

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
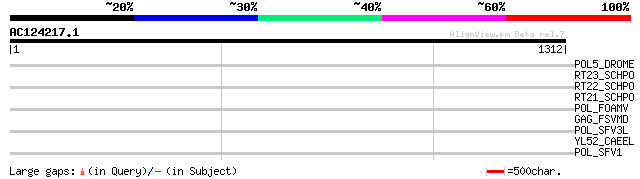
Query= AC124217.1 - phase: 0 /pseudo
(1312 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 40 0.032
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 36 0.79
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 36 0.79
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 36 0.79
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 34 3.0
GAG_FSVMD (P03340) Gag polyprotein [Contains: Core protein p15; ... 33 3.9
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 33 6.7
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 32 8.7
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 32 8.7
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 40.4 bits (93), Expect = 0.032
Identities = 19/52 (36%), Positives = 32/52 (61%)
Query: 1261 QVILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
QV LG+++ GI+ + K +AIS P ++ KEL+ FLG ++ +FI +
Sbjct: 323 QVEFLGYIVTADGIKADPKKVRAISEMPPPTSVKELKRFLGMTSYYRKFIQD 374
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 35.8 bits (81), Expect = 0.79
Identities = 18/50 (36%), Positives = 28/50 (56%)
Query: 1261 QVILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFI 1310
QV +G+ I++KG Q + K +KEL+ FLG +N+L +FI
Sbjct: 607 QVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFI 656
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 35.8 bits (81), Expect = 0.79
Identities = 18/50 (36%), Positives = 28/50 (56%)
Query: 1261 QVILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFI 1310
QV +G+ I++KG Q + K +KEL+ FLG +N+L +FI
Sbjct: 607 QVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFI 656
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 35.8 bits (81), Expect = 0.79
Identities = 18/50 (36%), Positives = 28/50 (56%)
Query: 1261 QVILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFI 1310
QV +G+ I++KG Q + K +KEL+ FLG +N+L +FI
Sbjct: 607 QVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFI 656
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 33.9 bits (76), Expect = 3.0
Identities = 22/52 (42%), Positives = 29/52 (55%), Gaps = 2/52 (3%)
Query: 1262 VILLGFVINKKGIEINQN-KTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
V LGF I K+G + KTK ++ T P K +LQ+ LG +NF FI N
Sbjct: 159 VEFLGFNITKEGRGLTDTFKTKLLNITPPKDLK-QLQSILGLLNFARNFIPN 209
>GAG_FSVMD (P03340) Gag polyprotein [Contains: Core protein p15;
Core protein p12; Core protein p30; Core protein p10]
Length = 536
Score = 33.5 bits (75), Expect = 3.9
Identities = 20/71 (28%), Positives = 37/71 (51%), Gaps = 2/71 (2%)
Query: 467 RKRKW-TDNKPKFIFNKRGVPYKDTPPTRHQNRGQVRTFTPSSKSPTEKWVFSSGKKSSY 525
RK+KW T + +++ G P + TPP + ++ + R F P ++ + + +S
Sbjct: 108 RKKKWITLCEAEWVMMNVGWPREGTPPLDNTSQVEKRIFAPGPHGHPDQVPYITTWRSLA 167
Query: 526 TSPPAKWVKRF 536
T PP+ WV+ F
Sbjct: 168 TDPPS-WVRPF 177
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 32.7 bits (73), Expect = 6.7
Identities = 18/52 (34%), Positives = 27/52 (51%)
Query: 1261 QVILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
+V LGF I K+G + + + + P K+LQ+ LG +NF FI N
Sbjct: 369 EVEFLGFNITKEGRGLTETFKQKLLNITPPRDLKQLQSILGLLNFARNFIPN 420
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 32.3 bits (72), Expect = 8.7
Identities = 17/52 (32%), Positives = 27/52 (51%)
Query: 1261 QVILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
+V LG + G+E + KT + + KELQ+FLG + + +FI N
Sbjct: 1137 EVEYLGHKVTLDGVETQEVKTDKMKQFSRPTNVKELQSFLGLVGYYRKFILN 1188
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 32.3 bits (72), Expect = 8.7
Identities = 18/52 (34%), Positives = 26/52 (49%)
Query: 1261 QVILLGFVINKKGIEINQNKTKAIS*TKPSSTKKELQTFLGKINFLIRFISN 1312
+V LGF I K+G + + + P K+LQ+ LG +NF FI N
Sbjct: 367 EVEFLGFNITKEGRGLTDTFKQKLLNITPPKDLKQLQSILGLLNFARNFIPN 418
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.358 0.160 0.568
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 130,283,450
Number of Sequences: 164201
Number of extensions: 5011532
Number of successful extensions: 24581
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 24572
Number of HSP's gapped (non-prelim): 11
length of query: 1312
length of database: 59,974,054
effective HSP length: 122
effective length of query: 1190
effective length of database: 39,941,532
effective search space: 47530423080
effective search space used: 47530423080
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 72 (32.3 bits)
Medicago: description of AC124217.1


