

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124214.11 + phase: 0 /pseudo
(240 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
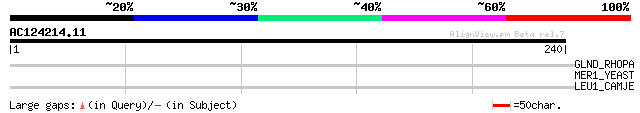
Score E
Sequences producing significant alignments: (bits) Value
GLND_RHOPA (P62223) [Protein-PII] uridylyltransferase (EC 2.7.7.... 32 1.4
MER1_YEAST (P16523) Meiotic recombination 1 protein 29 8.8
LEU1_CAMJE (Q9PLV9) 2-isopropylmalate synthase (EC 2.3.3.13) (Al... 29 8.8
>GLND_RHOPA (P62223) [Protein-PII] uridylyltransferase (EC 2.7.7.59)
(PII uridylyl-transferase) (Uridylyl removing enzyme)
(UTase)
Length = 929
Score = 32.0 bits (71), Expect = 1.4
Identities = 24/80 (30%), Positives = 38/80 (47%), Gaps = 1/80 (1%)
Query: 143 LKESGVRSSASASLLYLPLFLDYKRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEK 202
L E+ R ++S S L F+ + EI + DR +I V G++ + I K
Sbjct: 812 LPEAVARRASSGSKAKLRAFVVEPEV-EINNNWSDRYTVIEVSGLDRPGLLYQLTTAISK 870
Query: 203 LNLKVTNSSFMTFGSCALDI 222
LNL + ++ TFG A D+
Sbjct: 871 LNLNIASAHVATFGERARDV 890
>MER1_YEAST (P16523) Meiotic recombination 1 protein
Length = 270
Score = 29.3 bits (64), Expect = 8.8
Identities = 26/88 (29%), Positives = 38/88 (42%), Gaps = 15/88 (17%)
Query: 157 LYLPLFLDYKRLPEIEARFCDRNVLIRVHCEKIKGVVEKTIHKIEKLNLKVTNSSFMTFG 216
LYL L LDY + FC VH +KIKGV+ I L L + + F
Sbjct: 40 LYLKLALDYSFFRKNLLEFC-------VHLDKIKGVIRPNYDTIYILCLLEVDLLNLVFT 92
Query: 217 SCALDITI---IAQPTL-----SFYVFH 236
L+I + +++ L +FY +H
Sbjct: 93 DNILEICLPRFVSREDLRVFNNTFYTYH 120
>LEU1_CAMJE (Q9PLV9) 2-isopropylmalate synthase (EC 2.3.3.13)
(Alpha-isopropylmalate synthase) (Alpha-IPM synthetase)
Length = 511
Score = 29.3 bits (64), Expect = 8.8
Identities = 15/24 (62%), Positives = 16/24 (66%)
Query: 46 LLVTLHNTIHTFIQTSTLELQWKL 69
L V H IHTFI TSTL +Q KL
Sbjct: 89 LKVAKHFRIHTFIATSTLHMQDKL 112
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.364 0.163 0.551
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,028,052
Number of Sequences: 164201
Number of extensions: 629639
Number of successful extensions: 4529
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 4526
Number of HSP's gapped (non-prelim): 3
length of query: 240
length of database: 59,974,054
effective HSP length: 107
effective length of query: 133
effective length of database: 42,404,547
effective search space: 5639804751
effective search space used: 5639804751
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 64 (29.3 bits)
Medicago: description of AC124214.11


