

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124952.2 - phase: 0
(880 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
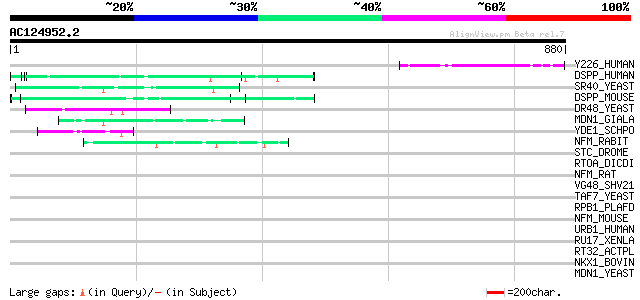
Score E
Sequences producing significant alignments: (bits) Value
Y226_HUMAN (Q92622) Hypothetical protein KIAA0226 (Fragment) 88 9e-17
DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contai... 67 2e-10
SR40_YEAST (P32583) Suppressor protein SRP40 59 7e-08
DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin... 57 2e-07
DR48_YEAST (P18899) DDR48 stress protein (DNA damage-responsive ... 47 2e-04
MDN1_GIALA (Q8T5T1) Midasin (MIDAS-containing protein) 47 2e-04
YDE1_SCHPO (Q10435) Probable ubiquitin fusion degradation protei... 45 6e-04
NFM_RABIT (P54938) Neurofilament triplet M protein (160 kDa neur... 45 6e-04
STC_DROME (P40798) Shuttle craft protein 44 0.001
RTOA_DICDI (P54681) Protein rtoA (Ratio-A) 44 0.002
NFM_RAT (P12839) Neurofilament triplet M protein (160 kDa neurof... 44 0.002
VG48_SHV21 (Q01033) Hypothetical gene 48 protein 43 0.003
TAF7_YEAST (Q05021) Transcription initiation factor TFIID subuni... 43 0.003
RPB1_PLAFD (P14248) DNA-directed RNA polymerase II largest subun... 43 0.003
NFM_MOUSE (P08553) Neurofilament triplet M protein (160 kDa neur... 43 0.003
URB1_HUMAN (Q7Z6Z7) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2... 42 0.005
RU17_XENLA (P09406) U1 small nuclear ribonucleoprotein 70 kDa (U... 42 0.007
RT32_ACTPL (P55131) RTX-III toxin determinant A from serotype 8 ... 42 0.009
NKX1_BOVIN (Q28139) Sodium/potassium/calcium exchanger 1 precurs... 42 0.009
MDN1_YEAST (Q12019) Midasin (MIDAS-containing protein) 42 0.009
>Y226_HUMAN (Q92622) Hypothetical protein KIAA0226 (Fragment)
Length = 1010
Score = 88.2 bits (217), Expect = 9e-17
Identities = 73/265 (27%), Positives = 115/265 (42%), Gaps = 23/265 (8%)
Query: 618 DSHPSSPVSNSPVSLSSFTRGENIRNSSTWGKTISLIVEIPSNKSTRQLLEA-QHHTCAG 676
DS P SP + + +R + W I+ TR++ A Q++ CAG
Sbjct: 704 DSLPISPDDGQHADI--YKLRIRVRGNLEWAPPRPQIIFNVHPAPTRKIAVAKQNYRCAG 761
Query: 677 CHRHFDDGNSSIWDFVQTFGWGKPRLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYP 736
C D D+++ R CEY G+ FC CH N +P+RVL WDF+ Y
Sbjct: 762 CGIRTDP------DYIKRL-----RYCEYLGKYFCQCCHENAQMAIPSRVLRKWDFSKYY 810
Query: 737 VSQLAKSYLDSIHEHPMLCVTAVNPFLVSKVPALLHVMSVRKKIGTML-PYVRCPFRRSI 795
VS +K L I P+ V +N L KV L V +R ++ M + C + +
Sbjct: 811 VSNFSKDLLIKIWNDPLFNVQDINSALYRKVKLLNQVRLLRVQLCHMKNMFKTCRLAKEL 870
Query: 796 NKGVGN-RRYLLESNDFFALRDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVG 854
+L E ++L DL KG P + E ++R H+ ++C++C G
Sbjct: 871 LDSFDTVPGHLTEDLHLYSLNDLTATRKGELG--PRLAE-LTRAGATHV-ERCMLCQAKG 926
Query: 855 IPCSARQDCSDPSSLIFPFQVCICQ 879
C + C + +IFPF++ C+
Sbjct: 927 FIC---EFCQNEDDIIFPFELHKCR 948
>DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contains:
Dentin phosphoprotein (Dentin phosphophoryn) (DPP);
Dentin sialoprotein (DSP)]
Length = 1253
Score = 67.4 bits (163), Expect = 2e-10
Identities = 92/467 (19%), Positives = 170/467 (35%), Gaps = 10/467 (2%)
Query: 24 SRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNRK 83
S+ S ++SDS+ S SNS + N+ S S + D SD +S S + DS+
Sbjct: 627 SKSESDSSDSDSKSDSSDSNSSDSSDNSDSSDSSNSSNSSDSSDSSDSSDSSSSSDSSSS 686
Query: 84 SFQYGSSGLELYGDEGDEL-SMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEE 142
S SS D + S DSS+ ++ + S+ N + + ++ +
Sbjct: 687 SDSSNSSDSSDSSDSSNSSESSDSSDSSDSDSSDSSDSSNSNSSDSDSSNSSDSSDSSDS 746
Query: 143 EEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMN 202
+ S DS + S D+ + S + S+ S+ S + + + N + N
Sbjct: 747 SDSSNSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSNDSSNSSDSSDSSNSSD-SSN 805
Query: 203 SSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVE 262
SS + S D D N S S + S++ + S + + +
Sbjct: 806 SSDSSDSSDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSS 865
Query: 263 ETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITD 322
++ D D + D D + ++ S + + +S D D S N +D
Sbjct: 866 DSSDSSDSSNSSDSSDSSDSSDSNESSNSSDSSDSSNSSDSDSSDSSNSSDSSDSSNSSD 925
Query: 323 GGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSN------VSVH 376
+ S + ++ S D+ + ++ + + N D S SN S
Sbjct: 926 SSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDSSNS 985
Query: 377 VENVDAKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVISKIEDSELSEFYDEVVQ 436
++ D+ + SN SS S+N + S+ + S+ S+ D
Sbjct: 986 SDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDS 1045
Query: 437 EMEEILLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSSTDDA 483
+S DS + S G+ D S S D ++ +S + D+
Sbjct: 1046 SNSSDSSDSSDSSDSSDSSGSS--DSSDSSDSSDSSDSSDSSDSSDS 1090
Score = 60.5 bits (145), Expect = 2e-08
Identities = 92/466 (19%), Positives = 162/466 (34%), Gaps = 27/466 (5%)
Query: 20 SDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDD 79
S S S SDS ++S+S + ++ S S + D SD D ++SS D
Sbjct: 770 SSDSSDSSDSSNSSDSNDSSNSSDSSDSSNSSDSSNSSDSSDSSDSSDSDSSNSS----D 825
Query: 80 SNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEE 139
S+ S SS D D + DSS ++ +S + + + ++
Sbjct: 826 SSNSSDSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSSDSSDSSDSSNSSDSSDSSDS 885
Query: 140 EEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPL 199
+ E S DS D N ++ S N S SS S E + +
Sbjct: 886 SDSNESSNSSDSS-------DSSNSSDSDSSDSSNSSDSSDSSNSSDSSESSNSSDNS-- 936
Query: 200 LMNSSVAFGSHDLDDF--LLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIV 257
NSS + S D D + S D + S++ + SS + DS
Sbjct: 937 --NSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDSSNSSDSSDSSDS 994
Query: 258 NDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKS 317
+D + + ++ + D D + S +D+S D D S
Sbjct: 995 SDSSDSSDSSNSSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDS 1054
Query: 318 LNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHV 377
+ +D + S + ++ S D+ + D+ + S + D+ D +S+ S
Sbjct: 1055 SDSSDSSDSSGSSDSSDSSDSSDSSDSSDSSDSSDSSD--SSESSDSSDSSDSSDSSDSS 1112
Query: 378 ENVDAKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVISKIEDSELSEFYDEVVQE 437
++ D+ + SN SS S++ + S + S+ S+ D
Sbjct: 1113 DSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSS 1172
Query: 438 MEEILLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSSTDDA 483
+S DS N D S S D ++ +S + D+
Sbjct: 1173 DSSDSSDSSDS--------NESSDSSDSSDSSDSSNSSDSSDSSDS 1210
Score = 58.5 bits (140), Expect = 7e-08
Identities = 89/462 (19%), Positives = 166/462 (35%), Gaps = 14/462 (3%)
Query: 24 SRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNRK 83
S SS ++SDS S+S+S ++S S S + D S+ +S S DS+
Sbjct: 560 SSDSSDSSDSDSSDSNSSSDSDSSDSDSSDSSDSDSS---DSSNSSDSSDSSDSSDSSDS 616
Query: 84 SFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEE 143
S S + S + DSS+ ++ + SD +++ + + +
Sbjct: 617 SDSSDSKSDSSKSESDSSDSDSKSDSSDSNSSDSSDNSDSSDSSNSSNSSDSSDSSDSSD 676
Query: 144 EFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNS 203
S D S + S +++ S E SS+ S + + N N +S
Sbjct: 677 SSSSSDSSSSSDSSNSSDSSDSSDSSNSSE--SSDSSDSSDSDSSDSSDSSNSNSSDSDS 734
Query: 204 SVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEE 263
S + S D D + S SD + SN+ S + + +++
Sbjct: 735 SNSSDSSDSSD----SSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSNDSSN 790
Query: 264 TKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDG 323
+ D D + + D D+ S +++S D D S + +D
Sbjct: 791 SSDSSDSSNSSDSSNSSDSSDSSDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSSDSSDS 850
Query: 324 GSEGKGNR--YSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVD 381
NR S + ++ S D+ + D+ + N D S S+ S ++ D
Sbjct: 851 SDSDSSNRSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSNESSNSSDSSDSSNSSDSDSSD 910
Query: 382 AKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVISKIEDSELSEFYDEVVQEMEEI 441
+ + + SN SS ++N + S+ + S+ S D
Sbjct: 911 SSNSSDSSD---SSNSSDSSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSN 967
Query: 442 LLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSSTDDA 483
+S DS ++ S + D S S D ++ +S++ D+
Sbjct: 968 SSDSSDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDS 1009
Score = 57.4 bits (137), Expect = 2e-07
Identities = 92/460 (20%), Positives = 159/460 (34%), Gaps = 30/460 (6%)
Query: 24 SRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNRK 83
S SS ++SDS +SNS + +++ S D SD +S S D SNR
Sbjct: 809 SSDSSDSSDSDSSNSSDSSNSSDSSDSSNSS---------DSSDSSDSSDSSDSDSSNRS 859
Query: 84 SFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEE 143
S + S DSS+ ++ +S + N + + + +
Sbjct: 860 DSSNSSDSSDSSDSSNSSDSSDSSDSSDSNESSNSSDSSDSSNSSDSDSSDSSNSSDSSD 919
Query: 144 EFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNS 203
+ D S S DN N S + S SS S G+ N + +S
Sbjct: 920 SSNSSDSS----ESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSS----DS 971
Query: 204 SVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVND--EV 261
S + S D + S S + +S+N + SS + DS +D
Sbjct: 972 SDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSNSSD----SSDSSDSSDSSNSSDSSNS 1027
Query: 262 EETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNIT 321
++ D D + D D D+ S +D+S D D S + +
Sbjct: 1028 SDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDS-----SDSSGSSDSSDSSDSSDSSDSS 1082
Query: 322 DGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVD 381
D + S++ ++ S D+ + D+ + S D+ D +SN S ++ D
Sbjct: 1083 DSSDSSDSSDSSESSDSSDSSDSSDSSDSSDSSD--SSDSSDSSDSSDSSNSSDSSDSSD 1140
Query: 382 AKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVISKIEDSELSEFYDEVVQEMEEI 441
+ + S+ SS S++ + S + +E S+ D
Sbjct: 1141 SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNESSDSSDSSDSSDSSN 1200
Query: 442 LLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSSTD 481
+S DS + S + + S +G S S +D
Sbjct: 1201 SSDSSDSSDSSDSTSDSNDESDSQSKSGNGNNNGSDSDSD 1240
Score = 54.7 bits (130), Expect = 1e-06
Identities = 91/471 (19%), Positives = 161/471 (33%), Gaps = 36/471 (7%)
Query: 20 SDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFD- 78
SD S SS SDS +S+S + ++ S S S+ D + S D
Sbjct: 583 SDSDSSDSSDSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSKSDSSKSESDSSDSDSKSDS 642
Query: 79 -DSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIET 137
DSN S + S DSS+ ++ S + N + + +
Sbjct: 643 SDSNSSDSSDNSDSSDSSNSSNSSDSSDSSDSSDSSSSSDSSSSSDSSNSSDSSDSSDSS 702
Query: 138 EEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNEN 197
E + S+ DS + S N ++ S + S SS S + + + + +
Sbjct: 703 NSSESSDSSDSSDSDSSDSSDSSNSNSSDSDSSNSSDSSDSSDSSDSSNSSDSSDSSDSS 762
Query: 198 PLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIV 257
NSS + S D D S N +SN+ SS + DS
Sbjct: 763 ----NSSDSSDSSDSSD-----------SSDSSNSSDSND-------SSNSSDSSDS--- 797
Query: 258 NDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQK- 316
++ + D + D D D + S +D+S D D
Sbjct: 798 -SNSSDSSNSSDSSDSSDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSSDSSDSSDSDSS 856
Query: 317 ----SLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSN 372
S N +D + S + ++ S D+ + D+ + + D+ D ++S+
Sbjct: 857 NRSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSNESSNSSDSSDSSNSSDSDSSDSSNSSD 916
Query: 373 VSVHVENVDAKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVISKIEDSELSEFYD 432
S + D+ N S+ SS S+N + S + + S S+ D
Sbjct: 917 SSDSSNSSDSSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSS---NSGDSSNSSDSSD 973
Query: 433 EVVQEMEEILLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSSTDDA 483
+ + S S ++ S + D S S D ++ +S++ D+
Sbjct: 974 SNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSNSSDS 1024
Score = 53.9 bits (128), Expect = 2e-06
Identities = 90/466 (19%), Positives = 166/466 (35%), Gaps = 31/466 (6%)
Query: 27 SSFGAESDSERYCSAS---NSMMGT-PNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNR 82
S+ G++S+S+ Y S SM G PN+S + + D A+S + S+R
Sbjct: 439 SNTGSDSNSDGYDSYDFDDKSMQGDDPNSS----------DESNGNDDANSESDNNSSSR 488
Query: 83 KSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEE 142
Y S + G+ D DS N + + NG + ++ + + +
Sbjct: 489 GDASYNSDESKDNGNGSDSKGAEDDDSDSTSDTNNSDSNGNGNNGNDDNDKSDSGKGKSD 548
Query: 143 EEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMN 202
S+ DS + S D+++ + ++ SS+ SS S +++ N
Sbjct: 549 SSDSDSSDSSNSSDSSDSSDSDSSDSNSSSDSDSSDSDSSDSSD--------SDSSDSSN 600
Query: 203 SSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVE 262
SS + S D D + SD + ES++ + S DS +D +
Sbjct: 601 SSDSSDSSDSSDSSDSSDSSDSKSD--SSKSESDSSDSDSKSDSSDSNSSDSSDNSDSSD 658
Query: 263 ETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITD 322
+ ++ + D D + S +D+S + D S +
Sbjct: 659 SSNSSNSSDSSDSSDSSDSSSSSDSSSSSDSSNSSDSSDSSDSSNSSESSDSSDSSDSDS 718
Query: 323 GGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDA 382
S N S + ++ S D+ + D+ S D+ D ++S+ S ++ D+
Sbjct: 719 SDSSDSSNSNSSDSDSSNSSDSSDSSDSSDS--SNSSDSSDSSDSSNSSDSSDSSDSSDS 776
Query: 383 KSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVISKIE-----DSELSEFYDEVVQE 437
N SN SS S+N + S + S+ S D
Sbjct: 777 SDSSNSSDSNDSSNSSDSSDSSNSSDSSNSSDSSDSSDSSDSDSSNSSDSSNSSDSSDSS 836
Query: 438 MEEILLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSSTDDA 483
+S DS + S + D S S D ++++S + D+
Sbjct: 837 NSSDSSDSSDSSDSSDSDSSNRSDSSNSSDSSDSSDSSNSSDSSDS 882
Score = 46.6 bits (109), Expect = 3e-04
Identities = 75/366 (20%), Positives = 126/366 (33%), Gaps = 14/366 (3%)
Query: 2 NGSPDPPLDSFPRLRLHQSDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTV 61
N S DS S S S SD+ +SNS + ++ S S +
Sbjct: 902 NSSDSDSSDSSNSSDSSDSSNSSDSSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSN 961
Query: 62 FHDFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEES 121
D S+ +S S + D S+ + S + S DSS+ ++ S
Sbjct: 962 SGDSSNSSDSSDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSNS 1021
Query: 122 DGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYS 181
+ N + + ++ + + S DS + S D+ S + S S
Sbjct: 1022 SDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSGSSDSSDSSDSSDSSDS 1081
Query: 182 SRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEE 241
S S + + +E+ +SS + S D D + S SD + SN+ +
Sbjct: 1082 SDSSDSSDSSD-SSESSDSSDSSDSSDSSDSSD----SSDSSDSSDSSDSSDSSNS--SD 1134
Query: 242 HGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCAD 301
SS + DS +D ++ D D + D D D+ S + +D
Sbjct: 1135 SSDSSDSSDSSDSSDSSDS-SDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNESSD--SSD 1191
Query: 302 NSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQF 361
+S D D S + +D S+ ++D SG+ D D+ SD
Sbjct: 1192 SSDSSDSSNSSDSSDSSDSSDSTSDSN----DESDSQSKSGNGNNNGSDSDSDSEGSDSN 1247
Query: 362 CDNRDD 367
DD
Sbjct: 1248 HSTSDD 1253
Score = 34.3 bits (77), Expect = 1.5
Identities = 23/67 (34%), Positives = 35/67 (51%), Gaps = 3/67 (4%)
Query: 117 GIEESDGNENGGEVGEREIE-TEEEEEEEFSEGDDSMFNY--GSGCDGDNENEFYSMKGE 173
G++ SDG+ +G E E E + ++E+EE G DS N G D E++ S G+
Sbjct: 238 GLDNSDGSPSGNGADEDEDEGSGDDEDEEAGNGKDSSNNSKGQEGQDHGKEDDHDSSIGQ 297
Query: 174 NVSSEFY 180
N S+ Y
Sbjct: 298 NSDSKEY 304
>SR40_YEAST (P32583) Suppressor protein SRP40
Length = 406
Score = 58.5 bits (140), Expect = 7e-08
Identities = 81/366 (22%), Positives = 137/366 (37%), Gaps = 30/366 (8%)
Query: 9 LDSFPRLRLHQSDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDI 68
+D P+L + + + + SS + S S S+S+S + + S S+ + SD
Sbjct: 8 VDEVPKLSVKEKEIEEKSSSSSSSSSSSSSSSSSSSSSSSSSGESSSSSSSSSSSSSSDS 67
Query: 69 DFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGG 128
+S S + S+ S SS E D S SS ++ E S +E+
Sbjct: 68 SDSSDSESSSSSSSSSSSSSSSSDSESSSESDSSSSGSSSSS---SSSSDESSSESESED 124
Query: 129 EVGEREIETEEEEEEEFS------EGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSS 182
E +R E++ E+ +E E S + SG +E+E G S+ SS
Sbjct: 125 ETKKRARESDNEDAKETKKAKTEPESSSSSESSSSGSSSSSESE----SGSESDSDSSSS 180
Query: 183 RSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEH 242
S S E + +++ +SS + S D D S SD + +S++
Sbjct: 181 SSSSSDSESDSESDSQSSSSSSSSDSSSDSD---------SSSSD---SSSDSDSSSSSS 228
Query: 243 GVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADN 302
SS + DS +D + ++ + D D D D+ S E++ +
Sbjct: 229 SSSSDSDSDSDSSSDSDSSGSSDSSSSSDSSSDESTSSDSSDSDSDSDSGSSSELETKEA 288
Query: 303 SPDLDLLPEEDPQKSLNIT----DGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKS 358
+ D + EE P S T S K N + DE +D F++
Sbjct: 289 TAD-ESKAEETPASSNESTPSASSSSSANKLNIPAGTDEIKEGQRKHFSRVDRSKINFEA 347
Query: 359 DQFCDN 364
+ DN
Sbjct: 348 WELTDN 353
>DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin
matrix protein-3) (DMP-3) [Contains: Dentin
phosphoprotein (Dentin phosphophoryn) (DPP); Dentin
sialoprotein (DSP)]
Length = 934
Score = 57.0 bits (136), Expect = 2e-07
Identities = 92/481 (19%), Positives = 165/481 (34%), Gaps = 51/481 (10%)
Query: 3 GSPDPPLDSFPRLRLHQSDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVF 62
G D P + + H + G SR S + Y SM G S +
Sbjct: 407 GESDKPQGTAEKSAAHSNLGHSRIGSSSNSDGHDSYEFDDESMQGDDPKSSDESNG---- 462
Query: 63 HDFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESD 122
SD +S + +R Y S E D+ D S G D S ++S
Sbjct: 463 ---SDESDTNSESANESGSRGDASYTSD--ESSDDDNDSDSHAGEDDSS-------DDSS 510
Query: 123 GNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSS 182
G+ + G+ + E+E+++E + S+ D+S + D+ ++ + SS+ S
Sbjct: 511 GDGDSDSNGDGDSESEDKDESDSSDHDNSSDSESKSDSSDSSDDSSDSSDSSDSSDSSDS 570
Query: 183 RSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEH 242
S + +++ +SS + GS D D
Sbjct: 571 SDSSDSSDSSDSSDSNSSSDSSDSSGSSDSSD---------------------------- 602
Query: 243 GVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADN 302
SS + DS +D ++ D D + D D D+ S C+D+
Sbjct: 603 --SSDTCDSSDSSDSSDS-SDSSDSSDSSDSSDSSDSSDSSDSSSSSDS--SDSSSCSDS 657
Query: 303 SPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFC 362
S D D S + + S N +D + +S + + + S
Sbjct: 658 SDSSDSSDSSDSSDSSDSSSSDSSSSSNSSDSSDSSDSSSSSDSSDSSDSSDSSDSSGSS 717
Query: 363 DNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRKILERSSTSTNVLEKSHVISKI 422
D+ D ++S+ S ++ D+ S + S+ SS S++ + S
Sbjct: 718 DSSDSSASSDSSSSSDSSDSSSSSDSSDSSDSSDSSDSSESSDSSNSSDSSDSSDSSDSS 777
Query: 423 EDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSSTDD 482
+ S+ S+ D +S DS + S + D S S D ++ +S + D
Sbjct: 778 DSSDSSDSSDSSDSSNSSDSSDSSDSSDS--SDSSNSSDSSDSSDSSDSSDSSDSSDSSD 835
Query: 483 A 483
+
Sbjct: 836 S 836
Score = 50.4 bits (119), Expect = 2e-05
Identities = 70/349 (20%), Positives = 122/349 (34%), Gaps = 17/349 (4%)
Query: 2 NGSPDPPLDSFPRLRLHQSDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTV 61
+GS D DS S S S SDS +S+S + ++S S S +
Sbjct: 595 SGSSDSS-DSSDTCDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSSSSDSSDSSS 653
Query: 62 FHDFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEES 121
D SD +S S DS+ S SS S + DSS+ ++ +S
Sbjct: 654 CSDSSDSSDSSDSSDSSDSSDSSSSDSSSSSNSSDSSDSSDSSSSSDSSDSSDSSDSSDS 713
Query: 122 DGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYS 181
G+ + + + + + S DS + S D + +E + SS+
Sbjct: 714 SGSSDSSDSSASSDSSSSSDSSDSSSSSDSSDSSDSS-DSSDSSESSDSSNSSDSSDSSD 772
Query: 182 SRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEE 241
S S + +++ NSS + S D D S N +S++ +
Sbjct: 773 SSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSD-----------SSDSSNSSDSSDSSD- 820
Query: 242 HGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCAD 301
SS + DS +D + + ++ + D D + S +D
Sbjct: 821 ---SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSD 877
Query: 302 NSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLD 350
+S D + S + +DG S+ N ++ + D+ E D
Sbjct: 878 SSNSSDSSDSDSKDSSSDSSDGDSKSGNGNSDSNSDSNSDSDSDSEGSD 926
Score = 48.9 bits (115), Expect = 6e-05
Identities = 76/363 (20%), Positives = 129/363 (34%), Gaps = 22/363 (6%)
Query: 18 HQSDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDID---FASSS 74
+ S S S SDS C +S+S + ++ S S + D SD +SSS
Sbjct: 586 NSSSDSSDSSGSSDSSDSSDTCDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSSS 645
Query: 75 RTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGERE 134
DS+ S SS D D + DSS ++ +S + + + +
Sbjct: 646 SDSSDSSSCSDSSDSSDSSDSSDSSDSSDSSSSDSSSSSNSSDSSDSSDSSSSSDSSDSS 705
Query: 135 IETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVR 194
++ + S+ DS + S D+ + S + S SS S
Sbjct: 706 DSSDSSDSSGSSDSSDSSASSDSSSSSDSSDSSSSSDSSDSSDSSDSSDS---------- 755
Query: 195 NENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDS 254
+E+ NSS + S D D + S SD + SN+ + SS + DS
Sbjct: 756 SESSDSSNSSDSSDSSDSSD----SSDSSDSSDSSDSSDSSNS--SDSSDSSDSSDSSDS 809
Query: 255 LIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDP 314
+D + + ++ + D D + S +D+S D D
Sbjct: 810 SNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDS 869
Query: 315 QKSLNITDGGSEGKGNRYSKNDEAGAS--GDAQRENLDLD-NFEFKSDQFCDNRDDVSTS 371
S + +D + + D + S GD++ N + D N + SD D+ S
Sbjct: 870 SDSSDSSDSSNSSDSSDSDSKDSSSDSSDGDSKSGNGNSDSNSDSNSDSDSDSEGSDSNH 929
Query: 372 NVS 374
+ S
Sbjct: 930 STS 932
>DR48_YEAST (P18899) DDR48 stress protein (DNA damage-responsive
protein 48) (DDRP 48) (YP 75) (Flocculent specific
protein)
Length = 429
Score = 47.4 bits (111), Expect = 2e-04
Identities = 51/245 (20%), Positives = 102/245 (40%), Gaps = 19/245 (7%)
Query: 25 RRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSR---TFDDSN 81
++SS+G+ +D S +N G+ N + + +D + SS++ ++ +N
Sbjct: 125 KKSSYGSNNDDSYGSSNNNDSYGSNNNDSYGSNNNDSYGSNNDDSYGSSNKNKSSYGSNN 184
Query: 82 RKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEE 141
S YGS+ + YG + S G +++ G+N + N N +
Sbjct: 185 DDS--YGSNNDDSYGSSNKKKSSYGSSNNDSYGSNNDDSYGSNNNDSYGSNNDDSYGSSN 242
Query: 142 EEEFSEGDDSMFNYGSGCD----GDNENEFYSMKGENVSS--------EFYSSRSVSLYE 189
+++ S G ++ +YGS + G N ++ Y +N SS + SS + Y
Sbjct: 243 KKKSSYGSNNDDSYGSSNNNDSYGSNNDDSYGSSNKNKSSYGSSSNDDSYGSSNNDDSYG 302
Query: 190 EGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRK 249
+ ++ N+ ++GS++ D + N S + SNN + +G S+ +K
Sbjct: 303 SSN-KKKSSYGSNNDDSYGSNNDDSYGSSNKKKSSYGSSNNDSYGSNND-DSYGSSNKKK 360
Query: 250 EEKDS 254
S
Sbjct: 361 SSYGS 365
Score = 41.6 bits (96), Expect = 0.009
Identities = 30/142 (21%), Positives = 60/142 (42%), Gaps = 13/142 (9%)
Query: 26 RSSFGAESDSERYCSASNS-MMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNRKS 84
+SS+G+ S+ + Y S++N G+ N S + + D +++ ++ SN+K
Sbjct: 280 KSSYGSSSNDDSYGSSNNDDSYGSSNKKKSSYGS-----NNDDSYGSNNDDSYGSSNKKK 334
Query: 85 FQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEE 144
YGSS + YG D+ + G+N + + N G ++
Sbjct: 335 SSYGSSNNDSYGSNNDDSYGSSNKKKSSYGSNNDDSYGSSNNNDSYG-------SNNDDS 387
Query: 145 FSEGDDSMFNYGSGCDGDNENE 166
+ + + +YGS G + N+
Sbjct: 388 YGSSNRNKNSYGSSNYGSSNND 409
Score = 41.2 bits (95), Expect = 0.012
Identities = 36/177 (20%), Positives = 78/177 (43%), Gaps = 11/177 (6%)
Query: 71 ASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEV 130
++++ ++ +N S YGS+ + YG + S G ++ + G++ +S G+ N
Sbjct: 97 SNNNDSYGSNNNDS--YGSNNDDSYGSSNKKKSSYGSNNDDSYGSSNNNDSYGSNNNDSY 154
Query: 131 GEREIET-EEEEEEEFSEGDDSMFNYGSGCD---GDNENEFYSMKGENVSSEFYSSRSVS 186
G ++ ++ + + + +YGS D G N ++ Y + SS Y S +
Sbjct: 155 GSNNNDSYGSNNDDSYGSSNKNKSSYGSNNDDSYGSNNDDSYGSSNKKKSS--YGSSNND 212
Query: 187 LYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHG 243
Y N++ N++ ++GS++ D + N S + S+N + +G
Sbjct: 213 SYGS---NNDDSYGSNNNDSYGSNNDDSYGSSNKKKSSYGSNNDDSYGSSNNNDSYG 266
Score = 37.4 bits (85), Expect = 0.18
Identities = 43/172 (25%), Positives = 78/172 (45%), Gaps = 20/172 (11%)
Query: 70 FASSSRTFDDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGE 129
F+++S +DSN YGS+ + YG ++ G ++++ G+N +S G+ N
Sbjct: 63 FSNTSINDNDSNNND-SYGSNNNDSYGSNNND--SYGSNNNDSYGSNN-NDSYGSNNDDS 118
Query: 130 VGEREIETEEEEEEEFSEGDDSMFNYGSGCD----GDNENEFYSMKGENVSSEFYSSRSV 185
G + +++ S DDS YGS + G N N+ Y G N +++ Y S +
Sbjct: 119 YG----SSNKKKSSYGSNNDDS---YGSSNNNDSYGSNNNDSY---GSN-NNDSYGSNND 167
Query: 186 SLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNN 237
Y +N++ N+ ++GS++ D + N S + SNN
Sbjct: 168 DSYGSSN-KNKSSYGSNNDDSYGSNNDDSYGSSNKKKSSYGSSNNDSYGSNN 218
>MDN1_GIALA (Q8T5T1) Midasin (MIDAS-containing protein)
Length = 4835
Score = 47.0 bits (110), Expect = 2e-04
Identities = 70/310 (22%), Positives = 123/310 (39%), Gaps = 40/310 (12%)
Query: 78 DDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIET 137
D++ +G L+ D +E S G D E++ E+ G+E G + ER
Sbjct: 4088 DEAIEMQNDFGGMSESLHHDIEEEDS-NGSDEEEVL-----EKEMGDEQGEAIDERNYNE 4141
Query: 138 EEEEEEEFSE--------GDDSMFNYGSGCDGDNEN-EFYSMKGENVSSEFYSSRSVSLY 188
+++ EE+S+ +D+M D D+ N E GE + ++ S
Sbjct: 4142 DDDSAEEYSDEQKKNTNNNNDTMELAAQEKDQDSINSELSDPSGEQHEEQADATGSTDEQ 4201
Query: 189 EEGEVRN---ENPLLMNSSVAFGSHDLDDFLLQ---NGPVSVVSDLFYNPRESNNRVEEH 242
+ + N + L S ++ D +D + +SDL NP + +EE
Sbjct: 4202 AQEDDYNDLDDKNLSGQSDLSVPEADGEDETVNEELEEEQQQMSDL-SNPDQDACAIEED 4260
Query: 243 GVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCEVQCADN 302
+ ++++ +DE E DI D EA +E + D DRD +S + Q ++
Sbjct: 4261 DDRDLPSSDENA-EEHDEHEAPVDIDDNEASDE---QSTYNDNDRDDAINISAQQQATND 4316
Query: 303 SPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFC 362
++ E D + NITD N +A G ++ DN +F+ +
Sbjct: 4317 EEEMQKDTEYDQE---NITD-----------SNPDANEVGTNDQKQTHEDNDQFRQENIE 4362
Query: 363 DNRDDVSTSN 372
D + ST N
Sbjct: 4363 DQWEAESTEN 4372
>YDE1_SCHPO (Q10435) Probable ubiquitin fusion degradation protein
C12B10.01c
Length = 1647
Score = 45.4 bits (106), Expect = 6e-04
Identities = 41/163 (25%), Positives = 69/163 (42%), Gaps = 25/163 (15%)
Query: 44 SMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGDELS 103
S+M N + + +S +T F + + + S + ++ S QY +Y D D
Sbjct: 165 SVMEPKNITFNDKSGLTKFGNSNSYETTYSDSSNYHTSTDSSQYNDQDDHVYDDTND--- 221
Query: 104 MTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDN 163
G D NN + D + +G E ER+ + +EEEEE+ E +D D ++
Sbjct: 222 --GTDDDI---NNNEYKDDESSSGSESYERDEDVDEEEEEDDDENNDE-------GDDED 269
Query: 164 ENEFYSMKGENVS----------SEFYSSRSVSLYEEGEVRNE 196
ENE ++ EN S E + + Y+E + NE
Sbjct: 270 ENENDELRSENGSPNPSAAVINEQENMNMSGIEEYDETNLLNE 312
>NFM_RABIT (P54938) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M) (Fragment)
Length = 644
Score = 45.4 bits (106), Expect = 6e-04
Identities = 68/344 (19%), Positives = 140/344 (39%), Gaps = 42/344 (12%)
Query: 118 IEESDGNENGGEVGEREIETEEEEEEEFSEG--DDSMFNYGSGCDGDNENEFYSMKGENV 175
I+E +G E+E E +EEEEEE EG D GS +G ++NE +GE
Sbjct: 307 IKEEEG--------EKEEEGQEEEEEEEDEGVKSDQAEEGGSEKEGSSKNEGEQEEGETE 358
Query: 176 SSEFYSSRSVSLYEEGEVRNENPLLMNSSVA---FGSHDLDDFLLQNGPVSVVSDLFY-- 230
+ ++ E ++E V G + + PV V
Sbjct: 359 AEGEVEEAEAKEEKKTEEKSEEVAAKEEPVTEAKVGKPEKAKSPVPKSPVEEVKPKAEAT 418
Query: 231 ----NPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRD 286
+E +VEE + ++ K+ + E E+ KD+ ++A V++ +
Sbjct: 419 AGKGEQKEEEEKVEEEKKKAAKESPKEEKVEKKE-EKPKDVPKKKAESPVKEEAAEEAAT 477
Query: 287 RDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSE----GKGNRYSKNDEAGASG 342
+ V E + + L +++ +K +GGSE +G++ +K ++ +G
Sbjct: 478 ITKPTKVGLEKETKEGEKPL----QQEKEKEKAGEEGGSEEEGSDQGSKRAKKEDIAVNG 533
Query: 343 DAQ-RENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNLKPIVLPSNGGTRK 401
+ + +E + + E S + + V T+ + + + ++ + +V+ T+K
Sbjct: 534 EGEGKEEEEPETKEKGSGR--EEEKGVVTNGLDLSPADEKKGGDRSEEKVVV-----TKK 586
Query: 402 I----LERSSTSTNVLEKSHVISKIEDSELSEFYDEVVQEMEEI 441
+ E +T + KS K+E+ E E ++E + +++
Sbjct: 587 VEKITTEGGDGATKYITKSVTAQKVEEHE--ETFEEKLVSTKKV 628
>STC_DROME (P40798) Shuttle craft protein
Length = 1106
Score = 44.3 bits (103), Expect = 0.001
Identities = 44/162 (27%), Positives = 61/162 (37%), Gaps = 32/162 (19%)
Query: 233 RESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDA- 291
R +NN+ H R+ E+D +D + RDRDRDRDRDRDRD+
Sbjct: 222 RGANNQCSNHNYERERERERD-----------RDRDRERDRDRDRDRDRDRDRDRDRDSR 270
Query: 292 PVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDL 351
P + Q + D D + +D K RY D R N
Sbjct: 271 PGNTRQQRRSDYRD-------DREDRYERSDRRRPQKQQRY----------DNHRSNKRR 313
Query: 352 DNFEFKSDQ---FCDNRDDVSTSNVSVHVENVDAKSFKNLKP 390
D++ D+ F DD+ TSN S H + + P
Sbjct: 314 DDWNRNRDRINGFPRAVDDLDTSNESAHPSPEKQSQLQQISP 355
>RTOA_DICDI (P54681) Protein rtoA (Ratio-A)
Length = 400
Score = 43.5 bits (101), Expect = 0.002
Identities = 44/165 (26%), Positives = 63/165 (37%), Gaps = 5/165 (3%)
Query: 24 SRRSSFGAESDSERYCSASNSMMGTPNTSM--SIRSAVTVFHDFSDIDFASSSRTFDDSN 81
S S+ G++S ++ S S G+ N+ S S+ + +D + S + SN
Sbjct: 125 SGSSNSGSQSSTDSSNSGSQGSTGSSNSGSQSSTDSSNSGSQSSTDSSNSGSQGSTGSSN 184
Query: 82 RKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEE 141
S GSS G E S +G +SS N+G E S G+ N G E
Sbjct: 185 SGSESSGSSNS---GSESSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSE 241
Query: 142 EEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVS 186
S S + GS G + S G SS +S S S
Sbjct: 242 SSSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSES 286
Score = 37.7 bits (86), Expect = 0.13
Identities = 43/169 (25%), Positives = 63/169 (36%), Gaps = 12/169 (7%)
Query: 25 RRSSFGAESDS--ERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNR 82
+ S G+ S S S+ S G+ +T+ S S + + S S S SN
Sbjct: 53 KNGSIGSSSQSLSSEVDSSDISSSGSNSTASSEGSVSSSSNSGSQSTSNSGSEASGSSNS 112
Query: 83 KSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGERE-----IET 137
S +SG E G S +G SS N+G + S G+ N G + ++
Sbjct: 113 GSQSTSNSGSEASGS-----SNSGSQSSTDSSNSGSQGSTGSSNSGSQSSTDSSNSGSQS 167
Query: 138 EEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVS 186
+ S+G N GS G + + S N SE S S S
Sbjct: 168 STDSSNSGSQGSTGSSNSGSESSGSSNSGSESSGSSNSGSESSSGSSNS 216
Score = 37.7 bits (86), Expect = 0.13
Identities = 50/217 (23%), Positives = 81/217 (37%), Gaps = 7/217 (3%)
Query: 20 SDGLSRRSSFGAESDSERYCSASNS--MMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTF 77
S+ S+ S+ + S S+ +SNS T +++ +S+ + S SS+
Sbjct: 128 SNSGSQSSTDSSNSGSQGSTGSSNSGSQSSTDSSNSGSQSSTDSSNSGSQGSTGSSNSGS 187
Query: 78 DDSNRKSFQYGSSGLELYGDEGDE-LSMTGLDSSELIGNNGIEESDGNENGGEVGEREIE 136
+ S + SSG G E S +G +SS N+G E S G+ N G
Sbjct: 188 ESSGSSNSGSESSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSS 247
Query: 137 TEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNE 196
E S S + GS G + S G SS +S S S + G +
Sbjct: 248 NSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSE-SSGSSNSGSESSSDSGSSSDG 306
Query: 197 NPLLMNSSVAFGSHDLDDFLLQ---NGPVSVVSDLFY 230
++ + +DD ++ G +SD Y
Sbjct: 307 KTTCISFHDTLSINTVDDDEIECTGKGETRCISDNNY 343
>NFM_RAT (P12839) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M)
Length = 845
Score = 43.5 bits (101), Expect = 0.002
Identities = 70/319 (21%), Positives = 120/319 (36%), Gaps = 35/319 (10%)
Query: 120 ESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEF 179
E+ E G + E E + EEEEE+E + D + +G +E E S K E E
Sbjct: 508 EAKEEEEGEKEEEEEGQEEEEEEDEGVKSDQAE-------EGGSEKEGSSEKDEGEQEEE 560
Query: 180 YSSRSVSLYEEGEVRNE-------NPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNP 232
+ + EE E + E + + + + + PV V
Sbjct: 561 GETEAEGEGEEAEAKEEKKTEGKVEEMAIKEEIKVEKPEKAKSPVPKSPVEEVKPKPEAK 620
Query: 233 RESNNRVEEHGV---SSVRKEEKDSLIVNDEVEETKDIGDREALEE-VRDRDRDRDRDRD 288
+ + EE V V KE V + E+ KD+ D++ E V+++ +
Sbjct: 621 AGKDEQKEEEKVEEKKEVAKESPKEEKVEKKEEKPKDVPDKKKAESPVKEKAVEEMITIT 680
Query: 289 RDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSE----GKGNRYSKNDEAGASGDA 344
+ VS E + P +++ K +GGSE K + SK ++ +G+
Sbjct: 681 KSVKVSLEKDTKEEKPQ-----QQEKVKEKAEEEGGSEEEVGDKSPQESKKEDIAINGEV 735
Query: 345 Q-RENLDLDNFEFKSDQ-----FCDNRDDVSTSNVSVHVENVDAKSF--KNLKPIVLPSN 396
+ +E + + E S Q N DVS + + D K K ++ I
Sbjct: 736 EGKEEEEQETQEKGSGQEEEKGVVTNGLDVSPAEEKKGEDRSDDKVVVTKKVEKITSEGG 795
Query: 397 GGTRKILERSSTSTNVLEK 415
G K + +S T T +E+
Sbjct: 796 DGATKYITKSVTVTQKVEE 814
>VG48_SHV21 (Q01033) Hypothetical gene 48 protein
Length = 797
Score = 43.1 bits (100), Expect = 0.003
Identities = 39/124 (31%), Positives = 53/124 (42%), Gaps = 12/124 (9%)
Query: 78 DDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIET 137
+D G E GDEGDE G D E G+ G +E D E+ G+ GE E +
Sbjct: 591 EDEGEDEGDEGEDEGEDEGDEGDEGEDEG-DEGEDEGDEGEDEGDEGEDEGDEGEDEGDE 649
Query: 138 EEEE----EEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEV 193
E+E E+E EGD+ G D +E E +GE+ E +EGE
Sbjct: 650 GEDEGDEGEDEGDEGDE-------GEDEGDEGEDEGDEGEDEGDEGDEGDEGDEGDEGED 702
Query: 194 RNEN 197
E+
Sbjct: 703 EGED 706
Score = 42.0 bits (97), Expect = 0.007
Identities = 24/70 (34%), Positives = 37/70 (52%), Gaps = 4/70 (5%)
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYG 156
DEGD+ G D E G + +E D + G + GE E + E+E E+E EGD+
Sbjct: 436 DEGDD----GEDEGEDEGEDEGDEGDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDEGD 491
Query: 157 SGCDGDNENE 166
G +G++E +
Sbjct: 492 EGDEGEDEGD 501
Score = 42.0 bits (97), Expect = 0.007
Identities = 31/101 (30%), Positives = 45/101 (43%), Gaps = 2/101 (1%)
Query: 78 DDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIET 137
+D + G G + DEGDE G D + + G +E D E+ GE + E
Sbjct: 544 EDEGDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGE--DEGDEG 601
Query: 138 EEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSE 178
E+E E+E EGD+ G D +E E +GE+ E
Sbjct: 602 EDEGEDEGDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDE 642
Score = 41.6 bits (96), Expect = 0.009
Identities = 29/79 (36%), Positives = 41/79 (51%), Gaps = 8/79 (10%)
Query: 96 GDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNY 155
GDEGDE G D + G+ G +E D + G E E E E E+E +E EGD+
Sbjct: 500 GDEGDE----GKDEGDE-GDEGKDEGDEGDEGDEGDEGEDEWEDEGDEGEDEGDE---GE 551
Query: 156 GSGCDGDNENEFYSMKGEN 174
G +G++E E +GE+
Sbjct: 552 DEGDEGEDEGEDEGDEGED 570
Score = 40.0 bits (92), Expect = 0.027
Identities = 31/96 (32%), Positives = 46/96 (47%), Gaps = 6/96 (6%)
Query: 79 DSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETE 138
D + G G E GDEG++ D E G+ G +E D E+ GE + E E
Sbjct: 514 DEGKDEGDEGDEGDE--GDEGEDEWEDEGDEGEDEGDEGEDEGDEGEDEGE--DEGDEGE 569
Query: 139 EEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGEN 174
+E E+E EG+D G +G++E E +GE+
Sbjct: 570 DEGEDEGDEGEDE--GEDEGDEGEDEGEDEGDEGED 603
Score = 40.0 bits (92), Expect = 0.027
Identities = 31/106 (29%), Positives = 46/106 (43%), Gaps = 9/106 (8%)
Query: 78 DDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEE-----SDGNENGGEVGE 132
D+ + G G + DEGDE G D E G+ G +E +G + G + G+
Sbjct: 555 DEGEDEGEDEGDEGEDEGEDEGDE----GEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGD 610
Query: 133 REIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSE 178
E E+E +E EGD+ G D +E E +GE+ E
Sbjct: 611 EGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDE 656
Score = 40.0 bits (92), Expect = 0.027
Identities = 36/145 (24%), Positives = 60/145 (40%), Gaps = 14/145 (9%)
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYG 156
DEGDE G D E G+ G +E D + G + G+ + +E E+E EGD+
Sbjct: 638 DEGDEGEDEG-DEGEDEGDEGEDEGDEGDEGEDEGDEGEDEGDEGEDEGDEGDEG----D 692
Query: 157 SGCDGDNENEFYSMKGENVSSEFYSSRSVSL---------YEEGEVRNENPLLMNSSVAF 207
G +GD + +GE+ E + + Y ++ ++ N S+
Sbjct: 693 EGDEGDEGEDEGEDEGEDEGDEGTKDKEGNANKVVQNPFDYYNWLQKSTLSIVHNKSIHA 752
Query: 208 GSHDLDDFLLQNGPVSVVSDLFYNP 232
H L+D L + S + L+ P
Sbjct: 753 TEHMLEDQLQHSLTASFETHLYIKP 777
Score = 38.9 bits (89), Expect = 0.060
Identities = 69/297 (23%), Positives = 103/297 (34%), Gaps = 28/297 (9%)
Query: 96 GDEGDELSMTGLDSS------ELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGD 149
GDEGDE G D E G+ G E D + G E GE E + +E ++E EGD
Sbjct: 456 GDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDEGDEGDE-GEDEGDEGDEGKDEGDEGD 514
Query: 150 DSMFNYGSGCDGDNENEFY-SMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFG 208
+ G +GD +E + E E +EGE E+
Sbjct: 515 EGKDEGDEGDEGDEGDEGEDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDE---GDEGEDE 571
Query: 209 SHDLDDFLLQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIG 268
D D G + + E+ G + E + DE +E +D G
Sbjct: 572 GEDEGDEGEDEG------------EDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEG 619
Query: 269 DREALEEVRDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGK 328
D E +E + + + D D E D D E++ + D G EG+
Sbjct: 620 D-EGEDEGDEGEDEGDEGEDEGDEGEDE---GDEGEDEGDEGEDEGDEGDEGEDEGDEGE 675
Query: 329 GNRYSKNDEAGASGDAQRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSF 385
DE G GD E + D E + + ++ D T + + V F
Sbjct: 676 DEGDEGEDE-GDEGDEGDEGDEGDEGEDEGEDEGEDEGDEGTKDKEGNANKVVQNPF 731
Score = 38.9 bits (89), Expect = 0.060
Identities = 26/101 (25%), Positives = 45/101 (43%), Gaps = 6/101 (5%)
Query: 78 DDSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIET 137
D+ + G G + DEGDE G D + + G +E D + G + G+ +
Sbjct: 566 DEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDE 625
Query: 138 EEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSE 178
+E E+E EG+D G +G++E + +G+ E
Sbjct: 626 GDEGEDEGDEGED------EGDEGEDEGDEGEDEGDEGEDE 660
Score = 38.9 bits (89), Expect = 0.060
Identities = 28/91 (30%), Positives = 43/91 (46%), Gaps = 4/91 (4%)
Query: 88 GSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSE 147
G E GDEG++ G D + + G +E D E+ GE + E E+E E+E E
Sbjct: 532 GEDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDEGDEGEDEGE--DEGDEGEDEGEDEGDE 589
Query: 148 GDDSMFNYGSGCDGDNENEFYSMKGENVSSE 178
G+D G +G++E E +G+ E
Sbjct: 590 GEDE--GEDEGDEGEDEGEDEGDEGDEGEDE 618
Score = 35.0 bits (79), Expect = 0.87
Identities = 74/322 (22%), Positives = 119/322 (35%), Gaps = 33/322 (10%)
Query: 52 SMSIRSAVTVFHDFSDIDFASSSRTFDD-SNRKSFQYGSSGLELYGDEGDELSMTGLDSS 110
S+++R +TV ++ ++ DD +R +Y EL +E + D
Sbjct: 369 SVALRQVLTVDKQANEKEYKKIIDKSDDRDDRDKDEY-----ELENEEYNR------DEE 417
Query: 111 ELIGNNGIEESDGNENGGEVGER-EIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYS 169
E G + +E D E G + G+ E E E+E E+E EGD+ G D D+E +
Sbjct: 418 EDEGEDEEDEKDEKEEGEDEGDDGEDEGEDEGEDEGDEGDEGDEGEDEGEDEDDEEDEGE 477
Query: 170 MKG-ENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDL 228
+G E E +EG+ +E D D G D
Sbjct: 478 DEGDEGDEGEDEGDEGDEGEDEGDEGDEGK---------DEGDEGDEGKDEGDEGDEGD- 527
Query: 229 FYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREA--LEEVRDRDRDRDRD 286
E + E+ G ++E D DE +E +D G+ E E+ + + D D
Sbjct: 528 --EGDEGEDEWEDEGDEG--EDEGDE--GEDEGDEGEDEGEDEGDEGEDEGEDEGDEGED 581
Query: 287 RDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQR 346
D E + D + + E++ + D G EG+ DE D
Sbjct: 582 EGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGD 641
Query: 347 ENLDL-DNFEFKSDQFCDNRDD 367
E D D E + D+ D D+
Sbjct: 642 EGEDEGDEGEDEGDEGEDEGDE 663
>TAF7_YEAST (Q05021) Transcription initiation factor TFIID subunit 7
(TBP-associated factor 7) (TBP-associated factor 67 kDa)
(TAFII-67)
Length = 590
Score = 43.1 bits (100), Expect = 0.003
Identities = 58/240 (24%), Positives = 96/240 (39%), Gaps = 33/240 (13%)
Query: 78 DDSNRKSFQYGSSGLELYGDE-GDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIE 136
D S ++ Q S E + +E G+ LS L E+ + I + G + E GE +E
Sbjct: 312 DKSELQARQERVSSWENFKEEPGEPLSRPALKKEEI---HTIASAVGKQGAEEEGEEGME 368
Query: 137 TEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSS---------------EFYS 181
EEEE+ + +S GSG +GD E + + G+ V E
Sbjct: 369 EEEEEDLDLGAAFESE-EEGSGAEGDKEQQQEEV-GDEVDQDTGGEDDDDDDDGDIEAAG 426
Query: 182 SRSVSLYEEGEVRNENPLL------MNSSVAFGSHDLDDF---LLQNGPVSVVSDLFYNP 232
S S E+ E R LL + +++A H L LL++ + + L
Sbjct: 427 GESESDDEKDENRQHTELLADELNELETTLAHTKHKLSKATNPLLKSRFIDSIKKLEKEA 486
Query: 233 RESNNRVE--EHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRD 290
+++ E V + D+ N+ EE ++ + E +EV D D + D + D D
Sbjct: 487 ELKRKQLQQTEDSVQKQHQHRSDAETANNVEEEEEEEEEEEEEDEV-DEDEEDDEENDED 545
>RPB1_PLAFD (P14248) DNA-directed RNA polymerase II largest subunit
(EC 2.7.7.6)
Length = 2452
Score = 43.1 bits (100), Expect = 0.003
Identities = 29/106 (27%), Positives = 56/106 (52%), Gaps = 6/106 (5%)
Query: 97 DEGDELSMTG---LDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMF 153
++G+E ++ G ++S L GN ++ D N + + EEEEEE+F GD ++
Sbjct: 1710 EDGNEGALRGGGDSNTSALFGNKNSQKEDNIVNNNN-DNNDDDDEEEEEEDFLFGDHNVS 1768
Query: 154 --NYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNEN 197
N G + + N+ + + +N S +S + + Y++G+V N+N
Sbjct: 1769 PKNTKDGKNKNTNNKSNNNENKNKKSGNNNSNNSNTYDDGDVDNDN 1814
>NFM_MOUSE (P08553) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M)
Length = 848
Score = 43.1 bits (100), Expect = 0.003
Identities = 67/321 (20%), Positives = 124/321 (37%), Gaps = 36/321 (11%)
Query: 120 ESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEF 179
E+ E GE E E EEEEEE+ D GS +G +E + +GE E
Sbjct: 508 EAKEEEEEGEKEEEEEGQEEEEEEDEGVKSDQAEEGGSEKEGSSEKD----EGEQEEEE- 562
Query: 180 YSSRSVSLYEEGEVRNENPL-------LMNSSVAFGSHDLDDFLLQNGPVSVV-----SD 227
+ + EE E + E + + + + + PV V +
Sbjct: 563 GETEAEGEGEEAEAKEEKKIEGKVEEVAVKEEIKVEKPEKAKSPMPKSPVEEVKPKPEAK 622
Query: 228 LFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEE-VRDRDRDRDRD 286
++ +VEE ++ K+ + E E+ KD+ D++ E V+++ +
Sbjct: 623 AGKGEQKEEEKVEEEKKEVTKESPKEEKVEKKE-EKPKDVADKKKAESPVKEKAVEEVIT 681
Query: 287 RDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGN----RYSKNDEAGASG 342
+ VS E + P P+E ++ +GGSE +G+ + SK ++ +G
Sbjct: 682 ISKSVKVSLEKDTKEEKPQ----PQEKVKEKAE-EEGGSEEEGSDRSPQESKKEDIAING 736
Query: 343 DA------QRENLDLDNFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSF--KNLKPIVLP 394
+ ++E + + + N DVS + ++ D K K ++ I
Sbjct: 737 EVEGKEEEEQETQEKGSGREEEKGVVTNGLDVSPAEEKKGEDSSDDKVVVTKKVEKITSE 796
Query: 395 SNGGTRKILERSSTSTNVLEK 415
G K + +S T T +E+
Sbjct: 797 GGDGATKYITKSVTVTQKVEE 817
>URB1_HUMAN (Q7Z6Z7) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-)
(HSPC272)
Length = 3360
Score = 42.4 bits (98), Expect = 0.005
Identities = 46/215 (21%), Positives = 89/215 (41%), Gaps = 11/215 (5%)
Query: 103 SMTGLDSSELIGNNGIEESDGNENG-GEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDG 161
S + + SE ++S N+ GE GE E++ E+ + + D + + + D
Sbjct: 1254 SASSKNKSEQDAQGASQDSSSNQQDPGEPGEAEVQEEDHDVTQTEVADGDIMDGEAETDS 1313
Query: 162 ---DNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQ 218
+ E S + V +E L +G N ++ S +D L+
Sbjct: 1314 VVIAGQPEVLSSQEMQVENELEDLIDELLERDGGSGNSTIIVSRSGE---DESQEDVLMD 1370
Query: 219 NGP--VSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEV 276
P +S S L N +S N ++ +EE DS N++ ++++D + E +E
Sbjct: 1371 EAPSNLSQASTLQANREDSMNILDPEDEEEHTQEE-DSSGSNEDEDDSQDEEEEEEEDEE 1429
Query: 277 RDRDRDRDRDRDR-DAPVSCEVQCADNSPDLDLLP 310
D++ D + D D E++ ++ PD++ P
Sbjct: 1430 DDQEDDEGEEGDEDDDDDGSEMELDEDYPDMNASP 1464
>RU17_XENLA (P09406) U1 small nuclear ribonucleoprotein 70 kDa (U1
snRNP 70 kDa) (snRNP70) (U1-70K)
Length = 471
Score = 42.0 bits (97), Expect = 0.007
Identities = 36/120 (30%), Positives = 55/120 (45%), Gaps = 10/120 (8%)
Query: 233 RESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAP 292
R+S +R +E R++ KD + E+ KD DR+ R R+R R+RDRDR+
Sbjct: 257 RKSRSREKEER-KRTREKSKDK-----DKEKDKDNKDRDRKRRSRSRERKRERDRDREKK 310
Query: 293 VSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLD 352
E + P+ D P++D Q ++ G E K K+ E D RE + D
Sbjct: 311 ---EERVEAEVPEADDAPQDDAQIG-DLGIDGIELKQEPEEKSRERDRERDRDREKGEKD 366
Score = 37.4 bits (85), Expect = 0.18
Identities = 22/66 (33%), Positives = 34/66 (51%), Gaps = 1/66 (1%)
Query: 225 VSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRD 284
+ DL + E EE R+ ++D + ++ +D DR+ RDRDR++D
Sbjct: 331 IGDLGIDGIELKQEPEEKSRERDRERDRDREKGEKDRDKDRD-RDRDRRRSHRDRDREKD 389
Query: 285 RDRDRD 290
RDRDRD
Sbjct: 390 RDRDRD 395
Score = 34.3 bits (77), Expect = 1.5
Identities = 22/64 (34%), Positives = 28/64 (43%)
Query: 227 DLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRD 286
+L P E + + K EKD D + + E+ RDRDRDR RD
Sbjct: 340 ELKQEPEEKSRERDRERDRDREKGEKDRDKDRDRDRDRRRSHRDRDREKDRDRDRDRRRD 399
Query: 287 RDRD 290
RDRD
Sbjct: 400 RDRD 403
Score = 33.1 bits (74), Expect = 3.3
Identities = 38/147 (25%), Positives = 58/147 (38%), Gaps = 21/147 (14%)
Query: 233 RESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKD-------IGDR-----EALEEVRDRD 280
R+ +R E R EK V EV E D IGD E +E ++
Sbjct: 290 RKRRSRSRERKRERDRDREKKEERVEAEVPEADDAPQDDAQIGDLGIDGIELKQEPEEKS 349
Query: 281 RDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKND---E 337
R+RDR+RDRD + + D D + D ++S D + +R + D +
Sbjct: 350 RERDRERDRDR------EKGEKDRDKDRDRDRDRRRSHRDRDREKDRDRDRDRRRDRDRD 403
Query: 338 AGASGDAQRENLDLDNFEFKSDQFCDN 364
D +RE D E + ++ DN
Sbjct: 404 RERDKDHKRERDRGDRSEKREERVPDN 430
Score = 32.3 bits (72), Expect = 5.6
Identities = 29/108 (26%), Positives = 44/108 (39%), Gaps = 8/108 (7%)
Query: 233 RESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAP 292
RE + + + +KD D +D + + RDR RDRDRDR+RD
Sbjct: 350 RERDRERDRDREKGEKDRDKDRDRDRDRRRSHRDRDREKDRDRDRDRRRDRDRDRERDKD 409
Query: 293 VSCEVQCADNS-------PDLDLLPEEDPQKSLNI-TDGGSEGKGNRY 332
E D S PD ++ E+ + S ++ D S G+ Y
Sbjct: 410 HKRERDRGDRSEKREERVPDNGMVMEQAEETSQDMYLDQESMQSGDGY 457
>RT32_ACTPL (P55131) RTX-III toxin determinant A from serotype 8
(APX-IIIA) (Cytolysin IIIA) (CLY-IIIA)
Length = 1052
Score = 41.6 bits (96), Expect = 0.009
Identities = 56/275 (20%), Positives = 107/275 (38%), Gaps = 45/275 (16%)
Query: 24 SRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDDSNRK 83
+++S+ G ++ Y + G N A H +I ++ F S +
Sbjct: 695 NQKSAVGKREETLEYRDYELTQSGNSNLK-----AHDELHSVEEIIGSNQRDEFKGSKFR 749
Query: 84 SFQYGSSGLELY-GDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEE 142
+G+ G +L G++GD++ + EL G+NG ++ GGE ++ +
Sbjct: 750 DIFHGADGDDLLNGNDGDDILYGDKGNDELRGDNGNDQL----YGGEGNDKLLGGNGNNY 805
Query: 143 EEFSEGDDSMFNYGSGCD----GDNENEFYSMKGENV--------------SSEFYSSRS 184
+G+D + G+G + G +++ Y G ++ S+FY RS
Sbjct: 806 LSGGDGNDELQVLGNGFNVLRGGKGDDKLYGSSGSDLLDGGEGNDYLEGGDGSDFYVYRS 865
Query: 185 VS----LYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPRESNNRVE 240
S +Y++G+ + + L L DF V V D SN
Sbjct: 866 TSGNHTIYDQGKSSDLDKLY-----------LSDFSFDRLLVEKVDDNLV--LRSNESSH 912
Query: 241 EHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEE 275
+GV +++ K+ N ++E+ D R+ E
Sbjct: 913 NNGVLTIKDWFKEGNKYNHKIEQIVDKNGRKLTAE 947
>NKX1_BOVIN (Q28139) Sodium/potassium/calcium exchanger 1 precursor
(Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod
Na-Ca+K exchanger)
Length = 1216
Score = 41.6 bits (96), Expect = 0.009
Identities = 39/141 (27%), Positives = 57/141 (39%), Gaps = 13/141 (9%)
Query: 64 DFSDIDFASSSRTFDDSNRKSFQYGSSGLELYGDEGD----ELSMTGLDSSELIGNNGIE 119
D +I + D + Q G G E+ GDEG+ E + E+ G
Sbjct: 882 DEGEIQAGEAGEVEGDEDEGEIQAGEGG-EVKGDEGEIQAGEAGEVEGEDGEVEGGEDEG 940
Query: 120 ESDGNENG-GEVGEREIETEEEEEEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSE 178
E E G GE GE+E+ E + E + + ++ + G G GD+E+E + E E
Sbjct: 941 EIQAGEGGEGETGEQELNAEIQGEAK--DDEEGVDGEGGGDGGDSEDEEEEDEEEEDEEE 998
Query: 179 FYSSRSVSLYEEGEVRNENPL 199
EE E NE PL
Sbjct: 999 EEEEE-----EEEEEENEQPL 1014
Score = 34.3 bits (77), Expect = 1.5
Identities = 54/231 (23%), Positives = 87/231 (37%), Gaps = 38/231 (16%)
Query: 79 DSNRKSFQYGSSGLELYGDEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETE 138
D + Q G G E+ GDE DE + + E+ G +E +G GE GE E
Sbjct: 812 DEDEGEIQAGEGG-EVEGDE-DEGEIQAGEGGEVEG----DEDEGEIQAGEAGEVE---G 862
Query: 139 EEEEEEFS-------EGDDSMFNYGSGCDG-----DNENEFYSMKGENVSSEFYSSRSVS 186
+E+E E EGD+ +G G ++E E + +G V + +
Sbjct: 863 DEDEGEIQAGEGGEVEGDEDEGEIQAGEAGEVEGDEDEGEIQAGEGGEVKGD---EGEIQ 919
Query: 187 LYEEGEVRNENPLLMNSSVAFGSHDLDDFLLQNGPVSVVSDLFYNPR-ESNNRVEEHGVS 245
E GEV E+ G D + G + N + + +E GV
Sbjct: 920 AGEAGEVEGED------GEVEGGEDEGEIQAGEGGEGETGEQELNAEIQGEAKDDEEGVD 973
Query: 246 SVRKEEKDSLIVNDEVEETKDIGDREALEEVRDRDRDRDRDRDRDAPVSCE 296
+ D DE EE D E +E + + + + + + + P+S E
Sbjct: 974 G--EGGGDGGDSEDEEEE-----DEEEEDEEEEEEEEEEEEEENEQPLSLE 1017
>MDN1_YEAST (Q12019) Midasin (MIDAS-containing protein)
Length = 4910
Score = 41.6 bits (96), Expect = 0.009
Identities = 62/277 (22%), Positives = 110/277 (39%), Gaps = 50/277 (18%)
Query: 97 DEGDELSMTGLDSSELIGNNGIEESDGNENGGEVGEREIETEEEEEEEFSEGDDSMFNYG 156
+E D + M G + EL + EE+D + E E + E ++ E++ + DD M++
Sbjct: 4104 NEDDAVEMEGDMAGELEDLSNGEENDDEDTDSEEEELDEEIDDLNEDDPNAIDDKMWDDK 4163
Query: 157 SGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEEGEVRNENPLLMNSSVAFGSHDLDDFL 216
+ + ++ ++ G+N V E E + +N D D
Sbjct: 4164 ASDNSKEKDTDQNLDGKN------QEEDVQAAENDEQQRDNK---------EGGDEDPNA 4208
Query: 217 LQNGPVSVVSDLFYNPRESNNRVEEHGVSSVRKEEKDSLIVNDEVEETKDIGDREALEEV 276
++G + +D N E N+ E+ V+ EE + L N ET D+ +
Sbjct: 4209 PEDGDEEIEND--ENAEEENDVGEQE--DEVKDEEGEDLEANVPEIETLDLPE------- 4257
Query: 277 RDRDRDRDRDRDRDAPVSCEVQCADNSPDLDLLPEEDPQKSLNI-TDGGSEGKGNRYSKN 335
D + D + + + +V +D PD DL EE + + + G E ++N
Sbjct: 4258 -DMNLDSEHEESDE-----DVDMSDGMPD-DLNKEEVGNEDEEVKQESGIESD----NEN 4306
Query: 336 DEAGASGDA------------QRENLDLDNFEFKSDQ 360
DE G DA E++D+ N E K D+
Sbjct: 4307 DEPGPEEDAGETETALDEEEGAEEDVDMTNDEGKEDE 4343
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.133 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 108,356,736
Number of Sequences: 164201
Number of extensions: 4962020
Number of successful extensions: 18344
Number of sequences better than 10.0: 258
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 196
Number of HSP's that attempted gapping in prelim test: 15658
Number of HSP's gapped (non-prelim): 1172
length of query: 880
length of database: 59,974,054
effective HSP length: 119
effective length of query: 761
effective length of database: 40,434,135
effective search space: 30770376735
effective search space used: 30770376735
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 70 (31.6 bits)
Medicago: description of AC124952.2


