

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123571.7 - phase: 0
(247 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
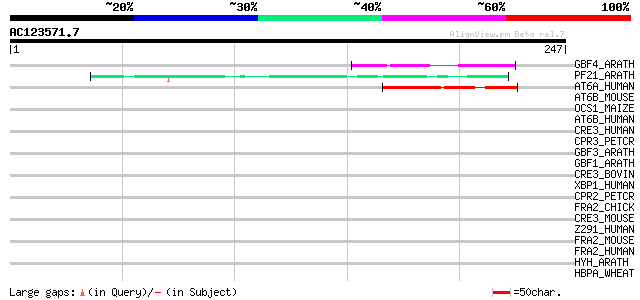
Score E
Sequences producing significant alignments: (bits) Value
GBF4_ARATH (P42777) G-box binding factor 4 (AtbZIP40) 48 3e-05
PF21_ARATH (Q04088) Possible transcription factor PosF21 (AtbZIP59) 45 2e-04
AT6A_HUMAN (P18850) Cyclic-AMP-dependent transcription factor AT... 44 5e-04
AT6B_MOUSE (O35451) Cyclic-AMP-dependent transcription factor AT... 42 0.001
OCS1_MAIZE (P24068) Ocs-element binding factor 1 (OCSBF-1) 42 0.002
AT6B_HUMAN (Q99941) Cyclic-AMP-dependent transcription factor AT... 40 0.004
CRE3_HUMAN (O43889) Cyclic-AMP responsive element binding protei... 40 0.007
CPR3_PETCR (Q99091) Light-inducible protein CPRF-3 40 0.007
GBF3_ARATH (P42776) G-box binding factor 3 (AtbZIP55) 39 0.009
GBF1_ARATH (P42774) G-box binding factor 1 (AtbZIP41) 39 0.009
CRE3_BOVIN (Q8SQ19) Cyclic-AMP responsive element binding protei... 39 0.009
XBP1_HUMAN (P17861) X box binding protein-1 (XBP-1) (TREB5 protein) 39 0.015
CPR2_PETCR (Q99090) Light-inducible protein CPRF-2 38 0.020
FRA2_CHICK (P18625) Fos-related antigen 2 38 0.026
CRE3_MOUSE (Q61817) Cyclic-AMP responsive element binding protei... 38 0.026
Z291_HUMAN (Q9BY12) Zinc finger protein 291 37 0.034
FRA2_MOUSE (P47930) Fos-related antigen 2 37 0.034
FRA2_HUMAN (P15408) Fos-related antigen 2 37 0.034
HYH_ARATH (Q8W191) Transcription factor HY5-like (HY5 homolog) 37 0.044
HBPA_WHEAT (P23922) Transcription factor HBP-1a (Histone-specifi... 37 0.044
>GBF4_ARATH (P42777) G-box binding factor 4 (AtbZIP40)
Length = 270
Score = 47.8 bits (112), Expect = 3e-05
Identities = 33/73 (45%), Positives = 42/73 (57%), Gaps = 13/73 (17%)
Query: 153 GKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLL 212
G+ EA D + +R+ KRMIKNRESAARSR RKQ AY ELE L
Sbjct: 176 GRVMMEAMDKAAAQRQ-KRMIKNRESAARSRERKQ------------AYQVELETLAAKL 222
Query: 213 QEENARLRRQQQE 225
+EEN +L ++ +E
Sbjct: 223 EEENEQLLKEIEE 235
>PF21_ARATH (Q04088) Possible transcription factor PosF21 (AtbZIP59)
Length = 398
Score = 45.1 bits (105), Expect = 2e-04
Identities = 56/189 (29%), Positives = 75/189 (39%), Gaps = 41/189 (21%)
Query: 37 QDFLTRPLTLDPTKSLDYSSNNSSSSVASDQNN---NNASFYCPISTTPPPLVTALSLNS 93
+D L+ L +D S S SS+ V N +T P T SL
Sbjct: 96 EDLLSMYLDMDKFNS----SATSSAQVGEPSGTAWKNETMMQTGTGSTSNPQNTVNSLGE 151
Query: 94 RPDFLYDPLIRHNKHNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFG 153
RP IRH Q Q G N++ ++ + D + S S T
Sbjct: 152 RPR------IRH----------QHSQSMDGSMNINEMLMSGNEDDSAIDAKKSMSAT--- 192
Query: 154 KRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQ 213
+ E I P +R KR+ NR+SAARS+ RK T ++F ELE+KVQ LQ
Sbjct: 193 -KLAELALIDP--KRAKRIWANRQSAARSKERK----TRYIF--------ELERKVQTLQ 237
Query: 214 EENARLRRQ 222
E L Q
Sbjct: 238 TEATTLSAQ 246
>AT6A_HUMAN (P18850) Cyclic-AMP-dependent transcription factor ATF-6
alpha (Activating transcription factor 6 alpha)
(ATF6-alpha)
Length = 670
Score = 43.5 bits (101), Expect = 5e-04
Identities = 27/60 (45%), Positives = 39/60 (65%), Gaps = 5/60 (8%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
RR +RMIKNRESA +SR +K+E + L ++ A +E EQ L++EN L+RQ E+
Sbjct: 308 RRQQRMIKNRESACQSRKKKKEYMLG-LEARLKAALSENEQ----LKKENGTLKRQLDEV 362
>AT6B_MOUSE (O35451) Cyclic-AMP-dependent transcription factor ATF-6
beta (Activating transcription factor 6 beta)
(ATF6-beta) (cAMP responsive element binding
protein-like 1) (cAMP response element binding
protein-related protein) (Creb-rp)
Length = 699
Score = 42.4 bits (98), Expect = 0.001
Identities = 27/66 (40%), Positives = 40/66 (59%), Gaps = 5/66 (7%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R +RMIKNRESA +SR +K+E + L ++ A + +Q L+ ENA LRR+ + L
Sbjct: 324 KRQQRMIKNRESACQSRRKKKEYLQG-LEARLQAVLADNQQ----LRRENAALRRRLEAL 378
Query: 227 WEAESG 232
SG
Sbjct: 379 LAENSG 384
>OCS1_MAIZE (P24068) Ocs-element binding factor 1 (OCSBF-1)
Length = 151
Score = 41.6 bits (96), Expect = 0.002
Identities = 24/60 (40%), Positives = 35/60 (58%), Gaps = 12/60 (20%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
RR KR + NRESA RSR RKQ+ + +EL Q+V LQ +NAR+ + +++
Sbjct: 26 RREKRRLSNRESARRSRLRKQQ------------HLDELVQEVARLQADNARVAARARDI 73
>AT6B_HUMAN (Q99941) Cyclic-AMP-dependent transcription factor ATF-6
beta (Activating transcription factor 6 beta)
(ATF6-beta) (cAMP responsive element binding
protein-like 1) (cAMP response element binding
protein-related protein) (Creb-rp) (G13 protein)
Length = 703
Score = 40.4 bits (93), Expect = 0.004
Identities = 23/60 (38%), Positives = 34/60 (56%), Gaps = 12/60 (20%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R +RMIKNRESA +SR +K+E Y LE ++Q + +N +LRR+ L
Sbjct: 327 KRQQRMIKNRESACQSRRKKKE------------YLQGLEARLQAVLADNQQLRRENAAL 374
>CRE3_HUMAN (O43889) Cyclic-AMP responsive element binding protein 3
(Luman protein) (Transcription factor LZIP-alpha)
Length = 395
Score = 39.7 bits (91), Expect = 0.007
Identities = 23/62 (37%), Positives = 37/62 (59%), Gaps = 2/62 (3%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSF--LFSKFSAYTNELEQKVQLLQEENARLRRQQQ 224
+R +R I+N+ SA SR +K+ + K++A EL+ KVQLL+E+N L Q +
Sbjct: 176 KRVRRKIRNKRSAQESRRKKKVYVGGLESRVLKYTAQNMELQNKVQLLEEQNLSLLDQLR 235
Query: 225 EL 226
+L
Sbjct: 236 KL 237
>CPR3_PETCR (Q99091) Light-inducible protein CPRF-3
Length = 296
Score = 39.7 bits (91), Expect = 0.007
Identities = 24/63 (38%), Positives = 35/63 (55%), Gaps = 12/63 (19%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R +R NRESA RSR RKQ K ++EL++++ L +EN LR+ Q +
Sbjct: 198 KRQRRKQSNRESARRSRLRKQAK------------SDELQERLDNLSKENRILRKNLQRI 245
Query: 227 WEA 229
EA
Sbjct: 246 SEA 248
>GBF3_ARATH (P42776) G-box binding factor 3 (AtbZIP55)
Length = 382
Score = 39.3 bits (90), Expect = 0.009
Identities = 26/62 (41%), Positives = 32/62 (50%), Gaps = 12/62 (19%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R +R NRESA RSR RKQ A T EL +KV+ L EN LR + +L
Sbjct: 261 KRERRKQSNRESARRSRLRKQ------------AETEELARKVEALTAENMALRSELNQL 308
Query: 227 WE 228
E
Sbjct: 309 NE 310
>GBF1_ARATH (P42774) G-box binding factor 1 (AtbZIP41)
Length = 315
Score = 39.3 bits (90), Expect = 0.009
Identities = 25/60 (41%), Positives = 31/60 (51%), Gaps = 12/60 (20%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R KR NRESA RSR RKQ A +L+Q+V+ L EN LR + Q L
Sbjct: 224 KRQKRKQSNRESARRSRLRKQ------------AECEQLQQRVESLSNENQSLRDELQRL 271
>CRE3_BOVIN (Q8SQ19) Cyclic-AMP responsive element binding protein 3
(Luman protein)
Length = 365
Score = 39.3 bits (90), Expect = 0.009
Identities = 23/62 (37%), Positives = 37/62 (59%), Gaps = 2/62 (3%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSF--LFSKFSAYTNELEQKVQLLQEENARLRRQQQ 224
+R +R I+N++SA SR +K+ + K++A EL+ KVQLL+E+N L Q +
Sbjct: 152 KRVRRKIRNKKSAQESRRKKKVYVGGLESRVLKYTAQNLELQNKVQLLEEQNLSLLDQLR 211
Query: 225 EL 226
L
Sbjct: 212 RL 213
>XBP1_HUMAN (P17861) X box binding protein-1 (XBP-1) (TREB5 protein)
Length = 260
Score = 38.5 bits (88), Expect = 0.015
Identities = 25/71 (35%), Positives = 39/71 (54%), Gaps = 12/71 (16%)
Query: 162 ISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRR 221
+SP E+ +R +KNR +A +R RK+ +++ ELEQ+V L+EEN +L
Sbjct: 66 LSPEEKALRRKLKNRVAAQTARDRKKARMS------------ELEQQVVDLEEENQKLLL 113
Query: 222 QQQELWEAESG 232
+ Q L E G
Sbjct: 114 ENQLLREKTHG 124
>CPR2_PETCR (Q99090) Light-inducible protein CPRF-2
Length = 401
Score = 38.1 bits (87), Expect = 0.020
Identities = 23/60 (38%), Positives = 34/60 (56%), Gaps = 12/60 (20%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R +RM+ NRESA RSR RKQ A+ ELE +V L+ EN+ L ++ ++
Sbjct: 200 KRVRRMLSNRESARRSRRRKQ------------AHMTELETQVSQLRVENSSLLKRLTDI 247
>FRA2_CHICK (P18625) Fos-related antigen 2
Length = 323
Score = 37.7 bits (86), Expect = 0.026
Identities = 27/88 (30%), Positives = 46/88 (51%), Gaps = 5/88 (5%)
Query: 139 QNVGVPASSSFTCFGKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKF 198
Q GV + T +R E E+R R +N+ +AA+ R R++E L K
Sbjct: 98 QRPGVIKTIGTTVGRRRRDEQLSPEEEEKRRIRRERNKLAAAKCRNRRRE-----LTEKL 152
Query: 199 SAYTNELEQKVQLLQEENARLRRQQQEL 226
A T LE++ +LQ+E A L++++++L
Sbjct: 153 QAETEVLEEEKSVLQKEIAELQKEKEKL 180
>CRE3_MOUSE (Q61817) Cyclic-AMP responsive element binding protein 3
(Transcription factor LZIP)
Length = 404
Score = 37.7 bits (86), Expect = 0.026
Identities = 21/62 (33%), Positives = 36/62 (57%), Gaps = 2/62 (3%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSF--LFSKFSAYTNELEQKVQLLQEENARLRRQQQ 224
+R +R I+N+ +A SR +K+ + K++A EL+ KVQ L+E+N L Q +
Sbjct: 187 KRVRRKIRNKRAAQESRKKKKVYVVGLESRVLKYTAQNRELQNKVQRLEEQNLSLLDQLR 246
Query: 225 EL 226
+L
Sbjct: 247 KL 248
>Z291_HUMAN (Q9BY12) Zinc finger protein 291
Length = 1399
Score = 37.4 bits (85), Expect = 0.034
Identities = 21/60 (35%), Positives = 34/60 (56%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
E++ + K RE AAR RAR +E+ + L + EL++K+QL +E+ R +Q E
Sbjct: 702 EQQRQEKEKAREDAARERARDREERLAALTAAQQEAMEELQKKIQLKHDESIRRHMEQIE 761
>FRA2_MOUSE (P47930) Fos-related antigen 2
Length = 326
Score = 37.4 bits (85), Expect = 0.034
Identities = 22/61 (36%), Positives = 38/61 (62%), Gaps = 5/61 (8%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
E+R R +N+ +AA+ R R++E L K A T ELE++ LQ+E A L++++++
Sbjct: 125 EKRRIRRERNKLAAAKCRNRRRE-----LTEKLQAETEELEEEKSGLQKEIAELQKEKEK 179
Query: 226 L 226
L
Sbjct: 180 L 180
>FRA2_HUMAN (P15408) Fos-related antigen 2
Length = 326
Score = 37.4 bits (85), Expect = 0.034
Identities = 22/61 (36%), Positives = 38/61 (62%), Gaps = 5/61 (8%)
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
E+R R +N+ +AA+ R R++E L K A T ELE++ LQ+E A L++++++
Sbjct: 125 EKRRIRRERNKLAAAKCRNRRRE-----LTEKLQAETEELEEEKSGLQKEIAELQKEKEK 179
Query: 226 L 226
L
Sbjct: 180 L 180
>HYH_ARATH (Q8W191) Transcription factor HY5-like (HY5 homolog)
Length = 149
Score = 37.0 bits (84), Expect = 0.044
Identities = 37/128 (28%), Positives = 61/128 (46%), Gaps = 18/128 (14%)
Query: 105 HNKH----NNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVP----ASSSFTCFGKRF 156
H KH ++ +LL+ G S C +++S D V P + + +R
Sbjct: 16 HKKHKTEESDEELLMVPDMEAAG----STCVLSSS-ADDGVNNPELDQTQNGVSTAKRRR 70
Query: 157 GEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTN---ELEQKVQLLQ 213
G P + R KR+++NR SA ++R RK+ + S L S+ + N +LE+K+ L
Sbjct: 71 GRNP-VDKEYRSLKRLLRNRVSAQQARERKKVYV-SDLESRANELQNNNDQLEEKISTLT 128
Query: 214 EENARLRR 221
EN LR+
Sbjct: 129 NENTMLRK 136
>HBPA_WHEAT (P23922) Transcription factor HBP-1a (Histone-specific
transcription factor HBP1)
Length = 349
Score = 37.0 bits (84), Expect = 0.044
Identities = 22/54 (40%), Positives = 29/54 (52%), Gaps = 12/54 (22%)
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLR 220
++ KR + NRESA RSR RKQ A EL Q+ + L+ EN+ LR
Sbjct: 254 KKQKRKLSNRESARRSRLRKQ------------AECEELGQRAEALKSENSSLR 295
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.127 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,438,513
Number of Sequences: 164201
Number of extensions: 1153114
Number of successful extensions: 5158
Number of sequences better than 10.0: 171
Number of HSP's better than 10.0 without gapping: 40
Number of HSP's successfully gapped in prelim test: 131
Number of HSP's that attempted gapping in prelim test: 4948
Number of HSP's gapped (non-prelim): 311
length of query: 247
length of database: 59,974,054
effective HSP length: 107
effective length of query: 140
effective length of database: 42,404,547
effective search space: 5936636580
effective search space used: 5936636580
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC123571.7


