

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.10 - phase: 0
(543 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
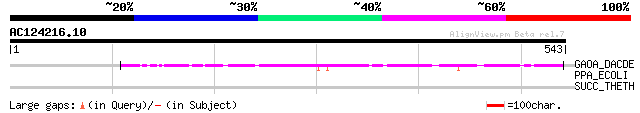
Score E
Sequences producing significant alignments: (bits) Value
GAOA_DACDE (Q01745) Galactose oxidase precursor (EC 1.1.3.9) (GAO) 90 1e-17
PPA_ECOLI (P07102) Periplasmic appA protein precursor [Includes:... 32 3.2
SUCC_THETH (P25126) Succinyl-CoA synthetase beta chain (EC 6.2.1... 31 9.4
>GAOA_DACDE (Q01745) Galactose oxidase precursor (EC 1.1.3.9) (GAO)
Length = 680
Score = 90.1 bits (222), Expect = 1e-17
Identities = 118/449 (26%), Positives = 190/449 (42%), Gaps = 51/449 (11%)
Query: 109 DVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNNCDWQEFDGGLAARRWYATNHILPD 168
D++C S++ G +V TGG ND +T ++D+ ++ W + R Y ++ + D
Sbjct: 266 DMFCPGISMDGNGQIVVTGG-NDAKKT-SLYDSSSD-SWIP-GPDMQVARGYQSSATMSD 321
Query: 169 GRQIIIGGRKQFNYEFYPKNNIGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANN 228
GR IGG ++ + KN VY T+ A+ N +L D ++N
Sbjct: 322 GRVFTIGG--SWSGGVFEKNG-EVYSPSSKTWTSLPNAKVN----PMLTADKQGLYRSDN 374
Query: 229 RAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGS 288
A LF + K V F P Y +SGSG + QS A +CG A
Sbjct: 375 HAWLFGWKKGSV---FQAGPSTAMNWYYTSGSGDVKSAGKRQSNRGVAPDAMCGNAVMYD 431
Query: 289 YQKASKREFLGA-------LNTCARIKI-----TDPNPTWVVETMPRARVMGDMVMLPNG 336
K F G+ T A I T PN + + AR V+LP+G
Sbjct: 432 AVKGKILTFGGSPDYQDSDATTNAHIITLGEPGTSPNTVFASNGLYFARTFHTSVVLPDG 491
Query: 337 DVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTPRMYHSTAILVRDG 396
I G G + PV P +Y P F QNP+ R+YHS ++L+ DG
Sbjct: 492 STFITGGQRRGIPFED--STPVFTPEIYV---PEQDTFYKQNPNSIVRVYHSISLLLPDG 546
Query: 397 RVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANV--RPRIVASTSELQKHGQK 454
RV GG N+ + F+P+YL N+ RP+I ++++ K G +
Sbjct: 547 RVFNGGGGLCGDCTTNH------FDAQIFTPNYLYNSNGNLATRPKITRTSTQSVKVGGR 600
Query: 455 LGLRFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPG 514
+ + + D ++ +++ TH+ + +QR + L G +Y QV P
Sbjct: 601 ITI------STDSSISKASLIRYGTATHTVNTDQRRIPLTLTNNG---GNSYSFQV--PS 649
Query: 515 SPILAPPGFYLLFVVHKE-IPSEGIWIQI 542
+A PG+++LFV++ +PS I++
Sbjct: 650 DSGVALPGYWMLFVMNSAGVPSVASTIRV 678
>PPA_ECOLI (P07102) Periplasmic appA protein precursor [Includes:
Phosphoanhydride phosphohydrolase (EC 3.1.3.2) (pH 2.5
acid phosphatase) (AP); 4-phytase (EC 3.1.3.26)]
Length = 432
Score = 32.3 bits (72), Expect = 3.2
Identities = 20/86 (23%), Positives = 36/86 (41%), Gaps = 1/86 (1%)
Query: 458 RFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVE-GTTYDVQVTMPGSP 516
R + LD +T P + ++ +L + + N+ G ++ T+PG P
Sbjct: 287 RSRATPLLDLIKTALTPHPPQKQAYGVTLPTSVLFIAGHDTNLANLGGALELNWTLPGQP 346
Query: 517 ILAPPGFYLLFVVHKEIPSEGIWIQI 542
PPG L+F + + WIQ+
Sbjct: 347 DNTPPGGELVFERWRRLSDNSQWIQV 372
>SUCC_THETH (P25126) Succinyl-CoA synthetase beta chain (EC 6.2.1.5)
(SCS-beta)
Length = 378
Score = 30.8 bits (68), Expect = 9.4
Identities = 22/76 (28%), Positives = 36/76 (46%), Gaps = 12/76 (15%)
Query: 192 VYRLPFLEQTNDAGAENNL------YPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFP 245
++R P L + + AE+ L Y F + +DGN+ I N ++ YT ++V R
Sbjct: 214 LFRHPDLAELREVEAEHPLEVEASNYGFAYVKLDGNIGIIGNGAGLVM-YTLDLVNRV-- 270
Query: 246 QIPGGDPRSYPSSGSG 261
GG P ++ G G
Sbjct: 271 ---GGKPANFLDIGGG 283
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,003,199
Number of Sequences: 164201
Number of extensions: 3223250
Number of successful extensions: 6271
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 6265
Number of HSP's gapped (non-prelim): 3
length of query: 543
length of database: 59,974,054
effective HSP length: 115
effective length of query: 428
effective length of database: 41,090,939
effective search space: 17586921892
effective search space used: 17586921892
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC124216.10


