

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
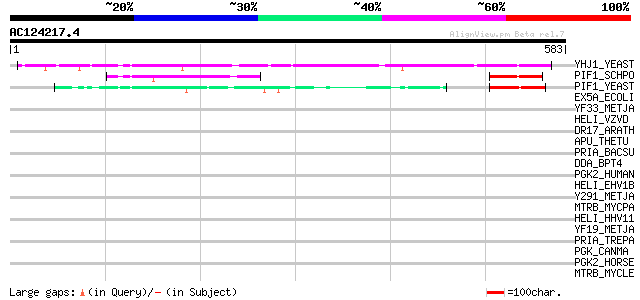
Query= AC124217.4 + phase: 0
(583 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 78 6e-14
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 61 9e-09
PIF1_YEAST (P07271) DNA repair and recombination protein PIF1, m... 54 1e-06
EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 37 0.11
YF33_METJA (Q58928) Hypothetical protein MJ1533 36 0.32
HELI_VZVD (P09303) Probable helicase 35 0.41
DR17_ARATH (O64790) Putative disease resistance protein At1g61300 35 0.71
APU_THETU (P38536) Amylopullulanase precursor (Alpha-amylase/pul... 35 0.71
PRIA_BACSU (P94461) Primosomal protein N' (Replication factor Y) 34 0.92
DDA_BPT4 (P32270) DNA helicase 34 0.92
PGK2_HUMAN (P07205) Phosphoglycerate kinase, testis specific (EC... 34 1.2
HELI_EHV1B (P28934) Probable helicase 34 1.2
Y291_METJA (Q57739) Probable signal recognition particle protein... 32 3.5
MTRB_MYCPA (Q93CB7) Sensor histidine kinase mtrB (EC 2.7.3.-) 32 3.5
HELI_HHV11 (P10189) Probable helicase 32 3.5
YF19_METJA (Q58914) Hypothetical protein MJ1519 32 4.6
PRIA_TREPA (O83258) Primosomal protein N' (Replication factor Y) 32 4.6
PGK_CANMA (P41757) Phosphoglycerate kinase (EC 2.7.2.3) 32 4.6
PGK2_HORSE (Q8MIF7) Phosphoglycerate kinase, testis specific (EC... 32 4.6
MTRB_MYCLE (Q9CCJ1) Sensor histidine kinase mtrB (EC 2.7.3.-) 32 4.6
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 78.2 bits (191), Expect = 6e-14
Identities = 141/612 (23%), Positives = 247/612 (40%), Gaps = 81/612 (13%)
Query: 9 SFLHQCYTLPLTYVNKCYVFDADLTLTD--------EQLQIYALADVEKLLRSFGKSLSD 60
S LH+ T + K Y FD D TL + QL + ++E S S
Sbjct: 128 SSLHK--TASASTTQKTYHFDEDETLREVTSVKSNSRQLSFTSTINIED--SSMKLSTDS 183
Query: 61 YPPMPTADPSL--------MPDLQNHFIHEEINYNRRELQEEHNQLMSKMTDEQHKVYDT 112
P + PS+ +P + + ++ N + +T EQ +V +
Sbjct: 184 ERPAKRSKPSMEFQGLKLTVPKKIKPLLRKTVS-NMDSMNHRSASSPVVLTMEQERVVNL 242
Query: 113 IMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSR--GEIVLTVASSGIAALLIPGGR 170
I+ +K+ +F+ G GTGK+ I + + + S E + AS+G+AA+ I GG
Sbjct: 243 IV----KKRTNVFYT-GSAGTGKSVILQTIIRQLSSLYGKESIAITASTGLAAVTI-GGS 296
Query: 171 TAHSRFGIPF---TVDECSTCGVKPNTPLAQLVVQARLIIWDEAPMMHKFCFEALDRSCR 227
T H GI T+D+ ++ L +++I DE M+ + L++ R
Sbjct: 297 TLHKWSGIGIGNKTIDQLVK-KIQSQKDLLAAWRYTKVLIIDEISMVDGNLLDKLEQIAR 355
Query: 228 DVMKEVDKRNKNIPFGGKVVVFGGDFRQILPVIPRGIRPEIVHATINSSHLWRHC--EVL 285
+ K D PFGG +V GDF Q+ PV + + S +W+ C + +
Sbjct: 356 RIRKNDD------PFGGIQLVLTGDFFQLPPVAKKDEHNVVKFCF--ESEMWKRCIQKTI 407
Query: 286 RLTKNMRLLNGASEADIEDRRKFSEWVLSIGDGTIGEDNLVDKTITI-PTDLLVEVDGCP 344
LTK R + DI + ++ E + I + +D I PT+L
Sbjct: 408 LLTKVFRQQDN-KLIDILNAIRYGELTVDIAKTIRNLNRDIDYADGIAPTELYATRREVE 466
Query: 345 IASIIR-STYPNLLASMGDIEYFQNR--AILTSKNTIVEKINEYMLDMVPGEEKVYLSYD 401
++++ + + P L ++ R AIL S + +VEK+ D +V + +
Sbjct: 467 LSNVKKLQSLPGDLYEFKAVDNAPERYQAILDS-SLMVEKVVALKED-----AQVMMLKN 520
Query: 402 SPDERNVCGD---------------------AMDD--VHTPKFLNTIVASGLPNHKLRLK 438
PD V G +DD V + ++ ++ + L +
Sbjct: 521 KPDVELVNGSLGKVLFFVTESLVVKMKEIYKIVDDEVVMDMRLVSRVIGNPLLKESKEFR 580
Query: 439 EGVPVMLLRNLDTKNGLCNGTRLIITRMGRYVLEGKVISGSNVGDRVFIPRLSLSPSDV- 497
+ + L L+ L N I ++ + + + +P P D+
Sbjct: 581 QDLNARPLARLERLKILINYAVKISPHKEKFPYVRWTVGKNKYIHELMVP--ERFPIDIP 638
Query: 498 RIPFKFQRKQFPLAVSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLK 557
R +R Q PL + +A++I+K+QGQ++Q + V L +F GQ+YVA+SR + L+
Sbjct: 639 RENVGLERTQIPLMLCWALSIHKAQGQTIQRLKVDL-RRIFEAGQVYVALSRAVTMDTLQ 697
Query: 558 ILITDEDGDDTN 569
+L D TN
Sbjct: 698 VLNFDPGKIRTN 709
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 60.8 bits (146), Expect = 9e-09
Identities = 49/168 (29%), Positives = 82/168 (48%), Gaps = 18/168 (10%)
Query: 102 MTDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSR----GEIVLTVA 157
++DEQ ++ D ++ E++ IFF G GTGK+ + + + ++S+ + V A
Sbjct: 310 LSDEQKRILDMVV----EQQHSIFFT-GSAGTGKSVLLRKIIEVLKSKYRKQSDRVAVTA 364
Query: 158 SSGIAALLIPGGRTAHSRFGIPFTVDECS--TCGVKPNTPLAQLVVQARLIIWDEAPMMH 215
S+G+AA I GG T HS G+ + +K N ++ R++I DE M+
Sbjct: 365 STGLAACNI-GGVTLHSFAGVGLARESVDLLVSKIKKNKKCVNRWLRTRVLIIDEVSMVD 423
Query: 216 KFCFEALDRSCRDVMKEVDKRNKNIPFGGKVVVFGGDFRQILPVIPRG 263
+ L+ R + K+ + PFGG +V GDF Q+ PV G
Sbjct: 424 AELMDKLEEVARVIRKD------SKPFGGIQLVLTGDFFQLPPVPENG 465
Score = 52.0 bits (123), Expect = 4e-06
Identities = 27/55 (49%), Positives = 40/55 (72%), Gaps = 1/55 (1%)
Query: 505 RKQFPLAVSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLKIL 559
R Q PL +++A++I+K+QGQ+L V V L VF GQ YVA+SR T++ GL++L
Sbjct: 708 RSQIPLILAYAISIHKAQGQTLDRVKVDL-GRVFEKGQAYVALSRATTQEGLQVL 761
>PIF1_YEAST (P07271) DNA repair and recombination protein PIF1,
mitochondrial precursor
Length = 857
Score = 53.5 bits (127), Expect = 1e-06
Identities = 105/431 (24%), Positives = 164/431 (37%), Gaps = 106/431 (24%)
Query: 48 EKLLRSFGKSLSDYPPMPTADPSLMPDLQNHFIHEEINYNRRELQEEHNQLMSKMTDEQH 107
EK FG+ ++ P+ +LQN ++ N N HN + K+
Sbjct: 192 EKKKMQFGEKIAVLTQRPS-----FTELQND--QDDSNLN------PHNGVKVKIPICLS 238
Query: 108 KVYDTIMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRS--RGEIVLTVASSGIAALL 165
K ++I+ ++ E IF+ G GTGK+ + + M ++ E V AS+G+AA
Sbjct: 239 KEQESII-KLAENGHNIFYT-GSAGTGKSILLREMIKVLKGIYGRENVAVTASTGLAACN 296
Query: 166 IPGGRTAHSRFGIPFTVDE---CSTCGVKPNTPLAQLVVQARLIIWDEAPMMHKFCFEAL 222
I GG T HS GI D G + L + L++ DE M+ + L
Sbjct: 297 I-GGITIHSFAGILGKGDADKLYKKVGRRSRKHLRRWENIGALVV-DEISMLDAELLDKL 354
Query: 223 DRSCRDVMKEVDKRNKNIPFGGKVVVFGGDFRQILPVIPRGIRP-----------EIVHA 271
D R + R + PFGG ++F GDF Q+ PV RP E V
Sbjct: 355 DFIARKI------RKNHQPFGGIQLIFCGDFFQLPPVSKDPNRPTKFAFESKAWKEGVKM 408
Query: 272 TINSSHLWRH---CEVLRLTKNMRLLNGASEADIEDRRKFSEWVLSIGDGTIGEDNLVDK 328
TI ++R + + + MRL N D E R+F + + D I
Sbjct: 409 TIMLQKVFRQRGDVKFIEMLNRMRLGN----IDDETEREFKKLSRPLPDDEI-------- 456
Query: 329 TITIPTDLLVEVDGCPIASIIRSTYPNLLASMGDIEYFQNRAILTSKNTIVEKINEYMLD 388
IP +L S VE+ N L
Sbjct: 457 ---IPAELY------------------------------------STRMEVERANNSRLS 477
Query: 389 MVPGEEKVYLSYDSPDERNVCGDAMDDVHTPKFLNTIVASGLPNHKLRLKEGVPVMLLRN 448
+PG+ ++ + D G A++D + ++ + L +L LK G VM+++N
Sbjct: 478 KLPGQVHIFNAID--------GGALED---EELKERLLQNFLAPKELHLKVGAQVMMVKN 526
Query: 449 LDTKNGLCNGT 459
LD L NG+
Sbjct: 527 LDAT--LVNGS 535
Score = 51.2 bits (121), Expect = 7e-06
Identities = 29/58 (50%), Positives = 40/58 (68%), Gaps = 1/58 (1%)
Query: 505 RKQFPLAVSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLKILITD 562
R Q PL ++++++I+KSQGQ+L V V L VF GQ YVA+SR SR GL++L D
Sbjct: 691 RVQLPLMLAWSLSIHKSQGQTLPKVKVDLRR-VFEKGQAYVALSRAVSREGLQVLNFD 747
>EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5) (Exodeoxyribonuclease V 67 kDa polypeptide)
Length = 608
Score = 37.4 bits (85), Expect = 0.11
Identities = 37/121 (30%), Positives = 54/121 (44%), Gaps = 22/121 (18%)
Query: 436 RLKEGVPVMLLRNLDTKNGLCNGTRLIITRMGRYVLEGKVISGSNVGDRVFIPRLSLSPS 495
R EG PVM+ RN D+ GL NG I G+ + N+ S+ PS
Sbjct: 475 RWYEGRPVMIARN-DSALGLFNGDIGIALDRGQGTRVWFAMPDGNIK--------SVQPS 525
Query: 496 DVRIPFKFQRKQFPLAVSFAMTINKSQGQSLQNVGVYLPA---PVFSHGQLYVAVSRVTS 552
R+P ++AMT++KSQG + + LP+ PV + +Y AV+R
Sbjct: 526 --RLPEH--------ETTWAMTVHKSQGSEFDHAALILPSQRTPVVTRELVYTAVTRARR 575
Query: 553 R 553
R
Sbjct: 576 R 576
>YF33_METJA (Q58928) Hypothetical protein MJ1533
Length = 642
Score = 35.8 bits (81), Expect = 0.32
Identities = 21/58 (36%), Positives = 36/58 (61%), Gaps = 1/58 (1%)
Query: 101 KMTDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGEIVLTVAS 158
K + E +++ D +M R+ E+ GIF + G G+GK+ A++ RS+G+IV T+ S
Sbjct: 264 KASLEDYELSDKLMERLKERAEGIF-VSGPPGSGKSTFVAALAEFYRSQGKIVKTMES 320
>HELI_VZVD (P09303) Probable helicase
Length = 881
Score = 35.4 bits (80), Expect = 0.41
Identities = 20/51 (39%), Positives = 28/51 (54%)
Query: 508 FPLAVSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLKI 558
+ L+ AMTI +SQG SL+ V + A +YVA+SR S LK+
Sbjct: 799 YGLSSKLAMTIARSQGLSLEKVAICFTADKLRLNSVYVAMSRTVSSRFLKM 849
>DR17_ARATH (O64790) Putative disease resistance protein At1g61300
Length = 766
Score = 34.7 bits (78), Expect = 0.71
Identities = 28/90 (31%), Positives = 45/90 (49%), Gaps = 7/90 (7%)
Query: 78 HFIHEEINYNRRELQEEHNQLMSKMTDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKTF 137
+F+ IN N ++E Q T Q ++ + R+ E + GI L+G GG GKT
Sbjct: 21 NFLCGNINRNSFGVEERPTQ----PTIGQEEMLEKAWNRLMEDRVGIMGLHGMGGVGKTT 76
Query: 138 IWKAMS---AAIRSRGEIVLTVASSGIAAL 164
++K + A + SR +IV+ + S A L
Sbjct: 77 LFKKIHNKFAKMSSRFDIVIWIVVSKGAKL 106
>APU_THETU (P38536) Amylopullulanase precursor
(Alpha-amylase/pullulanase) (Pullulanase type II)
[Includes: Alpha-amylase (EC 3.2.1.1)
(1,4-alpha-D-glucan glucanohydrolase); Pullulanase (EC
3.2.1.41) (1,4-alpha-D-glucan glucanohydrolase)
(Alpha-dextrin en
Length = 1861
Score = 34.7 bits (78), Expect = 0.71
Identities = 42/166 (25%), Positives = 65/166 (38%), Gaps = 21/166 (12%)
Query: 377 TIVEKINEYMLDMVPGEEKVYLSYDSPDERNVCGDAMDDVHTPKFLNTIVASGLPNHKLR 436
TI++ M D+ G+E PD+R +D F I S + N+
Sbjct: 768 TILQMGYPGMADIYYGDEAGVSGGKDPDDRRTFPWGNEDTTLQDFFKNI--SSIRNNNQV 825
Query: 437 LKEGVPVMLLRNLDTKNGLCNGTRLIITRMGRYVLEGKVISGSNVGDRVFIPRLSLSPSD 496
LK G L L +N + +GR ++ GK G++ D I ++ S SD
Sbjct: 826 LKTGD----LETLYAQND--------VYAIGRRIINGKDAFGTSYPDSAAIVAINRSKSD 873
Query: 497 VRIPF---KFQRKQFPLAVSFAMTINKSQGQSLQNVGVYLPAPVFS 539
+I KF R V+F IN + S+ N + + P S
Sbjct: 874 KQIAIDTTKFLRD----GVTFKDLINNNVSYSISNGQIVIDVPAMS 915
>PRIA_BACSU (P94461) Primosomal protein N' (Replication factor Y)
Length = 805
Score = 34.3 bits (77), Expect = 0.92
Identities = 15/50 (30%), Positives = 28/50 (56%)
Query: 102 MTDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGE 151
+TDEQ ++ I +D + +F L+G G+GKT I+ + ++G+
Sbjct: 268 LTDEQRAAFEPIRETLDSDEHKVFLLHGVTGSGKTEIYLQSIEKVLAKGK 317
>DDA_BPT4 (P32270) DNA helicase
Length = 439
Score = 34.3 bits (77), Expect = 0.92
Identities = 37/128 (28%), Positives = 57/128 (43%), Gaps = 12/128 (9%)
Query: 102 MTDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGEIVLTVASSGI 161
+T+ Q ++ +M + EKK + + G GTGKT + K + A+ S G + +A+
Sbjct: 6 LTEGQKNAFNIVMKAIKEKKHHVT-INGPAGTGKTTLTKFIIEALISTGGTGIILAAPTH 64
Query: 162 AALLI------PGGRTAHSRFGI-PFTVDECSTCGVKPNTPLAQLVVQARLIIWDEAPMM 214
AA I T HS I P T +E K LA + R++I DE M
Sbjct: 65 AAKKILSKLSGKEASTIHSILKINPVTYEENVLFEQKEVPDLA----KCRVLICDEVSMY 120
Query: 215 HKFCFEAL 222
+ F+ L
Sbjct: 121 DRKLFKIL 128
>PGK2_HUMAN (P07205) Phosphoglycerate kinase, testis specific (EC
2.7.2.3)
Length = 416
Score = 33.9 bits (76), Expect = 1.2
Identities = 22/65 (33%), Positives = 34/65 (51%), Gaps = 5/65 (7%)
Query: 188 CGVKPNTPLAQLVVQARLIIWDEAPMMHKFCFEALDRSCRDVMKEVDKRNKNIPFGGKVV 247
CG + N AQ+V QARLI+W+ + F ++A + + +M E+ K G V
Sbjct: 315 CGPESNKNHAQVVAQARLIVWNGP--LGVFEWDAFAKGTKALMDEIVKATSK---GCITV 369
Query: 248 VFGGD 252
+ GGD
Sbjct: 370 IGGGD 374
>HELI_EHV1B (P28934) Probable helicase
Length = 881
Score = 33.9 bits (76), Expect = 1.2
Identities = 19/51 (37%), Positives = 27/51 (52%)
Query: 508 FPLAVSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTSRGGLKI 558
+ L+ AMTI +SQG SL V + P +YVA+SR S L++
Sbjct: 799 YGLSSKLAMTIARSQGLSLDKVAICFPRNNLRINSVYVAMSRTVSSRFLRM 849
>Y291_METJA (Q57739) Probable signal recognition particle protein
MJ0291
Length = 409
Score = 32.3 bits (72), Expect = 3.5
Identities = 24/122 (19%), Positives = 49/122 (39%), Gaps = 4/122 (3%)
Query: 46 DVEKLLRSFGKSLSDYPPMPTADPSLMPDLQNHFIHEEI----NYNRRELQEEHNQLMSK 101
D+E +L +L + L+ +++N + +I N + N + +
Sbjct: 122 DIEDVLEELEIALLEADVALEVVEKLIENIKNELVGRKISPDDNVEEITINAVKNAIKNI 181
Query: 102 MTDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGEIVLTVASSGI 161
++ E+ + + I E KP + G GTGKT ++ ++ +G V+ A
Sbjct: 182 LSQEKIDIEEIIKKNKAEGKPTVIVFVGINGTGKTTTIAKLAYKLKQKGYSVVLAAGDTF 241
Query: 162 AA 163
A
Sbjct: 242 RA 243
>MTRB_MYCPA (Q93CB7) Sensor histidine kinase mtrB (EC 2.7.3.-)
Length = 565
Score = 32.3 bits (72), Expect = 3.5
Identities = 28/111 (25%), Positives = 57/111 (51%), Gaps = 14/111 (12%)
Query: 296 GASEADIE--DRRKFSEWVLSIGDGTIGEDNLVDKTITIPTD-LLVEVDGCPIASIIRST 352
G +E +E D R + LS G + ED ++ + +P + ++ EVD + I+R
Sbjct: 362 GVAELSVEAVDLRSTVQSALS-NVGHLAEDAGIELQVELPAEEVIAEVDTRRVERILR-- 418
Query: 353 YPNLLASMGDIEYFQNRAI----LTSKNTIVEKINEYMLDMVPGEEKVYLS 399
NL+A+ I++ +++ + ++T+ + +Y + + PGEEK+ S
Sbjct: 419 --NLIANA--IDHAEHKPVKIRMAADEDTVAVTVRDYGVGLRPGEEKLVFS 465
>HELI_HHV11 (P10189) Probable helicase
Length = 882
Score = 32.3 bits (72), Expect = 3.5
Identities = 23/64 (35%), Positives = 29/64 (44%), Gaps = 4/64 (6%)
Query: 508 FPLAVSFAMTINKSQGQSLQNVGVYLPAPVFSHGQLYVAVSRVTS----RGGLKILITDE 563
+ ++ AMTI +SQG SL V + YVA+SR TS R L L
Sbjct: 800 YGISSKLAMTITRSQGLSLDKVAICFTPGNLRLNSAYVAMSRTTSSEFLRMNLNPLRERH 859
Query: 564 DGDD 567
D DD
Sbjct: 860 DRDD 863
>YF19_METJA (Q58914) Hypothetical protein MJ1519
Length = 1175
Score = 32.0 bits (71), Expect = 4.6
Identities = 18/54 (33%), Positives = 32/54 (58%), Gaps = 5/54 (9%)
Query: 513 SFAMTINKSQGQSLQNVGVYLPAPV---FSHGQLYVAVSRVTSRGGLKILITDE 563
++A+TI+KSQG +NV + +P + S LY A++R R L +++ +E
Sbjct: 960 AYAITIHKSQGSGFENVILIIPKGLNKFVSKEMLYTAITRAKKR--LYVIVEEE 1011
>PRIA_TREPA (O83258) Primosomal protein N' (Replication factor Y)
Length = 657
Score = 32.0 bits (71), Expect = 4.6
Identities = 16/53 (30%), Positives = 30/53 (56%), Gaps = 3/53 (5%)
Query: 102 MTDEQHKVYDTIMARVDEKKPGIFFLYGYGGTGKTFIWKAMSAAIRSRGEIVL 154
++ EQ K D I A + F+++G G+GKT ++ + A+ +RG+ V+
Sbjct: 132 LSGEQRKAIDAITASTGARS---FYVHGVTGSGKTEVFLRAAEAVLARGKSVI 181
>PGK_CANMA (P41757) Phosphoglycerate kinase (EC 2.7.2.3)
Length = 417
Score = 32.0 bits (71), Expect = 4.6
Identities = 18/65 (27%), Positives = 29/65 (43%), Gaps = 5/65 (7%)
Query: 188 CGVKPNTPLAQLVVQARLIIWDEAPMMHKFCFEALDRSCRDVMKEVDKRNKNIPFGGKVV 247
CG K A V +A+ I+W+ P + +F D+ +D K+ G V+
Sbjct: 315 CGPKSRKLFADAVAKAKTIVWNGPPGVFEF-----DKFAEGTKSLLDDAVKSAEVGNIVI 369
Query: 248 VFGGD 252
+ GGD
Sbjct: 370 IGGGD 374
>PGK2_HORSE (Q8MIF7) Phosphoglycerate kinase, testis specific (EC
2.7.2.3)
Length = 416
Score = 32.0 bits (71), Expect = 4.6
Identities = 19/65 (29%), Positives = 34/65 (52%), Gaps = 5/65 (7%)
Query: 188 CGVKPNTPLAQLVVQARLIIWDEAPMMHKFCFEALDRSCRDVMKEVDKRNKNIPFGGKVV 247
CG + N AQ++ QA+LI+W+ + F ++A + + +M E+ K G +
Sbjct: 315 CGPETNKKYAQVMAQAKLIVWNGP--VGVFEWDAFAKGTKALMDEIVKATSR---GCITI 369
Query: 248 VFGGD 252
+ GGD
Sbjct: 370 IGGGD 374
>MTRB_MYCLE (Q9CCJ1) Sensor histidine kinase mtrB (EC 2.7.3.-)
Length = 562
Score = 32.0 bits (71), Expect = 4.6
Identities = 21/87 (24%), Positives = 47/87 (53%), Gaps = 11/87 (12%)
Query: 318 GTIGEDNLVDKTITIPTD-LLVEVDGCPIASIIRSTYPNLLASMGDIEYFQNRAI----L 372
G + E+ ++ + +P D ++ EVD + I+R NL+A+ I++ +++ +
Sbjct: 385 GHLAEEAGIELLVDMPVDEVIAEVDARRVERILR----NLIANA--IDHSEHKPVRIRMA 438
Query: 373 TSKNTIVEKINEYMLDMVPGEEKVYLS 399
++T+ + +Y + + PGEEK+ S
Sbjct: 439 ADEDTVAVTVRDYGIGLRPGEEKLVFS 465
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,128,281
Number of Sequences: 164201
Number of extensions: 3094042
Number of successful extensions: 8512
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 8492
Number of HSP's gapped (non-prelim): 33
length of query: 583
length of database: 59,974,054
effective HSP length: 116
effective length of query: 467
effective length of database: 40,926,738
effective search space: 19112786646
effective search space used: 19112786646
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC124217.4


