

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124953.18 + phase: 0
(391 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
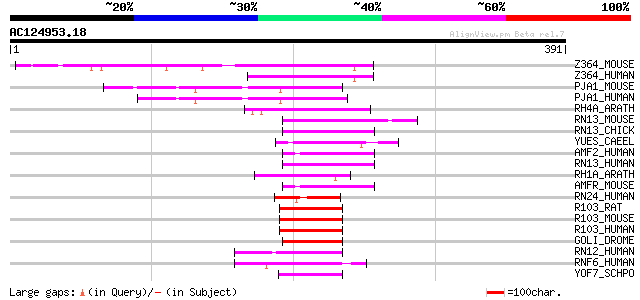
Score E
Sequences producing significant alignments: (bits) Value
Z364_MOUSE (Q9D0C1) Zinc finger protein 364 80 7e-15
Z364_HUMAN (Q9Y4L5) Zinc finger protein 364 (Fragment) 75 2e-13
PJA1_MOUSE (O55176) Ubiquitin protein ligase Praja1 (EC 6.3.2.-) 67 1e-10
PJA1_HUMAN (Q8NG27) Ubiquitin protein ligase Praja1 (EC 6.3.2.-) 66 2e-10
RH4A_ARATH (Q84TF5) RING-H2 zinc finger protein RHA4a 59 2e-08
RN13_MOUSE (O54965) RING finger protein 13 57 8e-08
RN13_CHICK (Q90972) RING finger protein 13 (C-RZF) 57 1e-07
YUES_CAEEL (P90859) Hypothetical RING finger protein F26E4.11 in... 56 1e-07
AMF2_HUMAN (Q9UKV5) Autocrine motility factor receptor, isoform ... 56 1e-07
RN13_HUMAN (O43567) RING finger protein 13 55 2e-07
RH1A_ARATH (Q9SUS4) RING-H2 zinc finger protein RHA1a 54 9e-07
AMFR_MOUSE (Q9R049) Autocrine motility factor receptor (EC 6.3.2... 54 9e-07
RN24_HUMAN (Q9Y225) RING finger protein 24 52 3e-06
R103_RAT (Q9EPZ8) RING finger protein 103 (Zinc finger protein 1... 50 8e-06
R103_MOUSE (Q9R1W3) RING finger protein 103 (Zinc finger protein... 50 8e-06
R103_HUMAN (O00237) RING finger protein 103 (Zinc finger protein... 50 8e-06
GOLI_DROME (Q06003) Goliath protein (Protein g1) 50 8e-06
RN12_HUMAN (Q9NVW2) RING finger protein 12 (LIM domain interacti... 49 2e-05
RNF6_HUMAN (Q9Y252) RING finger protein 6 (RING-H2 protein) 49 2e-05
YOF7_SCHPO (Q9P7E1) RING finger protein P4H10.07 in chromosome II 49 3e-05
>Z364_MOUSE (Q9D0C1) Zinc finger protein 364
Length = 305
Score = 80.5 bits (197), Expect = 7e-15
Identities = 74/287 (25%), Positives = 119/287 (40%), Gaps = 47/287 (16%)
Query: 5 WCYRCNKFVRVWRLGMPICPDCDSGFLEDVEQSTHSANTVGGRRMRFPMAAA-------- 56
+C+ C V +L ICP CDSGF+E+V T ++ +GG R + A
Sbjct: 21 FCHFCKGEVNP-KLPEYICPRCDSGFIEEV---TDDSSFLGGGGSRTDNSTATHFAELWD 76
Query: 57 -----MYMIGHR----NNNYNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSRE 107
M++ R +N +Q+ R + + P P + S G++R
Sbjct: 77 HLDHTMFLQDFRPFLSSNPLDQDNRANERGHQTHTDFWGPSRPPRLPMTRRYRSRGSTRP 136
Query: 108 RE----ENNEFELFYEDGAGSGLRALPPRMS----------ELILG-SGFERVMEQLSHV 152
E ++F A S + P S + G +G + ++ QL
Sbjct: 137 DRSPAIEGIIQQIFAGFFANSAIPGSPHPFSWSGMLHSNPGDYAWGQTGLDAIVTQLL-- 194
Query: 153 EANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCK 212
+ N PA K + LPT+ + + +N C VCKE + + R++PC
Sbjct: 195 ------GQLENTGPPPADKEKITSLPTVTVTQEQVNTGLECPVCKEDYTVEEKVRQLPCN 248
Query: 213 HIYHNECILPWLAIQNSCPVCRHELPCES---PQINNEISNSNEDEN 256
H +H+ CI+PWL + ++CPVCR L E ++E S SN N
Sbjct: 249 HFFHSSCIVPWLELHDTCPVCRKSLNGEDSTRQTQSSEASASNRFSN 295
>Z364_HUMAN (Q9Y4L5) Zinc finger protein 364 (Fragment)
Length = 232
Score = 75.5 bits (184), Expect = 2e-13
Identities = 33/92 (35%), Positives = 51/92 (54%), Gaps = 3/92 (3%)
Query: 168 PALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQ 227
PA K + LPT+ + + +++ C VCKE + + R++PC H +H+ CI+PWL +
Sbjct: 131 PADKEKITSLPTVTVTQEQVDMGLECPVCKEDYTVEEEVRQLPCNHFFHSSCIVPWLELH 190
Query: 228 NSCPVCRHELPCES---PQINNEISNSNEDEN 256
++CPVCR L E + E S SN N
Sbjct: 191 DTCPVCRKSLNGEDSTRQSQSTEASASNRFSN 222
>PJA1_MOUSE (O55176) Ubiquitin protein ligase Praja1 (EC 6.3.2.-)
Length = 605
Score = 66.6 bits (161), Expect = 1e-10
Identities = 51/178 (28%), Positives = 76/178 (42%), Gaps = 17/178 (9%)
Query: 67 YNQNTFRRHRRNNVNGGDISPFNPIIMIRGGGGSSEGTSREREENNEFELFYEDGAGSGL 126
YN+N N I P + M+ G + +S + ++ LF DG GL
Sbjct: 402 YNENESSSEGDNESTHELIQP--GMFMLDGNNNLEDDSSVSEDLEVDWSLF--DGFADGL 457
Query: 127 RAL-------PPRMSELILGSGFERVMEQ-LSHVEANRSGNEGHNQQHLPALKSAVELLP 178
P ++ + L + ME L+H+E+ E N PA K +++ LP
Sbjct: 458 GVAEAISYVDPQFLTYMALEERLAQAMETALAHLESLAVDVEVANP---PASKESIDALP 514
Query: 179 TIEINESHMNV--ESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCR 234
I + E H V E C +C + G A E+PC H +H C+ WL +CPVCR
Sbjct: 515 EILVTEDHGTVGQEMCCPICCSEYVKGEVATELPCHHYFHKPCVSIWLQKSGTCPVCR 572
>PJA1_HUMAN (Q8NG27) Ubiquitin protein ligase Praja1 (EC 6.3.2.-)
Length = 643
Score = 65.9 bits (159), Expect = 2e-10
Identities = 46/158 (29%), Positives = 70/158 (44%), Gaps = 15/158 (9%)
Query: 91 IIMIRGGGGSSEGTSREREENNEFELFYEDGAGSGLRAL-------PPRMSELILGSGFE 143
+ M+ G + +S + ++ LF DG GL P ++ + L
Sbjct: 488 VFMLDGNNNLEDDSSVSEDLEVDWSLF--DGFADGLGVAEAISYVDPQFLTYMALEERLA 545
Query: 144 RVMEQ-LSHVEANRSGNEGHNQQHLPALKSAVELLPTIEINESHMNV--ESHCAVCKEPF 200
+ ME L+H+E+ E N PA K +++ LP I + E H V E C +C +
Sbjct: 546 QAMETALAHLESLAVDVEVANP---PASKESIDALPEILVTEDHGAVGQEMCCPICCSEY 602
Query: 201 ELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELP 238
G A E+PC H +H C+ WL +CPVCR P
Sbjct: 603 VKGEVATELPCHHYFHKPCVSIWLQKSGTCPVCRCMFP 640
>RH4A_ARATH (Q84TF5) RING-H2 zinc finger protein RHA4a
Length = 174
Score = 59.3 bits (142), Expect = 2e-08
Identities = 37/96 (38%), Positives = 49/96 (50%), Gaps = 7/96 (7%)
Query: 166 HLPA---LKSAVEL---LPTIEINESHMNVESHCAVCKEPFELGISAREMP-CKHIYHNE 218
HLP+ L VEL L + NE +S C VC FEL EMP CKHI+H +
Sbjct: 72 HLPSVCLLDVKVELKDKLHVVLFNEELGTRDSLCCVCLGEFELKEELVEMPLCKHIFHLD 131
Query: 219 CILPWLAIQNSCPVCRHELPCESPQINNEISNSNED 254
CI WL N+CP+CR + S + + + N + D
Sbjct: 132 CIHLWLYSHNTCPLCRSSVSISSTKTSVDDDNDHPD 167
>RN13_MOUSE (O54965) RING finger protein 13
Length = 381
Score = 57.0 bits (136), Expect = 8e-08
Identities = 30/96 (31%), Positives = 46/96 (47%), Gaps = 3/96 (3%)
Query: 193 CAVCKEPFELGISAREMPCKHIYHNECILPWLA-IQNSCPVCRHELPCESPQINNEISNS 251
CA+C E +E G R +PC H YH +C+ PWL + +CPVC+ ++ +++ +S
Sbjct: 240 CAICLEEYEDGDKLRILPCSHAYHCKCVDPWLTKTKKTCPVCKQKVVPSQGDSDSDTDSS 299
Query: 252 NEDENVGLTIWRLPGGGFAVGRFSGDGGGGENRMEH 287
E+ V LP A R G E+ H
Sbjct: 300 QEENQVSEHTPLLPPS--ASARTQSFGSLSESHSHH 333
>RN13_CHICK (Q90972) RING finger protein 13 (C-RZF)
Length = 381
Score = 56.6 bits (135), Expect = 1e-07
Identities = 23/66 (34%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Query: 193 CAVCKEPFELGISAREMPCKHIYHNECILPWLA-IQNSCPVCRHELPCESPQINNEISNS 251
CA+C + +E G R +PC H YH +C+ PWL + +CPVC+ ++ ++E +S
Sbjct: 240 CAICLDEYEDGDKLRILPCSHAYHCKCVDPWLTKTKKTCPVCKQKVVPSQGDSDSETDSS 299
Query: 252 NEDENV 257
E+ V
Sbjct: 300 QEENEV 305
>YUES_CAEEL (P90859) Hypothetical RING finger protein F26E4.11 in
chromosome I
Length = 564
Score = 56.2 bits (134), Expect = 1e-07
Identities = 32/89 (35%), Positives = 44/89 (48%), Gaps = 13/89 (14%)
Query: 188 NVESHCAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELPCESPQINN- 246
N + C VC +EL ++R +PC H +H+ C++ WLA +SCP CR +P QI
Sbjct: 330 NGDDRCVVC---WELLGTSRRLPCSHQFHDWCLMWWLAQDSSCPTCRCTIPSPQDQIRQP 386
Query: 247 -EISNSNEDENVGLTIWRLPGGGFAVGRF 274
E+ NS T R GG F F
Sbjct: 387 PEVGNS--------TRLRFNGGSFGFVHF 407
>AMF2_HUMAN (Q9UKV5) Autocrine motility factor receptor, isoform 2
(EC 6.3.2.-) (AMF receptor) (gp78)
Length = 643
Score = 56.2 bits (134), Expect = 1e-07
Identities = 24/66 (36%), Positives = 38/66 (57%), Gaps = 4/66 (6%)
Query: 193 CAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELP-CESPQINNEISNS 251
CA+C + + +AR++PC H++HN C+ WL SCP CR L ++ ++ E
Sbjct: 341 CAICWDSMQ---AARKLPCGHLFHNSCLRSWLEQDTSCPTCRMSLNIADNNRVREEHQGE 397
Query: 252 NEDENV 257
N DEN+
Sbjct: 398 NLDENL 403
>RN13_HUMAN (O43567) RING finger protein 13
Length = 381
Score = 55.5 bits (132), Expect = 2e-07
Identities = 22/66 (33%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Query: 193 CAVCKEPFELGISAREMPCKHIYHNECILPWLA-IQNSCPVCRHELPCESPQINNEISNS 251
CA+C + +E G R +PC H YH +C+ PWL + +CPVC+ ++ +++ +S
Sbjct: 240 CAICLDEYEDGDKLRILPCSHAYHCKCVDPWLTKTKKTCPVCKQKVVPSQGDSDSDTDSS 299
Query: 252 NEDENV 257
E+ V
Sbjct: 300 QEENEV 305
>RH1A_ARATH (Q9SUS4) RING-H2 zinc finger protein RHA1a
Length = 159
Score = 53.5 bits (127), Expect = 9e-07
Identities = 27/72 (37%), Positives = 39/72 (53%), Gaps = 4/72 (5%)
Query: 173 AVELLPTIEINESHMNVESHCAVCKEPFELGISAREMP-CKHIYHNECILPWLAIQN--S 229
A EL+P + ++ + E C VC FE R++P C H++H+ C+ W+ N
Sbjct: 66 ANELIPVVRFSDLPTDPEDCCTVCLSDFESDDKVRQLPKCGHVFHHYCLDRWIVDYNKMK 125
Query: 230 CPVCRHE-LPCE 240
CPVCRH LP E
Sbjct: 126 CPVCRHRFLPKE 137
>AMFR_MOUSE (Q9R049) Autocrine motility factor receptor (EC
6.3.2.19) (AMF receptor)
Length = 643
Score = 53.5 bits (127), Expect = 9e-07
Identities = 23/66 (34%), Positives = 36/66 (53%), Gaps = 4/66 (6%)
Query: 193 CAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCRHELP-CESPQINNEISNS 251
CA+C + + +AR++PC H++HN C+ WL SCP CR L + + +
Sbjct: 341 CAICWDSMQ---AARKLPCGHLFHNSCLRSWLEQDTSCPTCRMSLNIADGSRAREDHQGE 397
Query: 252 NEDENV 257
N DEN+
Sbjct: 398 NLDENL 403
>RN24_HUMAN (Q9Y225) RING finger protein 24
Length = 148
Score = 51.6 bits (122), Expect = 3e-06
Identities = 22/51 (43%), Positives = 31/51 (60%), Gaps = 8/51 (15%)
Query: 187 MNVESHCAVCKEPF----ELGISAREMPCKHIYHNECILPWLAIQNSCPVC 233
+N+ CAVC E F ELGI PCKH +H +C++ WL ++ CP+C
Sbjct: 72 LNLHELCAVCLEDFKPRDELGIC----PCKHAFHRKCLIKWLEVRKVCPLC 118
>R103_RAT (Q9EPZ8) RING finger protein 103 (Zinc finger protein 103)
(Zfp-103) (ADRG34 protein)
Length = 682
Score = 50.4 bits (119), Expect = 8e-06
Identities = 21/45 (46%), Positives = 28/45 (61%), Gaps = 1/45 (2%)
Query: 191 SHCAVCKEPFELGISAREMPCKHIYHNECILPWLA-IQNSCPVCR 234
+ C VC E FE G +PC H++H CI+ WLA ++ CPVCR
Sbjct: 616 TECVVCLENFENGCLLMGLPCGHVFHQNCIVMWLAGGRHCCPVCR 660
>R103_MOUSE (Q9R1W3) RING finger protein 103 (Zinc finger protein
103) (Zfp-103) (KF-1) (mKF-1)
Length = 683
Score = 50.4 bits (119), Expect = 8e-06
Identities = 21/45 (46%), Positives = 28/45 (61%), Gaps = 1/45 (2%)
Query: 191 SHCAVCKEPFELGISAREMPCKHIYHNECILPWLA-IQNSCPVCR 234
+ C VC E FE G +PC H++H CI+ WLA ++ CPVCR
Sbjct: 617 TECVVCLENFENGCLLMGLPCGHVFHQNCIVMWLAGGRHCCPVCR 661
>R103_HUMAN (O00237) RING finger protein 103 (Zinc finger protein
103 homolog) (Zfp-103) (KF-1) (hKF-1)
Length = 685
Score = 50.4 bits (119), Expect = 8e-06
Identities = 21/45 (46%), Positives = 28/45 (61%), Gaps = 1/45 (2%)
Query: 191 SHCAVCKEPFELGISAREMPCKHIYHNECILPWLA-IQNSCPVCR 234
+ C VC E FE G +PC H++H CI+ WLA ++ CPVCR
Sbjct: 619 TECVVCLENFENGCLLMGLPCGHVFHQNCIVMWLAGGRHCCPVCR 663
>GOLI_DROME (Q06003) Goliath protein (Protein g1)
Length = 286
Score = 50.4 bits (119), Expect = 8e-06
Identities = 18/42 (42%), Positives = 27/42 (63%)
Query: 193 CAVCKEPFELGISAREMPCKHIYHNECILPWLAIQNSCPVCR 234
CA+C E ++ + R +PCKH +H CI PWL +CP+C+
Sbjct: 128 CAICIEAYKPTDTIRILPCKHEFHKNCIDPWLIEHRTCPMCK 169
>RN12_HUMAN (Q9NVW2) RING finger protein 12 (LIM domain interacting
RING finger protein) (RING finger LIM domain-binding
protein) (R-LIM) (NY-REN-43 antigen)
Length = 624
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/76 (31%), Positives = 37/76 (48%), Gaps = 2/76 (2%)
Query: 159 NEGHNQQHLPALKSAVELLPTIEINESHMNVESHCAVCKEPFELGISAREMPCKHIYHNE 218
NE + Q K ++ L E+ + C+VC + G R++PC H YH
Sbjct: 538 NEDDDDQPRGLTKEQIDNLAMRSFGEN--DALKTCSVCITEYTEGNKLRKLPCSHEYHVH 595
Query: 219 CILPWLAIQNSCPVCR 234
CI WL+ ++CP+CR
Sbjct: 596 CIDRWLSENSTCPICR 611
>RNF6_HUMAN (Q9Y252) RING finger protein 6 (RING-H2 protein)
Length = 685
Score = 48.9 bits (115), Expect = 2e-05
Identities = 28/96 (29%), Positives = 45/96 (46%), Gaps = 9/96 (9%)
Query: 159 NEGHNQQHLPAL-KSAVELLPT--IEINESHMNVESHCAVCKEPFELGISAREMPCKHIY 215
NE + + L K ++ L T E N + C+VC + G R++PC H +
Sbjct: 595 NESDDDDRIRGLTKEQIDNLSTRHYEHNSIDSELGKICSVCISDYVTGNKLRQLPCMHEF 654
Query: 216 HNECILPWLAIQNSCPVCRHELPCESPQINNEISNS 251
H CI WL+ +CP+CR P + + I+N+
Sbjct: 655 HIHCIDRWLSENCTCPICR------QPVLGSNIANN 684
>YOF7_SCHPO (Q9P7E1) RING finger protein P4H10.07 in chromosome II
Length = 583
Score = 48.5 bits (114), Expect = 3e-05
Identities = 22/47 (46%), Positives = 27/47 (56%), Gaps = 2/47 (4%)
Query: 190 ESHCAVCKEPFELGISAREMP-CKHIYHNECILPWL-AIQNSCPVCR 234
+ C VC FEL R + C H +H ECI WL + QNSCP+CR
Sbjct: 522 DERCLVCLSNFELNDECRRLKQCNHFFHRECIDQWLTSSQNSCPLCR 568
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 52,128,483
Number of Sequences: 164201
Number of extensions: 2460230
Number of successful extensions: 6766
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 54
Number of HSP's successfully gapped in prelim test: 61
Number of HSP's that attempted gapping in prelim test: 6677
Number of HSP's gapped (non-prelim): 132
length of query: 391
length of database: 59,974,054
effective HSP length: 112
effective length of query: 279
effective length of database: 41,583,542
effective search space: 11601808218
effective search space used: 11601808218
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC124953.18


