

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124952.12 + phase: 2 /pseudo
(313 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
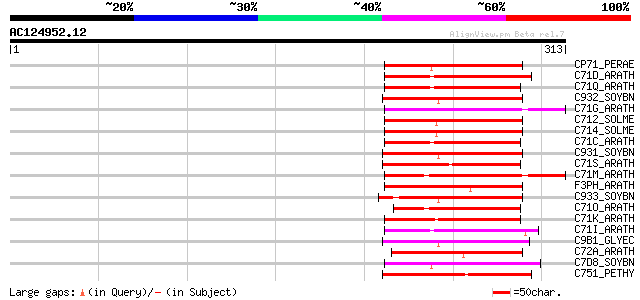
Score E
Sequences producing significant alignments: (bits) Value
CP71_PERAE (P24465) Cytochrome P450 71A1 (EC 1.14.-.-) (CYPLXXIA... 75 2e-13
C71D_ARATH (O49342) Cytochrome P450 71A13 (EC 1.14.-.-) 74 5e-13
C71Q_ARATH (Q9STK7) Cytochrome P450 71A26 (EC 1.14.-.-) 70 5e-12
C932_SOYBN (Q42799) Cytochrome P450 93A2 (EC 1.14.-.-) 69 1e-11
C71G_ARATH (Q9FH66) Cytochrome P450 71A16 (EC 1.14.-.-) 69 1e-11
C712_SOLME (P37118) Cytochrome P450 71A2 (EC 1.14.-.-) (CYPLXXIA... 69 1e-11
C714_SOLME (P37117) Cytochrome P450 71A4 (EC 1.14.-.-) (CYPLXXIA... 65 2e-10
C71C_ARATH (O49340) Cytochrome P450 71A12 (EC 1.14.-.-) 65 3e-10
C931_SOYBN (Q42798) Cytochrome P450 93A1 (EC 1.14.-.-) 64 4e-10
C71S_ARATH (P58047) Cytochrome P450 71A28 (EC 1.14.-.-) 64 6e-10
C71M_ARATH (Q9STL1) Cytochrome P450 71A22 (EC 1.14.-.-) 63 8e-10
F3PH_ARATH (Q9SD85) Flavonoid 3'-monooxygenase (EC 1.14.13.21) (... 62 2e-09
C933_SOYBN (O81973) Cytochrome P450 93A3 (EC 1.14.-.-) (P450 CP5) 62 2e-09
C71O_ARATH (Q9STK9) Cytochrome P450 71A24 (EC 1.14.-.-) 62 2e-09
C71K_ARATH (Q9T0K2) Cytochrome P450 71A20 (EC 1.14.-.-) 61 4e-09
C71I_ARATH (Q9SAB6) Cytochrome P450 71A18 (EC 1.14.-.-) 60 5e-09
C9B1_GLYEC (P93149) Cytochrome P450 93B1 (EC 1.14.-.-) ((2S)-fla... 60 7e-09
C72A_ARATH (Q9LVD2) Cytochrome P450 71B10 (EC 1.14.-.-) 60 7e-09
C7D8_SOYBN (O81974) Cytochrome P450 71D8 (EC 1.14.-.-) (P450 CP7) 60 9e-09
C751_PETHY (P48418) Flavonoid 3',5'-hydroxylase 1 (EC 1.14.-.-) ... 60 9e-09
>CP71_PERAE (P24465) Cytochrome P450 71A1 (EC 1.14.-.-) (CYPLXXIA1)
(ARP-2)
Length = 471
Score = 75.1 bits (183), Expect = 2e-13
Identities = 37/83 (44%), Positives = 57/83 (68%), Gaps = 5/83 (6%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRR-----DPKKKDFLDILLQLQDDGLTEFELTQNDLK 266
++K + E+D F+D VI +H +SR+ ++KD +D+LL LQ D L +N+LK
Sbjct: 236 RLKRNHGELDAFVDHVIDDHLLSRKANGSDGVEQKDLVDVLLHLQKDSSLGVHLNRNNLK 295
Query: 267 ALLMNMFLGGSDTTSTTLEWAMA 289
A++++MF GG+DTT+ TLEWAMA
Sbjct: 296 AVILDMFSGGTDTTAVTLEWAMA 318
>C71D_ARATH (O49342) Cytochrome P450 71A13 (EC 1.14.-.-)
Length = 497
Score = 73.9 bits (180), Expect = 5e-13
Identities = 35/83 (42%), Positives = 55/83 (66%), Gaps = 2/83 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
KIK+ S D +D+V+ EH + D K DF+DILL ++ D + F++ +ND+K ++++
Sbjct: 237 KIKEVSRGFSDLMDKVVQEHLEASND--KADFVDILLSIEKDKNSGFQVQRNDIKFMILD 294
Query: 272 MFLGGSDTTSTTLEWAMAACKKS 294
MF+GG+ TTST LEW M +S
Sbjct: 295 MFIGGTSTTSTLLEWTMTELIRS 317
>C71Q_ARATH (Q9STK7) Cytochrome P450 71A26 (EC 1.14.-.-)
Length = 489
Score = 70.5 bits (171), Expect = 5e-12
Identities = 31/77 (40%), Positives = 54/77 (69%), Gaps = 2/77 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
+++ ++ ++D F +RV+ +H RD DF+D+LL +Q D FE+ + +KA++MN
Sbjct: 230 QLEKTANDVDKFFERVVQDHVDGNRD--MTDFVDVLLAIQRDKTVGFEINRVSIKAIVMN 287
Query: 272 MFLGGSDTTSTTLEWAM 288
+F+GG+DT+ST +EWAM
Sbjct: 288 VFVGGTDTSSTLMEWAM 304
>C932_SOYBN (Q42799) Cytochrome P450 93A2 (EC 1.14.-.-)
Length = 502
Score = 69.3 bits (168), Expect = 1e-11
Identities = 33/86 (38%), Positives = 57/86 (65%), Gaps = 7/86 (8%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKK-------KDFLDILLQLQDDGLTEFELTQN 263
K+I+ + D LDR+I + + RR+ K+ KD LD+LL + +D +E +LT+
Sbjct: 228 KRIRKTRIRFDAVLDRIIKQREEERRNNKEIGGTRQFKDILDVLLDIGEDDSSEIKLTKE 287
Query: 264 DLKALLMNMFLGGSDTTSTTLEWAMA 289
++KA +M++F+ G+DT++ T+EWAMA
Sbjct: 288 NIKAFIMDIFVAGTDTSAATMEWAMA 313
>C71G_ARATH (Q9FH66) Cytochrome P450 71A16 (EC 1.14.-.-)
Length = 497
Score = 69.3 bits (168), Expect = 1e-11
Identities = 33/102 (32%), Positives = 61/102 (59%), Gaps = 3/102 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
K++ +++ DF+++V+ EH+ + D + DF+D+LL +Q D + +L ++DLK ++
Sbjct: 236 KLEKITKQFGDFIEKVLQEHEDTTADKETPDFVDMLLTIQRDETAQCQLDKSDLKVIIFE 295
Query: 272 MFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGINK 313
MFLG + TTS +EWAM + N E ++ R ++K
Sbjct: 296 MFLGSTTTTSAVIEWAMT---RLMRNPECLKKLQDEIRSVSK 334
>C712_SOLME (P37118) Cytochrome P450 71A2 (EC 1.14.-.-) (CYPLXXIA2)
(P-450EG4)
Length = 505
Score = 69.3 bits (168), Expect = 1e-11
Identities = 34/84 (40%), Positives = 56/84 (66%), Gaps = 6/84 (7%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPK------KKDFLDILLQLQDDGLTEFELTQNDL 265
K+K ++E+D FL+ VI EH + ++ + KDF+D+LL++Q+ T+F L ++ L
Sbjct: 238 KVKKVAKELDMFLEIVIEEHIIRKKKEEYTSTGEAKDFVDVLLEIQNGNETDFPLQRDSL 297
Query: 266 KALLMNMFLGGSDTTSTTLEWAMA 289
KA+L++ F G+DTT TL+W MA
Sbjct: 298 KAILLDSFAAGTDTTFATLDWTMA 321
>C714_SOLME (P37117) Cytochrome P450 71A4 (EC 1.14.-.-) (CYPLXXIA4)
(P-450EG2)
Length = 507
Score = 65.5 bits (158), Expect = 2e-10
Identities = 32/84 (38%), Positives = 54/84 (64%), Gaps = 6/84 (7%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPK------KKDFLDILLQLQDDGLTEFELTQNDL 265
K++ ++++D FLD VI EH + + + KDF+D+LL++Q+ T+F L ++ L
Sbjct: 239 KVEKIAKKLDTFLDSVIEEHIIRNKKEEYAITDEAKDFVDVLLEIQNGKETDFPLQRDSL 298
Query: 266 KALLMNMFLGGSDTTSTTLEWAMA 289
KA+L++ F G+DT T L+W MA
Sbjct: 299 KAILLDAFAAGTDTIYTNLDWTMA 322
>C71C_ARATH (O49340) Cytochrome P450 71A12 (EC 1.14.-.-)
Length = 497
Score = 64.7 bits (156), Expect = 3e-10
Identities = 29/77 (37%), Positives = 52/77 (66%), Gaps = 2/77 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
+IK+ S+ D +D+V+ EH + K+DF+DILL ++ + F+ ++D+K ++++
Sbjct: 237 RIKEVSQGFSDLMDKVVQEHLEAGNH--KEDFVDILLSIESEKSIGFQAQRDDIKFMILD 294
Query: 272 MFLGGSDTTSTTLEWAM 288
MF+GG+ T+ST LEW M
Sbjct: 295 MFIGGTSTSSTLLEWIM 311
>C931_SOYBN (Q42798) Cytochrome P450 93A1 (EC 1.14.-.-)
Length = 509
Score = 64.3 bits (155), Expect = 4e-10
Identities = 30/86 (34%), Positives = 55/86 (63%), Gaps = 7/86 (8%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKK-------KDFLDILLQLQDDGLTEFELTQN 263
+KIK++ + D +D +I + + RR K+ KD LD+LL + +D E +L +
Sbjct: 235 RKIKETRDRFDVVVDGIIKQRQEERRKNKETGTAKQFKDMLDVLLDMHEDENAEIKLDKK 294
Query: 264 DLKALLMNMFLGGSDTTSTTLEWAMA 289
++KA +M++F+ G+DT++ ++EWAMA
Sbjct: 295 NIKAFIMDIFVAGTDTSAVSIEWAMA 320
>C71S_ARATH (P58047) Cytochrome P450 71A28 (EC 1.14.-.-)
Length = 488
Score = 63.5 bits (153), Expect = 6e-10
Identities = 30/78 (38%), Positives = 53/78 (67%), Gaps = 1/78 (1%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
+K+++ S+ +FL+RV+ EH + + DF+DILL +Q + EFEL ++D++ +++
Sbjct: 238 RKMEEVSKTFVEFLERVVQEHVDEGENKETFDFVDILL-IQLEKTNEFELERSDIRLIIL 296
Query: 271 NMFLGGSDTTSTTLEWAM 288
FLGG+ TT T ++WAM
Sbjct: 297 EFFLGGTTTTFTAIDWAM 314
>C71M_ARATH (Q9STL1) Cytochrome P450 71A22 (EC 1.14.-.-)
Length = 490
Score = 63.2 bits (152), Expect = 8e-10
Identities = 31/102 (30%), Positives = 63/102 (61%), Gaps = 5/102 (4%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
++K + ++D+FL++V+ +H+ D ++ DF+D+LL++Q + FE+ + +KA++++
Sbjct: 231 QLKKTGNDLDEFLEKVVQDHEDG--DAQRTDFVDVLLRIQREKSVGFEIDRLSIKAIILD 288
Query: 272 MFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGINK 313
+ +GG+DT+ +EWAM + H E +E R I K
Sbjct: 289 VVVGGTDTSYALMEWAMT---ELLHRPECLNRLQEEVRTICK 327
>F3PH_ARATH (Q9SD85) Flavonoid 3'-monooxygenase (EC 1.14.13.21)
(Flavonoid 3'-hydroxylase) (AtF3'H) (Cytochrome P450
75B1) (TRANSPARENT TESTA 7 protein)
Length = 513
Score = 62.0 bits (149), Expect = 2e-09
Identities = 31/80 (38%), Positives = 50/80 (61%), Gaps = 2/80 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEF--ELTQNDLKALL 269
K+K + D FL ++ EH+M+ +D K D L L+ L+ L LT ++KALL
Sbjct: 237 KMKRLHKRFDAFLSSILKEHEMNGQDQKHTDMLSTLISLKGTDLDGDGGSLTDTEIKALL 296
Query: 270 MNMFLGGSDTTSTTLEWAMA 289
+NMF G+DT+++T++WA+A
Sbjct: 297 LNMFTAGTDTSASTVDWAIA 316
>C933_SOYBN (O81973) Cytochrome P450 93A3 (EC 1.14.-.-) (P450 CP5)
Length = 510
Score = 62.0 bits (149), Expect = 2e-09
Identities = 30/88 (34%), Positives = 55/88 (62%), Gaps = 10/88 (11%)
Query: 209 KLKKIKDSSEEMDDFLDRVIVEHKMSRRDPKK-------KDFLDILLQLQDDGLTEFELT 261
+L+KI+D D LDR+I + + RR+ + KD LD+L + +D +E +L
Sbjct: 237 RLEKIRDC---FDTVLDRIIKQREEERRNKNETVGKREFKDMLDVLFDISEDESSEIKLN 293
Query: 262 QNDLKALLMNMFLGGSDTTSTTLEWAMA 289
+ ++KA ++++ + G+DT++ T+EWAMA
Sbjct: 294 KENIKAFILDILIAGTDTSAVTMEWAMA 321
>C71O_ARATH (Q9STK9) Cytochrome P450 71A24 (EC 1.14.-.-)
Length = 486
Score = 62.0 bits (149), Expect = 2e-09
Identities = 29/72 (40%), Positives = 48/72 (66%), Gaps = 2/72 (2%)
Query: 217 SEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMNMFLGG 276
S ++D+FL+RV+ +H D K DF+D LL ++ + FE+ + +KA+++++F+G
Sbjct: 235 SNDLDEFLERVVQDHVDG--DGHKNDFVDFLLTIEREKSVGFEIDRLSIKAIILDVFVGD 292
Query: 277 SDTTSTTLEWAM 288
DTT T LEWAM
Sbjct: 293 MDTTYTLLEWAM 304
>C71K_ARATH (Q9T0K2) Cytochrome P450 71A20 (EC 1.14.-.-)
Length = 495
Score = 60.8 bits (146), Expect = 4e-09
Identities = 30/77 (38%), Positives = 51/77 (65%), Gaps = 1/77 (1%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
K++ + D+FL+RV+ EH+ + ++ + D +D LL +Q D +FEL ++ LK ++ +
Sbjct: 236 KMEVVDKRFDEFLERVVKEHEEADKETRS-DLVDKLLTIQSDKTGQFELEKSALKLIIWD 294
Query: 272 MFLGGSDTTSTTLEWAM 288
MFL G+ TT + LEWAM
Sbjct: 295 MFLAGTATTLSFLEWAM 311
>C71I_ARATH (Q9SAB6) Cytochrome P450 71A18 (EC 1.14.-.-)
Length = 497
Score = 60.5 bits (145), Expect = 5e-09
Identities = 32/93 (34%), Positives = 53/93 (56%), Gaps = 8/93 (8%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
KI + S D +++V+ EH + K DF++ILL ++ + F++ +ND+K ++++
Sbjct: 237 KIVEVSRAYSDLMEKVVQEHLEAGEH--KADFVNILLSIEKEKNNGFKVQRNDIKFMILD 294
Query: 272 MFLGGSDTTSTTLEWAMA------ACKKSSHNE 298
MF+GG T+ST LEW M C K NE
Sbjct: 295 MFIGGISTSSTLLEWIMTELIRNPECMKKLQNE 327
>C9B1_GLYEC (P93149) Cytochrome P450 93B1 (EC 1.14.-.-)
((2S)-flavanone 2-hydroxylase) (Licodione synthase)
(Flavone synthase II) (CYP GE-5)
Length = 523
Score = 60.1 bits (144), Expect = 7e-09
Identities = 30/96 (31%), Positives = 55/96 (57%), Gaps = 13/96 (13%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKK-------------KDFLDILLQLQDDGLTE 257
K+I+D + D ++R+I + + +R+D ++ +DFLDILL +D +E
Sbjct: 227 KRIEDLFQRFDTLVERIISKREQTRKDRRRNGKKGEQGSGDGIRDFLDILLDCTEDENSE 286
Query: 258 FELTQNDLKALLMNMFLGGSDTTSTTLEWAMAACKK 293
++ + +KAL+M+ F G+DTT+ + EWA+ K
Sbjct: 287 IKIQRVHIKALIMDFFTAGTDTTAISTEWALVELVK 322
>C72A_ARATH (Q9LVD2) Cytochrome P450 71B10 (EC 1.14.-.-)
Length = 502
Score = 60.1 bits (144), Expect = 7e-09
Identities = 30/76 (39%), Positives = 51/76 (66%), Gaps = 2/76 (2%)
Query: 216 SSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDG--LTEFELTQNDLKALLMNMF 273
S ++D F +++I H R+ + DF+D+LL+L+ + L +LT+N +KA+LMN+
Sbjct: 240 SVRDLDAFYEQMIDLHLQKNREESEDDFVDLLLRLEKEEAVLGYGKLTRNHIKAILMNIL 299
Query: 274 LGGSDTTSTTLEWAMA 289
LGG +T++ T+ WAMA
Sbjct: 300 LGGINTSAITMTWAMA 315
>C7D8_SOYBN (O81974) Cytochrome P450 71D8 (EC 1.14.-.-) (P450 CP7)
Length = 504
Score = 59.7 bits (143), Expect = 9e-09
Identities = 28/96 (29%), Positives = 57/96 (59%), Gaps = 8/96 (8%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRR--------DPKKKDFLDILLQLQDDGLTEFELTQN 263
K++ + D L+ ++ +H R + +++D +D+LL+L++ G E +T
Sbjct: 235 KVEHVHQRADKILEDILRKHMEKRTRVKEGNGSEAEQEDLVDVLLRLKESGSLEVPMTME 294
Query: 264 DLKALLMNMFLGGSDTTSTTLEWAMAACKKSSHNEE 299
++KA++ N+F G+DT+++TLEWAM+ K+ +E
Sbjct: 295 NIKAVIWNIFAAGTDTSASTLEWAMSEMMKNPKVKE 330
>C751_PETHY (P48418) Flavonoid 3',5'-hydroxylase 1 (EC 1.14.-.-)
(F3'5'H) (Cytochrome P450 75A1) (CYPLXXVA1)
Length = 506
Score = 59.7 bits (143), Expect = 9e-09
Identities = 31/85 (36%), Positives = 55/85 (64%), Gaps = 2/85 (2%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPK-KKDFLDILLQLQDDGLTEFELTQNDLKALL 269
K++K ++ D L ++ EHK + + K K DFLD++++ D+ E L+ ++KALL
Sbjct: 237 KRMKRLHKKFDALLTKMFDEHKATTYERKGKPDFLDVVMENGDNSEGE-RLSTTNIKALL 295
Query: 270 MNMFLGGSDTTSTTLEWAMAACKKS 294
+N+F G+DT+S+ +EWA+A K+
Sbjct: 296 LNLFTAGTDTSSSAIEWALAEMMKN 320
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.353 0.156 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,050,751
Number of Sequences: 164201
Number of extensions: 1057904
Number of successful extensions: 6781
Number of sequences better than 10.0: 329
Number of HSP's better than 10.0 without gapping: 138
Number of HSP's successfully gapped in prelim test: 191
Number of HSP's that attempted gapping in prelim test: 6465
Number of HSP's gapped (non-prelim): 363
length of query: 313
length of database: 59,974,054
effective HSP length: 110
effective length of query: 203
effective length of database: 41,911,944
effective search space: 8508124632
effective search space used: 8508124632
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 66 (30.0 bits)
Medicago: description of AC124952.12


