

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124951.10 - phase: 0
(446 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
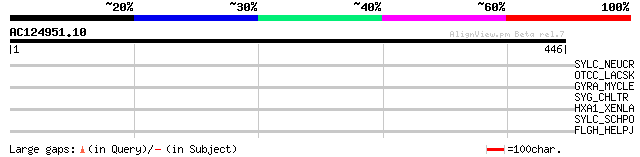
Score E
Sequences producing significant alignments: (bits) Value
SYLC_NEUCR (P10857) Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.... 33 1.1
OTCC_LACSK (O53089) Ornithine carbamoyltransferase, catabolic (E... 32 3.3
GYRA_MYCLE (Q57532) DNA gyrase subunit A (EC 5.99.1.3) [Contains... 32 3.3
SYG_CHLTR (Q46371) Glycyl-tRNA synthetase (EC 6.1.1.14) (GlyRS) ... 32 4.3
HXA1_XENLA (Q08821) Homeobox protein Hox-A1 (Hox.lab2) (Fragment) 31 7.4
SYLC_SCHPO (Q10490) Putative leucyl-tRNA synthetase, cytoplasmic... 30 9.6
FLGH_HELPJ (Q9ZMB3) Flagellar L-ring protein precursor (Basal bo... 30 9.6
>SYLC_NEUCR (P10857) Leucyl-tRNA synthetase, cytoplasmic (EC
6.1.1.4) (Leucine--tRNA ligase) (LeuRS)
Length = 1123
Score = 33.5 bits (75), Expect = 1.1
Identities = 15/39 (38%), Positives = 22/39 (55%), Gaps = 2/39 (5%)
Query: 237 LMNWSCAMAYGVGSQSPLSPSFNFSLELVKSSQFVASFY 275
L W+CA YG+GS+ P P NF +E + S ++Y
Sbjct: 608 LKQWACARTYGLGSKLPWDP--NFLVESLSDSTVYMAYY 644
>OTCC_LACSK (O53089) Ornithine carbamoyltransferase, catabolic (EC
2.1.3.3) (OTCase)
Length = 337
Score = 32.0 bits (71), Expect = 3.3
Identities = 15/34 (44%), Positives = 20/34 (58%)
Query: 397 PNFVMLTPSKEHLAEGIVWETGKRPMLQSDIDAG 430
P+ + T LAEG ETG + M+ SD+DAG
Sbjct: 188 PDSLQPTQEVRDLAEGYAKETGSKNMITSDVDAG 221
>GYRA_MYCLE (Q57532) DNA gyrase subunit A (EC 5.99.1.3) [Contains: Mle
gyrA intein]
Length = 1273
Score = 32.0 bits (71), Expect = 3.3
Identities = 27/91 (29%), Positives = 38/91 (41%), Gaps = 5/91 (5%)
Query: 120 LYVSTSDPVLQMRSCAYYPRYGFGAFGVFPLLLKNRTSSQIATFTADKVRQLYRETSQDS 179
L V+TS + S YP G G GV ++ R S + D+ +LY TS
Sbjct: 1144 LLVATSGGYAKRTSIEEYPMQGRGGKGVLTVMYDRRRGSLVGAIVVDEDSELYAITS-GG 1202
Query: 180 GVMGLR----YGSGNLSCGVTLMPFAKKDEL 206
GV+ +G + GV LM + D L
Sbjct: 1203 GVIRTTARQVRQAGRQTKGVRLMNLGEGDTL 1233
>SYG_CHLTR (Q46371) Glycyl-tRNA synthetase (EC 6.1.1.14) (GlyRS)
[Includes: Glycyl-tRNA synthetase alpha chain
(Glycine--tRNA ligase alpha chain); Glycyl-tRNA
synthetase beta chain (Glycine--tRNA ligase beta chain)]
Length = 1003
Score = 31.6 bits (70), Expect = 4.3
Identities = 38/169 (22%), Positives = 61/169 (35%), Gaps = 34/169 (20%)
Query: 39 MLESSYDVVFGKLALRCLFNDYFQQPKHFVTRIMLKPIDDPHVDLIATVSGPLDQKPDEN 98
++ + +D F L L + Q ++F T+ M I + LI + P D + N
Sbjct: 577 VISAQFDPAFCSLPKELLIAEMIQHQRYFPTQNMQGEITNRF--LIVCDNSPTDSIVEGN 634
Query: 99 --------INGNALFRWQSDVNDPHTFMDLYVSTSDPVLQMRSCAYYPRYGF-------- 142
+GN LF+ DL S V +++S Y+ G
Sbjct: 635 EKALAPRLTDGNFLFK-----------QDLLTPLSSFVEKLKSVTYFESLGSLADKTSRL 683
Query: 143 -----GAFGVFPLLLKNRTSSQIATFTADKVRQLYRETSQDSGVMGLRY 186
A+ + PL K + I AD V + E + G+MG Y
Sbjct: 684 KLHLEEAYALLPLCAKEDIDTAIHYCKADLVSSVVNEFPELQGIMGRYY 732
>HXA1_XENLA (Q08821) Homeobox protein Hox-A1 (Hox.lab2) (Fragment)
Length = 240
Score = 30.8 bits (68), Expect = 7.4
Identities = 20/80 (25%), Positives = 35/80 (43%)
Query: 241 SCAMAYGVGSQSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENTVVGITNYI 300
SC YG+ + SP F E SS F S Y + V++ ++ + G +YI
Sbjct: 6 SCGSNYGMQNFSPGYSHFPIHQETEVSSGFPQSVYSGNIASSVVQHQQHQSYIEGSAHYI 65
Query: 301 DFGFELQTSVDDAIAANNIS 320
+ + ++ A NN++
Sbjct: 66 HHSYGPEQNLSVANYNNNVA 85
>SYLC_SCHPO (Q10490) Putative leucyl-tRNA synthetase, cytoplasmic
(EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS)
Length = 1111
Score = 30.4 bits (67), Expect = 9.6
Identities = 13/39 (33%), Positives = 22/39 (56%), Gaps = 2/39 (5%)
Query: 237 LMNWSCAMAYGVGSQSPLSPSFNFSLELVKSSQFVASFY 275
L W+CA +YG+G++ P P F +E + S ++Y
Sbjct: 592 LSQWACARSYGLGTRLPWDP--QFLVESLTDSTIYMAYY 628
>FLGH_HELPJ (Q9ZMB3) Flagellar L-ring protein precursor (Basal body
L-ring protein)
Length = 237
Score = 30.4 bits (67), Expect = 9.6
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 4/53 (7%)
Query: 374 GQVQYGFGLQSESLREASYQRADPNFVMLTPSKEHLAE----GIVWETGKRPM 422
G V +G S E + PN+V TPSKE + E G ++ G+RP+
Sbjct: 8 GAVAFGVVFSMASANEPNIDFNPPNYVEETPSKEFIPELNKLGSLFGQGERPL 60
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.135 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,391,280
Number of Sequences: 164201
Number of extensions: 2163797
Number of successful extensions: 4784
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 4779
Number of HSP's gapped (non-prelim): 7
length of query: 446
length of database: 59,974,054
effective HSP length: 113
effective length of query: 333
effective length of database: 41,419,341
effective search space: 13792640553
effective search space used: 13792640553
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC124951.10


