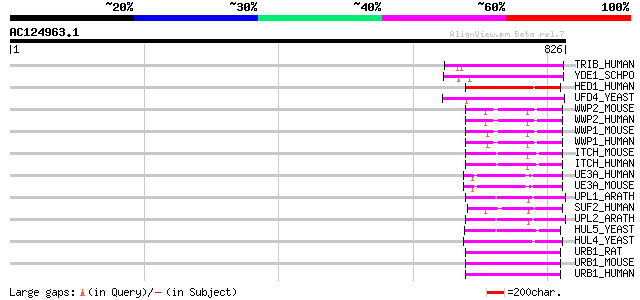
Score E
Sequences producing significant alignments: (bits) Value
TRIB_HUMAN (Q14669) Thyroid receptor interacting protein 12 (TRI... 150 1e-35
YDE1_SCHPO (Q10435) Probable ubiquitin fusion degradation protei... 137 9e-32
HED1_HUMAN (Q9ULT8) HECT domain containing protein 1 (Fragment) 132 5e-30
UFD4_YEAST (P33202) Ubiquitin fusion degradation protein 4 (UB f... 126 2e-28
WWP2_MOUSE (Q9DBH0) Nedd-4-like E3 ubiquitin-protein ligase WWP2... 105 4e-22
WWP2_HUMAN (O00308) Nedd-4-like E3 ubiquitin-protein ligase WWP2... 104 8e-22
WWP1_MOUSE (Q8BZZ3) Nedd-4-like E3 ubiquitin-protein ligase WWP1... 100 1e-20
WWP1_HUMAN (Q9H0M0) Nedd-4-like E3 ubiquitin-protein ligase WWP1... 100 1e-20
ITCH_MOUSE (Q8C863) Itchy E3 ubiquitin protein ligase (EC 6.3.2.-) 99 3e-20
ITCH_HUMAN (Q96J02) Itchy homolog E3 ubiquitin protein ligase (E... 97 1e-19
UE3A_HUMAN (Q05086) Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E... 94 2e-18
UE3A_MOUSE (O08759) Ubiquitin-protein ligase E3A (EC 6.3.2.-) (O... 89 6e-17
UPL1_ARATH (Q8GY23) E3 ubiquitin protein ligase UPL1 (EC 6.3.2.-... 88 8e-17
SUF2_HUMAN (Q9HAU4) Smad ubiquitination regulatory factor 2 (EC ... 86 5e-16
UPL2_ARATH (Q8H0T4) E3 ubiquitin protein ligase UPL2 (EC 6.3.2.-... 85 7e-16
HUL5_YEAST (P53119) Probable ubiquitin--protein ligase HUL5 (EC ... 84 1e-15
HUL4_YEAST (P40985) Probable ubiquitin--protein ligase HUL4 (EC ... 82 6e-15
URB1_RAT (P51593) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-... 82 8e-15
URB1_MOUSE (Q7TMY8) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2... 82 8e-15
URB1_HUMAN (Q7Z6Z7) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2... 82 8e-15
>TRIB_HUMAN (Q14669) Thyroid receptor interacting protein 12 (TRIP12)
Length = 1992
Score = 150 bits (380), Expect = 1e-35
Identities = 89/203 (43%), Positives = 120/203 (58%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ ++ + F GF
Sbjct: 1787 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVSRQFDSFRDGFE 1846
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1847 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1906
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1907 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1966
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1967 DYSSIEIMREKLLIAAREGQQSF 1989
>YDE1_SCHPO (Q10435) Probable ubiquitin fusion degradation protein
C12B10.01c
Length = 1647
Score = 137 bits (346), Expect = 9e-32
Identities = 82/203 (40%), Positives = 112/203 (54%), Gaps = 24/203 (11%)
Query: 646 KIEDLCLDFSLPGYDQTFLLP---------FSIY*ITNLYSFFLVG-------------F 683
K+EDL LDF+LPG+ L+P F++ N + VG F
Sbjct: 1442 KLEDLHLDFTLPGFPSIELIPDGASTPVTTFNVSDYLNYVIDYTVGKGVQQQLEAFQNGF 1501
Query: 684 NQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
+ VFP L+V +E EL + W++ L+K DHGYT SP I LL ++ +
Sbjct: 1502 SSVFPYTSLQVLTEHELVTLFGTVDEDWSYATLMKSIVADHGYTMESPTIQRLLTLMSQM 1561
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNH--TDTDLPSVMTCANYLKLP 801
N +E+R F+QF+T S +LP GG A L+P LTVV++++ D LPSVMTC NYLKLP
Sbjct: 1562 NFQEQRDFLQFITGSRKLPIGGFAGLNPPLTVVRRLNEPPYVPDDYLPSVMTCVNYLKLP 1621
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E + +L AI EG+G F
Sbjct: 1622 EYSSSEVLGSRLSKAILEGQGSF 1644
>HED1_HUMAN (Q9ULT8) HECT domain containing protein 1 (Fragment)
Length = 1620
Score = 132 bits (331), Expect = 5e-30
Identities = 68/142 (47%), Positives = 96/142 (66%), Gaps = 3/142 (2%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPN-SWTFNDLLKHFKFDHGYTARSPPIMNLL 737
F GFN+VFP+E L FS EE+++ILCG + SW D++ + + GYT SP + +
Sbjct: 1475 FRDGFNKVFPMEKLSSFSHEEVQMILCGNQSPSWAAEDIINYTEPKLGYTRDSPGFLRFV 1534
Query: 738 EILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANY 797
+L +++ER+AF+QF T LPPGGLA+L P+LTVV+K+ + TD PSV TC +Y
Sbjct: 1535 RVLCGMSSDERKAFLQFTTGCSTLPPGGLANLHPRLTVVRKV--DATDASYPSVNTCVHY 1592
Query: 798 LKLPPYSSKERMKQKLLYAITE 819
LKLP YSS+E M+++LL A E
Sbjct: 1593 LKLPEYSSEEIMRERLLAATME 1614
>UFD4_YEAST (P33202) Ubiquitin fusion degradation protein 4 (UB fusion
protein 4)
Length = 1483
Score = 126 bits (317), Expect = 2e-28
Identities = 78/206 (37%), Positives = 105/206 (50%), Gaps = 24/206 (11%)
Query: 644 NSKIEDLCLDFSLPGYDQTFLLPFSIY*ITNLYSF----------------------FLV 681
N +E L L F++PG D L+P N + F+
Sbjct: 1276 NMTLESLSLTFTVPGNDDIELIPGGCNKSLNSSNVEEYIHGVIDQILGKGIEKQLKAFIE 1335
Query: 682 GFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQ 741
GF++VF E + + +EL I W+ L + +HGYT S I + + I+
Sbjct: 1336 GFSKVFSYERMLILFPDELVDIFGRVEEDWSMATLYTNLNAEHGYTMDSSIIHDFISIIS 1395
Query: 742 EFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTD--LPSVMTCANYLK 799
F ERR F+QF+T SP+LP GG SL+PK TVV K + + D LPSVMTCANYLK
Sbjct: 1396 AFGKHERRLFLQFLTGSPKLPIGGFKSLNPKFTVVLKHAEDGLTADEYLPSVMTCANYLK 1455
Query: 800 LPPYSSKERMKQKLLYAITEGRGCFL 825
LP Y+SK+ M+ +L AI EG G FL
Sbjct: 1456 LPKYTSKDIMRSRLCQAIEEGAGAFL 1481
>WWP2_MOUSE (Q9DBH0) Nedd-4-like E3 ubiquitin-protein ligase WWP2
(EC 6.3.2.-) (WW domain-containing protein 2)
Length = 870
Score = 105 bits (263), Expect = 4e-22
Identities = 64/152 (42%), Positives = 88/152 (57%), Gaps = 17/152 (11%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCG----EPNSWTFNDLLKHFKFDHGYTARSPPIM 734
FL GFN+V P+E LR F E+ELEL+LCG + + W N + +H YT S I
Sbjct: 724 FLDGFNEVAPLEWLRYFDEKELELMLCGMQEIDMSDWQKNAIYRH------YTKSSKQIQ 777
Query: 735 NLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASL----DPKLTVVQKISYNHTDTDLPS 790
++++E +NE+R +QFVT + RLP GG A L P+ + ++ +T LP
Sbjct: 778 WFWQVVKEMDNEKRIRLLQFVTGTCRLPVGGFAELIGSNGPQKFCIDRVG---KETWLPR 834
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
TC N L LPPY S E++K+KLLYAI E G
Sbjct: 835 SHTCFNRLDLPPYKSYEQLKEKLLYAIEETEG 866
>WWP2_HUMAN (O00308) Nedd-4-like E3 ubiquitin-protein ligase WWP2
(EC 6.3.2.-) (WW domain-containing protein 2) (Atropin-1
interacting protein 2) (AIP2)
Length = 870
Score = 104 bits (260), Expect = 8e-22
Identities = 63/152 (41%), Positives = 88/152 (57%), Gaps = 17/152 (11%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCG----EPNSWTFNDLLKHFKFDHGYTARSPPIM 734
FL GFN+V P+E LR F E+ELEL+LCG + + W + + +H YT S I
Sbjct: 724 FLDGFNEVAPLEWLRYFDEKELELMLCGMQEIDMSDWQKSTIYRH------YTKNSKQIQ 777
Query: 735 NLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASL----DPKLTVVQKISYNHTDTDLPS 790
++++E +NE+R +QFVT + RLP GG A L P+ + K+ +T LP
Sbjct: 778 WFWQVVKEMDNEKRIRLLQFVTGTCRLPVGGFAELIGSNGPQKFCIDKVG---KETWLPR 834
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
TC N L LPPY S E++++KLLYAI E G
Sbjct: 835 SHTCFNRLDLPPYKSYEQLREKLLYAIEETEG 866
>WWP1_MOUSE (Q8BZZ3) Nedd-4-like E3 ubiquitin-protein ligase WWP1
(EC 6.3.2.-) (WW domain-containing protein 1)
Length = 918
Score = 100 bits (250), Expect = 1e-20
Identities = 62/152 (40%), Positives = 86/152 (55%), Gaps = 17/152 (11%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPN----SWTFNDLLKHFKFDHGYTARSPPIM 734
FL GFN+V P++ L+ F E+ELE++LCG W N + +H YT S I+
Sbjct: 772 FLDGFNEVVPLQWLQYFDEKELEVMLCGMQEVDLADWQRNTVYRH------YTRNSKQII 825
Query: 735 NLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASL----DPKLTVVQKISYNHTDTDLPS 790
+ ++E +NE R +QFVT + RLP GG A L P+ ++K+ DT LP
Sbjct: 826 WFWQFVKETDNEVRMRLLQFVTGTCRLPLGGFAELMGSNGPQKFCIEKVG---KDTWLPR 882
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
TC N L LPPY S E++K+KLL+AI E G
Sbjct: 883 SHTCFNRLDLPPYKSYEQLKEKLLFAIEETEG 914
>WWP1_HUMAN (Q9H0M0) Nedd-4-like E3 ubiquitin-protein ligase WWP1
(EC 6.3.2.-) (WW domain-containing protein 1) (Atropin-1
interacting protein 5) (AIP5)
Length = 922
Score = 100 bits (250), Expect = 1e-20
Identities = 62/152 (40%), Positives = 86/152 (55%), Gaps = 17/152 (11%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPN----SWTFNDLLKHFKFDHGYTARSPPIM 734
FL GFN+V P++ L+ F E+ELE++LCG W N + +H YT S I+
Sbjct: 776 FLDGFNEVVPLQWLQYFDEKELEVMLCGMQEVDLADWQRNTVYRH------YTRNSKQII 829
Query: 735 NLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASL----DPKLTVVQKISYNHTDTDLPS 790
+ ++E +NE R +QFVT + RLP GG A L P+ ++K+ DT LP
Sbjct: 830 WFWQFVKETDNEVRMRLLQFVTGTCRLPLGGFAELMGSNGPQKFCIEKVG---KDTWLPR 886
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
TC N L LPPY S E++K+KLL+AI E G
Sbjct: 887 SHTCFNRLDLPPYKSYEQLKEKLLFAIEETEG 918
>ITCH_MOUSE (Q8C863) Itchy E3 ubiquitin protein ligase (EC 6.3.2.-)
Length = 864
Score = 99.4 bits (246), Expect = 3e-20
Identities = 59/148 (39%), Positives = 85/148 (56%), Gaps = 9/148 (6%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F GFN++ P ++L+ F +ELE++LCG ND +H + H YT S IM +
Sbjct: 718 FFEGFNEILPQQYLQYFDAKELEVLLCGMQEI-DLNDWQRHAIYRH-YTRTSKQIMWFWQ 775
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASL----DPKLTVVQKISYNHTDTDLPSVMTC 794
++E +NE+R +QFVT + RLP GG A L P+ ++K+ + LP TC
Sbjct: 776 FVKEIDNEKRMRLLQFVTGTCRLPVGGFADLMGSNGPQKFCIEKVGKENW---LPRSHTC 832
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRG 822
N L LPPY S E++K+KLL+AI E G
Sbjct: 833 FNRLDLPPYKSYEQLKEKLLFAIEETEG 860
>ITCH_HUMAN (Q96J02) Itchy homolog E3 ubiquitin protein ligase (EC
6.3.2.-) (Itch) (Atrophin-1-interacting protein 4)
(AIP4) (NFE2-associated polypeptide 1) (NAPP1)
Length = 903
Score = 97.4 bits (241), Expect = 1e-19
Identities = 58/148 (39%), Positives = 84/148 (56%), Gaps = 9/148 (6%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F GFN++ P ++L+ F +ELE++LCG ND +H + H Y S IM +
Sbjct: 757 FFEGFNEILPQQYLQYFDAKELEVLLCGMQEI-DLNDWQRHAIYRH-YARTSKQIMWFWQ 814
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASL----DPKLTVVQKISYNHTDTDLPSVMTC 794
++E +NE+R +QFVT + RLP GG A L P+ ++K+ + LP TC
Sbjct: 815 FVKEIDNEKRMRLLQFVTGTCRLPVGGFADLMGSNGPQKFCIEKVGKENW---LPRSHTC 871
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRG 822
N L LPPY S E++K+KLL+AI E G
Sbjct: 872 FNRLDLPPYKSYEQLKEKLLFAIEETEG 899
>UE3A_HUMAN (Q05086) Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP
ubiquitin-protein ligase) (Oncogenic protein-associated
protein E6-AP) (Human papillomavirus E6-associated
protein)
Length = 875
Score = 93.6 bits (231), Expect = 2e-18
Identities = 62/151 (41%), Positives = 89/151 (58%), Gaps = 13/151 (8%)
Query: 676 YSFFLVGFNQVF---PIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPP 732
+ F GF+ V P+++L F EE+EL++CG N F L + ++D GYT S
Sbjct: 730 FKAFRRGFHMVTNESPLKYL--FRPEEIELLICGSRNL-DFQALEETTEYDGGYTRDSVL 786
Query: 733 IMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTD-LPSV 791
I EI+ F +E++R F+QF T + R P GGL KL ++ I+ N DT+ LP+
Sbjct: 787 IREFWEIVHSFTDEQKRLFLQFTTGTDRAPVGGLG----KLKMI--IAKNGPDTERLPTS 840
Query: 792 MTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
TC N L LP YSSKE++K++LL AIT +G
Sbjct: 841 HTCFNVLLLPEYSSKEKLKERLLKAITYAKG 871
>UE3A_MOUSE (O08759) Ubiquitin-protein ligase E3A (EC 6.3.2.-)
(Oncogenic protein-associated protein E6-AP)
Length = 885
Score = 88.6 bits (218), Expect = 6e-17
Identities = 60/151 (39%), Positives = 86/151 (56%), Gaps = 13/151 (8%)
Query: 676 YSFFLVGFNQVF---PIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPP 732
+ F GF+ V P+++L F EE+EL++CG N F L + ++D GYT S
Sbjct: 740 FKAFRRGFHMVTNESPLKYL--FRPEEIELLICGSRNL-DFQALEETTEYDGGYTRESVV 796
Query: 733 IMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTD-LPSV 791
I EI+ F +E++R F+ F T + R P GGL KL ++ I+ N DT+ LP+
Sbjct: 797 IREFWEIVHSFTDEQKRLFLLFTTGTDRAPVGGLG----KLKMI--IAKNGPDTERLPTS 850
Query: 792 MTCANYLKLPPYSSKERMKQKLLYAITEGRG 822
TC N L LP YSSKE++ +LL AIT +G
Sbjct: 851 HTCFNVLLLPEYSSKEKLNVRLLKAITYAKG 881
>UPL1_ARATH (Q8GY23) E3 ubiquitin protein ligase UPL1 (EC 6.3.2.-)
(Ubiquitin-protein ligase 1)
Length = 3684
Score = 88.2 bits (217), Expect = 8e-17
Identities = 57/152 (37%), Positives = 89/152 (58%), Gaps = 8/152 (5%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GFN++ P E + +F+++ELEL++ G P F+DL + ++ YTA SP I E
Sbjct: 3533 FLEGFNELIPRELVSIFNDKELELLISGLPEI-DFDDLKANTEYT-SYTAGSPVIHWFWE 3590
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLD----PKLTVVQKISYNHTDTDLPSVMTC 794
+++ F+ E+ F+QFVT + ++P G +L P+ + K +Y + LPS TC
Sbjct: 3591 VVKAFSKEDMARFLQFVTGTSKVPLEGFKALQGISGPQRLQIHK-AYGAPER-LPSAHTC 3648
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
N L LP Y SKE+++++LL AI E F F
Sbjct: 3649 FNQLDLPEYQSKEQLQERLLLAIHEASEGFGF 3680
>SUF2_HUMAN (Q9HAU4) Smad ubiquitination regulatory factor 2 (EC
6.3.2.-) (Ubiquitin--protein ligase SMURF2)
(Smad-specific E3 ubiquitin ligase 2) (hSMURF2)
Length = 748
Score = 85.5 bits (210), Expect = 5e-16
Identities = 59/150 (39%), Positives = 79/150 (52%), Gaps = 18/150 (12%)
Query: 682 GFNQVFPIEHLRVFSEEELELILCG----EPNSWTFNDLLKHFKFDHGYTARSPPIMNLL 737
GFN+V P L+ F E+ELELI+CG + N W N LKH D I+
Sbjct: 604 GFNEVIPQHLLKTFDEKELELIICGLGKIDVNDWKVNTRLKHCTPDSN-------IVKWF 656
Query: 738 EILQEFNNEERRA-FVQFVTRSPRLPPGGLASLD----PKLTVVQKISYNHTDTDLPSVM 792
EF +EERRA +QFVT S R+P G +L P+L + +I + +LP
Sbjct: 657 WKAVEFFDEERRARLLQFVTGSSRVPLQGFKALQGAAGPRLFTIHQI--DACTNNLPKAH 714
Query: 793 TCANYLKLPPYSSKERMKQKLLYAITEGRG 822
TC N + +PPY S E++ +KLL AI E G
Sbjct: 715 TCFNRIDIPPYESYEKLYEKLLTAIEETCG 744
>UPL2_ARATH (Q8H0T4) E3 ubiquitin protein ligase UPL2 (EC 6.3.2.-)
(Ubiquitin-protein ligase 2)
Length = 3658
Score = 85.1 bits (209), Expect = 7e-16
Identities = 55/152 (36%), Positives = 87/152 (57%), Gaps = 8/152 (5%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL G N++ P E + +F+++ELEL++ G P F+DL + ++ YT SP I E
Sbjct: 3510 FLEGLNELIPRELVSIFNDKELELLISGLPEI-DFDDLKANTEYT-SYTVGSPVIRWFWE 3567
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLD----PKLTVVQKISYNHTDTDLPSVMTC 794
+++ F+ E+ F+QFVT + ++P G +L P+ + K +Y + LPS TC
Sbjct: 3568 VVKAFSKEDMARFLQFVTGTSKVPLEGFKALQGISGPQRLQIHK-AYGSPER-LPSAHTC 3625
Query: 795 ANYLKLPPYSSKERMKQKLLYAITEGRGCFLF 826
N L LP Y SKE+++++LL AI E F F
Sbjct: 3626 FNQLDLPEYQSKEQVQERLLLAIHEANEGFGF 3657
>HUL5_YEAST (P53119) Probable ubiquitin--protein ligase HUL5 (EC
6.3.2.-)
Length = 910
Score = 84.3 bits (207), Expect = 1e-15
Identities = 48/144 (33%), Positives = 79/144 (54%), Gaps = 4/144 (2%)
Query: 677 SFFLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNL 736
S F G + + + +F+ EL++++ GE ++ +DL + ++ GY I++
Sbjct: 765 SAFHGGLSVIIAPHWMEMFNSIELQMLISGERDNIDLDDLKSNTEYG-GYKEEDQTIVDF 823
Query: 737 LEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCAN 796
E+L EF EE+ F++FVT P+ P G +LDPK + + + LP+ TC N
Sbjct: 824 WEVLNEFKFEEKLNFLKFVTSVPQAPLQGFKALDPKFGIRNAGTEKYR---LPTASTCVN 880
Query: 797 YLKLPPYSSKERMKQKLLYAITEG 820
LKLP Y +K +++KLLYAI G
Sbjct: 881 LLKLPDYRNKTILREKLLYAINSG 904
>HUL4_YEAST (P40985) Probable ubiquitin--protein ligase HUL4 (EC
6.3.2.-)
Length = 892
Score = 82.0 bits (201), Expect = 6e-15
Identities = 49/150 (32%), Positives = 86/150 (56%), Gaps = 7/150 (4%)
Query: 676 YSFFLVGFNQVFP-IEHLRVFSEEELELILCG--EPNSWTFNDLLKHFKFDHGYTARSPP 732
Y+ F+ GF +VF +++F+ EELE ++CG E + F L K+ G++ S
Sbjct: 743 YNKFVSGFKRVFAECNSIKLFNSEELERLVCGDEEQTKFDFKSLRSVTKYVGGFSDDSRA 802
Query: 733 IMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVM 792
+ EI++ ++ ++ +QFVT S R+P G++++ K++++ +H DLP
Sbjct: 803 VCWFWEIIESWDYPLQKKLLQFVTASDRIPATGISTIPFKISLLG----SHDSDDLPLAH 858
Query: 793 TCANYLKLPPYSSKERMKQKLLYAITEGRG 822
TC N + L YSSK++++ KLL+AI E G
Sbjct: 859 TCFNEICLWNYSSKKKLELKLLWAINESEG 888
>URB1_RAT (P51593) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-)
(Upstream regulatory element binding protein 1)
(Fragment)
Length = 322
Score = 81.6 bits (200), Expect = 8e-15
Identities = 53/144 (36%), Positives = 81/144 (55%), Gaps = 6/144 (4%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GF ++ P + +F+E+ELEL++ G P +DL + ++ H Y + S I
Sbjct: 174 FLEGFYEIIPKRLISIFTEQELELLISGLPTI-DIDDLKSNTEY-HKYQSNSIQIQWFWR 231
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTD--TD-LPSVMTCA 795
L+ F+ +R F+QFVT + ++P G A+L+ + +QK + D TD LPS TC
Sbjct: 232 ALRSFDQADRAKFLQFVTGTSKVPLQGFAALEG-MNGIQKFQIHRDDRSTDRLPSAHTCF 290
Query: 796 NYLKLPPYSSKERMKQKLLYAITE 819
N L LP Y S E+++ LL AI E
Sbjct: 291 NQLDLPAYESFEKLRHMLLLAIQE 314
>URB1_MOUSE (Q7TMY8) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-)
(Fragment)
Length = 2749
Score = 81.6 bits (200), Expect = 8e-15
Identities = 53/144 (36%), Positives = 81/144 (55%), Gaps = 6/144 (4%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GF ++ P + +F+E+ELEL++ G P +DL + ++ H Y + S I
Sbjct: 2601 FLEGFYEIIPKRLISIFTEQELELLISGLPTI-DIDDLKSNTEY-HKYQSNSIQIQWFWR 2658
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTD--TD-LPSVMTCA 795
L+ F+ +R F+QFVT + ++P G A+L+ + +QK + D TD LPS TC
Sbjct: 2659 ALRSFDQADRAKFLQFVTGTSKVPLQGFAALEG-MNGIQKFQIHRDDRSTDRLPSAHTCF 2717
Query: 796 NYLKLPPYSSKERMKQKLLYAITE 819
N L LP Y S E+++ LL AI E
Sbjct: 2718 NQLDLPAYESFEKLRHMLLLAIQE 2741
>URB1_HUMAN (Q7Z6Z7) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-)
(HSPC272)
Length = 3360
Score = 81.6 bits (200), Expect = 8e-15
Identities = 53/144 (36%), Positives = 81/144 (55%), Gaps = 6/144 (4%)
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
FL GF ++ P + +F+E+ELEL++ G P +DL + ++ H Y + S I
Sbjct: 3212 FLEGFYEIIPKRLISIFTEQELELLISGLPTI-DIDDLKSNTEY-HKYQSNSIQIQWFWR 3269
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTD--TD-LPSVMTCA 795
L+ F+ +R F+QFVT + ++P G A+L+ + +QK + D TD LPS TC
Sbjct: 3270 ALRSFDQADRAKFLQFVTGTSKVPLQGFAALEG-MNGIQKFQIHRDDRSTDRLPSAHTCF 3328
Query: 796 NYLKLPPYSSKERMKQKLLYAITE 819
N L LP Y S E+++ LL AI E
Sbjct: 3329 NQLDLPAYESFEKLRHMLLLAIQE 3352
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.358 0.160 0.582
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 81,508,723
Number of Sequences: 164201
Number of extensions: 2998592
Number of successful extensions: 12201
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 12079
Number of HSP's gapped (non-prelim): 53
length of query: 826
length of database: 59,974,054
effective HSP length: 119
effective length of query: 707
effective length of database: 40,434,135
effective search space: 28586933445
effective search space used: 28586933445
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC124963.1


