

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123570.7 + phase: 0
(601 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
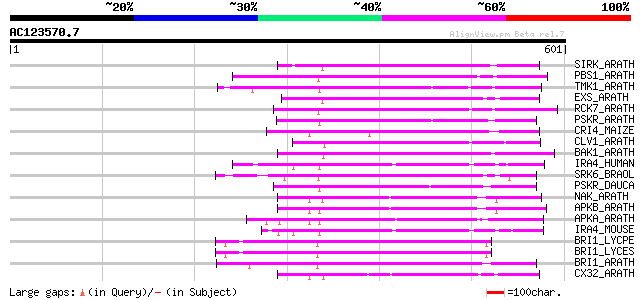
Score E
Sequences producing significant alignments: (bits) Value
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 207 8e-53
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 204 7e-52
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 202 2e-51
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 200 8e-51
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 199 2e-50
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 191 4e-48
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 189 2e-47
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 187 5e-47
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 187 9e-47
IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (... 184 6e-46
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 181 5e-45
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 181 5e-45
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 181 5e-45
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 180 8e-45
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 179 1e-44
IRA4_MOUSE (Q8R4K2) Interleukin-1 receptor-associated kinase-4 (... 177 9e-44
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 173 1e-42
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 172 2e-42
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 170 9e-42
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 166 2e-40
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 207 bits (526), Expect = 8e-53
Identities = 116/288 (40%), Positives = 169/288 (58%), Gaps = 14/288 (4%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKAT---KEFLAELKVL 347
F Y E+ N TNNF IG+GGFG+VY+ +NGE+ A+K + ++ KEF AE+ +L
Sbjct: 564 FKYSEVVNITNNFERV--IGKGGFGKVYHGVINGEQVAVKVLSEESAQGYKEFRAEVDLL 621
Query: 348 TRVHHVNLVRLIGYCVE-GSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSARG 406
RVHH NL L+GYC E + L+YEY+ N NLG +L LSW R+KI+LD+A+G
Sbjct: 622 MRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSFILSWEERLKISLDAAQG 681
Query: 407 LEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYM 466
LEY+H P +HRD+K NILL++ AK+ADFGL++ S + +AG+ GY+
Sbjct: 682 LEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQISTVVAGSIGYL 741
Query: 467 PPE-YAYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPI 525
PE Y+ ++ K DVY+ GVVL E+I+ + A+ + + V D
Sbjct: 742 DPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKTEKVHISDHVRSILANGD----- 796
Query: 526 EGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
++ +VD RL + Y + +KM+++A CT QRP MS VV+ L
Sbjct: 797 --IRGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVVMEL 842
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 204 bits (518), Expect = 7e-52
Identities = 129/350 (36%), Positives = 191/350 (53%), Gaps = 14/350 (4%)
Query: 242 RKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPANMIGIRVEKSGEFSYEELANATN 301
+K++ S+ I G E + T G S G+ + F++ ELA AT
Sbjct: 25 QKKQSQPTVSNNISGLPSGGEKLSSKTNGGSKRELLLPRDGLGQIAAHTFAFRELAAATM 84
Query: 302 NFNMANKIGQGGFGEVYYAELN--GEKAAIKKMD---MKATKEFLAELKVLTRVHHVNLV 356
NF+ +G+GGFG VY L+ G+ A+K++D ++ +EFL E+ +L+ +HH NLV
Sbjct: 85 NFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVLMLSLLHHPNLV 144
Query: 357 RLIGYCVEGSL-FLVYEYIDNGNLGQHLRS--SDGEPLSWSIRVKIALDSARGLEYIHEH 413
LIGYC +G LVYE++ G+L HL D E L W++R+KIA +A+GLE++H+
Sbjct: 145 NLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAGAAKGLEFLHDK 204
Query: 414 TVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY- 472
P I+RD KS NILLD+ F K++DFGL KL G S + + GT+GY PEYA
Sbjct: 205 ANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRVMGTYGYCAPEYAMT 264
Query: 473 GSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLV 532
G ++ K DVY+FGVV ELI+ + A+ G + LV +F+ KL
Sbjct: 265 GQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGE--QNLVAWARPLFNDRRK---FIKLA 319
Query: 533 DPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTTED 582
DPRL +P +++ +A +C RP ++ VV AL+ L + D
Sbjct: 320 DPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVVTALSYLANQAYD 369
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 202 bits (514), Expect = 2e-51
Identities = 146/380 (38%), Positives = 217/380 (56%), Gaps = 40/380 (10%)
Query: 226 LLSFCIYVTYYRKKKIRKQEFLSEESSAIFGQVKND---------EVSGNATY--GTSDS 274
LL FC Y KK +K+ SE S+A+ ++ V+G++ G SD+
Sbjct: 500 LLVFCWY------KKRQKRFSGSESSNAVVVHPRHSGSDNESVKITVAGSSVSVGGISDT 553
Query: 275 AS--PANMIG--IRVEKSGEF--SYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKA 327
+ + +G I++ ++G S + L + TNNF+ N +G GGFG VY EL+ G K
Sbjct: 554 YTLPGTSEVGDNIQMVEAGNMLISIQVLRSVTNNFSSDNILGSGGFGVVYKGELHDGTKI 613
Query: 328 AIKKMDM-----KATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQ 381
A+K+M+ K EF +E+ VLT+V H +LV L+GYC++G+ LVYEY+ G L +
Sbjct: 614 AVKRMENGVIAGKGFAEFKSEIAVLTKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSR 673
Query: 382 HLR--SSDG-EPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKV 438
HL S +G +PL W R+ +ALD ARG+EY+H ++IHRD+K NILL + AKV
Sbjct: 674 HLFEWSEEGLKPLLWKQRLTLALDVARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKV 733
Query: 439 ADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAA 497
ADFGL +L G S+ T +AGTFGY+ PEYA G V++K+DVY+FGV+L ELI+ + +
Sbjct: 734 ADFGLVRLAPEGKGSIET-RIAGTFGYLAPEYAVTGRVTTKVDVYSFGVILMELITGRKS 792
Query: 498 VIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLG-DNYPIDHVFKMAQLAKVCT 556
+ E + LV F+ ++ KK +D + D + V +A+LA C
Sbjct: 793 --LDESQPEESIHLVSWFKRMYINKE--ASFKKAIDTTIDLDEETLASVHTVAELAGHCC 848
Query: 557 NSDPQQRPNMSSVVVALTTL 576
+P QRP+M V L++L
Sbjct: 849 AREPYQRPDMGHAVNILSSL 868
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 200 bits (509), Expect = 8e-51
Identities = 126/289 (43%), Positives = 175/289 (59%), Gaps = 18/289 (6%)
Query: 295 ELANATNNFNMANKIGQGGFGEVYYAELNGEKA-AIKKMDMKAT---KEFLAELKVLTRV 350
++ AT++F+ N IG GGFG VY A L GEK A+KK+ T +EF+AE++ L +V
Sbjct: 909 DIVEATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQGNREFMAEMETLGKV 968
Query: 351 HHVNLVRLIGYC-VEGSLFLVYEYIDNGNLGQHLRSSDG--EPLSWSIRVKIALDSARGL 407
H NLV L+GYC LVYEY+ NG+L LR+ G E L WS R+KIA+ +ARGL
Sbjct: 969 KHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGAARGL 1028
Query: 408 EYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMP 467
++H +P IHRDIK+ NILLD +F KVADFGL +LI A S V TV +AGTFGY+P
Sbjct: 1029 AFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTV-IAGTFGYIP 1087
Query: 468 PEYAYGS-VSSKIDVYAFGVVLYELISAK--AAVIMGEDSGADLKGLVVLFEEVFDQPHP 524
PEY + ++K DVY+FGV+L EL++ K E G +L G + + +Q
Sbjct: 1088 PEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGNLVGWAI---QKINQGKA 1144
Query: 525 IEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
++ ++DP L + ++ Q+A +C P +RPNM V+ AL
Sbjct: 1145 VD----VIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKAL 1189
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 199 bits (505), Expect = 2e-50
Identities = 119/319 (37%), Positives = 185/319 (57%), Gaps = 16/319 (5%)
Query: 286 EKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN--GEKAAIKKMD---MKATKEF 340
+K+ F+++ELA AT NF +G+GGFG+V+ + + AIK++D ++ +EF
Sbjct: 86 KKAQTFTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIREF 145
Query: 341 LAELKVLTRVHHVNLVRLIGYCVEGSL-FLVYEYIDNGNLGQHLR--SSDGEPLSWSIRV 397
+ E+ L+ H NLV+LIG+C EG LVYEY+ G+L HL S +PL W+ R+
Sbjct: 146 VVEVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRM 205
Query: 398 KIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTV 457
KIA +ARGLEY+H+ P I+RD+K NILL +++ K++DFGL K+ +G + +
Sbjct: 206 KIAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVST 265
Query: 458 NMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFE 516
+ GT+GY P+YA G ++ K D+Y+FGVVL ELI+ + A+ + LV
Sbjct: 266 RVMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAI--DNTKTRKDQNLVGWAR 323
Query: 517 EVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTL 576
+F K+VDP L YP+ +++ ++ +C P RP +S VV+AL L
Sbjct: 324 PLFKDR---RNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVVLALNFL 380
Query: 577 TSTTEDWD--ITSIFKNPN 593
S+ D + +S KNP+
Sbjct: 381 ASSKYDPNSPSSSSGKNPS 399
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 191 bits (486), Expect = 4e-48
Identities = 118/290 (40%), Positives = 174/290 (59%), Gaps = 17/290 (5%)
Query: 290 EFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDM---KATKEFLAELK 345
E SY++L ++TN+F+ AN IG GGFG VY A L +G+K AIKK+ + +EF AE++
Sbjct: 721 ELSYDDLLDSTNSFDQANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQIEREFEAEVE 780
Query: 346 VLTRVHHVNLVRLIGYCV-EGSLFLVYEYIDNGNLGQHLRSSDGEP--LSWSIRVKIALD 402
L+R H NLV L G+C + L+Y Y++NG+L L + P L W R++IA
Sbjct: 781 TLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGPALLKWKTRLRIAQG 840
Query: 403 SARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGT 462
+A+GL Y+HE P +HRDIKS NILLD+NF + +ADFGL +L+ + V T ++ GT
Sbjct: 841 AAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVST-DLVGT 899
Query: 463 FGYMPPEYAYGSVSS-KIDVYAFGVVLYELISAKAAVIMGEDSGA-DLKGLVVLFEEVFD 520
GY+PPEY SV++ K DVY+FGVVL EL++ K V M + G DL VV +
Sbjct: 900 LGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMKHE-- 957
Query: 521 QPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVV 570
++ DP + +F++ ++A +C + +P+QRP +V
Sbjct: 958 -----SRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLV 1002
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 189 bits (480), Expect = 2e-47
Identities = 118/308 (38%), Positives = 184/308 (59%), Gaps = 23/308 (7%)
Query: 279 NMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELN-----GEKAAIKKMD 333
+M +++ ++ EFSYEEL AT F+ +++G+G F V+ L K AIK D
Sbjct: 481 DMEDLKIRRAQEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASD 540
Query: 334 MK-ATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDG--- 388
+K ++KEF EL +L+R++H +L+ L+GYC +GS LVYE++ +G+L QHL D
Sbjct: 541 VKKSSKEFHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLK 600
Query: 389 EPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLID 448
+ L+W+ RV IA+ +ARG+EY+H + P IHRDIKS NIL+D++ A+VADFGL+ L
Sbjct: 601 KRLNWARRVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGP 660
Query: 449 AGISSVPTVNMAGTFGYMPPE-YAYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGAD 507
A + + AGT GY+ PE Y +++K DVY+FGVVL E++S + A+ M + G
Sbjct: 661 ADSGTPLSELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFEEGNI 720
Query: 508 LKGLVVLFE--EVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPN 565
++ V L + ++F ++DP L ++ + K+A +A C + RP+
Sbjct: 721 VEWAVPLIKAGDIF----------AILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPS 770
Query: 566 MSSVVVAL 573
M V AL
Sbjct: 771 MDKVTTAL 778
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 187 bits (476), Expect = 5e-47
Identities = 116/276 (42%), Positives = 161/276 (58%), Gaps = 11/276 (3%)
Query: 307 NKIGQGGFGEVYYAEL-NGEKAAIKKMDMKATKE----FLAELKVLTRVHHVNLVRLIGY 361
N IG+GG G VY + N AIK++ + T F AE++ L R+ H ++VRL+GY
Sbjct: 696 NIIGKGGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRLLGY 755
Query: 362 CV-EGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIH 420
+ + L+YEY+ NG+LG+ L S G L W R ++A+++A+GL Y+H P +H
Sbjct: 756 VANKDTNLLLYEYMPNGSLGELLHGSKGGHLQWETRHRVAVEAAKGLCYLHHDCSPLILH 815
Query: 421 RDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAYG-SVSSKI 479
RD+KS NILLD +F A VADFGL K + G +S ++AG++GY+ PEYAY V K
Sbjct: 816 RDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAYTLKVDEKS 875
Query: 480 DVYAFGVVLYELISAKAAVIMGE-DSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGD 538
DVY+FGVVL ELI+ K V GE G D+ V EE QP + +VDPRL
Sbjct: 876 DVYSFGVVLLELIAGKKPV--GEFGEGVDIVRWVRNTEEEITQPSDAAIVVAIVDPRL-T 932
Query: 539 NYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALT 574
YP+ V + ++A +C + RP M VV LT
Sbjct: 933 GYPLTSVIHVFKIAMMCVEEEAAARPTMREVVHMLT 968
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 187 bits (474), Expect = 9e-47
Identities = 120/311 (38%), Positives = 179/311 (56%), Gaps = 16/311 (5%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDMKATK----EFLAELK 345
FS EL A++NF+ N +G+GGFG+VY L +G A+K++ + T+ +F E++
Sbjct: 277 FSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGELQFQTEVE 336
Query: 346 VLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRS--SDGEPLSWSIRVKIALD 402
+++ H NL+RL G+C+ + LVY Y+ NG++ LR PL W R +IAL
Sbjct: 337 MISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWPKRQRIALG 396
Query: 403 SARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGT 462
SARGL Y+H+H P IHRD+K+ NILLD+ F A V DFGL KL+D + V T + GT
Sbjct: 397 SARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTA-VRGT 455
Query: 463 FGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQ 521
G++ PEY + G S K DV+ +GV+L ELI+ + A + + D L+ + + +
Sbjct: 456 IGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVKGLLKE 515
Query: 522 PHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL--TTLTST 579
+ L+ LVD L NY + V ++ Q+A +CT S P +RP MS VV L L
Sbjct: 516 ----KKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRMLEGDGLAER 571
Query: 580 TEDWDITSIFK 590
E+W +F+
Sbjct: 572 WEEWQKEEMFR 582
>IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4) (NY-REN-64 antigen)
Length = 460
Score = 184 bits (467), Expect = 6e-46
Identities = 126/354 (35%), Positives = 190/354 (53%), Gaps = 31/354 (8%)
Query: 242 RKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPANM-IGIRVEKSGEFSYEELANAT 300
+KQ ++ + V+N E S Y DS+SP N + + + FS+ EL N T
Sbjct: 122 QKQMPFCDKDRTLMTPVQNLEQS----YMPPDSSSPENKSLEVSDTRFHSFSFYELKNVT 177
Query: 301 NNFNM------ANKIGQGGFGEVYYAELNGEKAAIKKMDM-------KATKEFLAELKVL 347
NNF+ NK+G+GGFG VY +N A+KK+ + ++F E+KV+
Sbjct: 178 NNFDERPISVGGNKMGEGGFGVVYKGYVNNTTVAVKKLAAMVDITTEELKQQFDQEIKVM 237
Query: 348 TRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGEP-LSWSIRVKIALDSAR 405
+ H NLV L+G+ +G L LVY Y+ NG+L L DG P LSW +R KIA +A
Sbjct: 238 AKCQHENLVELLGFSSDGDDLCLVYVYMPNGSLLDRLSCLDGTPPLSWHMRCKIAQGAAN 297
Query: 406 GLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGY 465
G+ ++HE+ +IHRDIKS NILLD+ F AK++DFGL + + +V T + GT Y
Sbjct: 298 GINFLHENH---HIHRDIKSANILLDEAFTAKISDFGLARASEKFAQTVMTSRIVGTTAY 354
Query: 466 MPPEYAYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPI 525
M PE G ++ K D+Y+FGVVL E+I+ AV D + + L+ + EE+ D+ I
Sbjct: 355 MAPEALRGEITPKSDIYSFGVVLLEIITGLPAV----DEHREPQLLLDIKEEIEDEEKTI 410
Query: 526 EGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTST 579
E +D ++ D V M +A C + +RP++ V L +T++
Sbjct: 411 E---DYIDKKMND-ADSTSVEAMYSVASQCLHEKKNKRPDIKKVQQLLQEMTAS 460
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 181 bits (459), Expect = 5e-45
Identities = 123/372 (33%), Positives = 195/372 (52%), Gaps = 51/372 (13%)
Query: 224 LILLSFCIYVTYYRKKKIRKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPANMIGI 283
L+L+ FC++ RKQ+ + +I +N + N ++
Sbjct: 458 LLLIMFCLWK--------RKQKRAKASAISIANTQRNQNLPMNEM-----------VLSS 498
Query: 284 RVEKSGEFSYEEL----------ANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKM 332
+ E SGE+ +EEL AT NF+ NK+GQGGFG VY L +G++ A+K++
Sbjct: 499 KREFSGEYKFEELELPLIEMETVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKRL 558
Query: 333 D---MKATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHL-RSSD 387
++ T EF+ E+ ++ R+ H+NLV+++G C+EG L+YEY++N +L +L +
Sbjct: 559 SKTSVQGTDEFMNEVTLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKTR 618
Query: 388 GEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLI 447
L+W+ R I ARGL Y+H+ + IHRD+K NILLDKN K++DFG+ ++
Sbjct: 619 RSKLNWNERFDITNGVARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIF 678
Query: 448 DAGISSVPTVNMAGTFGYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKA-AVIMGEDSG 505
+ + T+ + GT+GYM PEYA YG S K DV++FGV++ E++S K D
Sbjct: 679 ERDETEANTMKVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYE 738
Query: 506 ADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDN-------YPIDHVFKMAQLAKVCTNS 558
DL V + + +E +VDP + D+ + V K Q+ +C
Sbjct: 739 NDLLSYVW---SRWKEGRALE----IVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQE 791
Query: 559 DPQQRPNMSSVV 570
+ RP MSSVV
Sbjct: 792 LAEHRPAMSSVV 803
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 181 bits (459), Expect = 5e-45
Identities = 114/294 (38%), Positives = 170/294 (57%), Gaps = 17/294 (5%)
Query: 286 EKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDM---KATKEFL 341
+ + E S +++ +T++FN AN IG GGFG VY A L +G K AIK++ + +EF
Sbjct: 726 DSNNELSLDDILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQMDREFQ 785
Query: 342 AELKVLTRVHHVNLVRLIGYC-VEGSLFLVYEYIDNGNLGQHLRSS-DGEP-LSWSIRVK 398
AE++ L+R H NLV L+GYC + L+Y Y+DNG+L L DG P L W R++
Sbjct: 786 AEVETLSRAQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLR 845
Query: 399 IALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVN 458
IA +A GL Y+H+ P +HRDIKS NILL F A +ADFGL +LI + V T +
Sbjct: 846 IARGAAEGLAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTHV-TTD 904
Query: 459 MAGTFGYMPPEYAYGSVSS-KIDVYAFGVVLYELISAKAAVIMGEDSGA-DLKGLVVLFE 516
+ GT GY+PPEY SV++ K DVY+FGVVL EL++ + + + + G+ DL V+
Sbjct: 905 LVGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWVL--- 961
Query: 517 EVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVV 570
Q + ++ DP + D + + + ++A C +P+ RP +V
Sbjct: 962 ----QMKTEKRESEIFDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLV 1011
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 181 bits (459), Expect = 5e-45
Identities = 116/308 (37%), Positives = 173/308 (55%), Gaps = 32/308 (10%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELN-----------GEKAAIKKMDMKAT-- 337
FS EL +AT NF + +G+GGFG V+ ++ G A+K+++ +
Sbjct: 56 FSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLNQEGFQG 115
Query: 338 -KEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHL--RSSDGEPLSW 393
+E+LAE+ L ++ H NLV+LIGYC+E LVYE++ G+L HL R + +PLSW
Sbjct: 116 HREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFYQPLSW 175
Query: 394 SIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISS 453
+ RV++AL +ARGL ++H + P I+RD K+ NILLD N+ AK++DFGL + G +S
Sbjct: 176 NTRVRMALGAARGLAFLH-NAQPQVIYRDFKASNILLDSNYNAKLSDFGLARDGPMGDNS 234
Query: 454 VPTVNMAGTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLV 512
+ + GT GY PEY A G +S K DVY+FGVVL EL+S + A+ + G
Sbjct: 235 HVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVGE------ 288
Query: 513 VLFEEVFDQPHPI----EGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSS 568
+ D P L +++DPRL Y + K+A LA C + D + RP M+
Sbjct: 289 ---HNLVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAKSRPTMNE 345
Query: 569 VVVALTTL 576
+V + L
Sbjct: 346 IVKTMEEL 353
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 180 bits (457), Expect = 8e-45
Identities = 114/313 (36%), Positives = 178/313 (56%), Gaps = 32/313 (10%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELN-----------GEKAAIKKMDM---KA 336
F++ EL AT NF + +G+GGFG V+ ++ G A+KK++ +
Sbjct: 57 FTFAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTASKPGTGVVIAVKKLNQDGWQG 116
Query: 337 TKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHL--RSSDGEPLSW 393
+E+LAE+ L + H NLV+LIGYC+E LVYE++ G+L HL R S +PLSW
Sbjct: 117 HQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGSYFQPLSW 176
Query: 394 SIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISS 453
++R+K+AL +A+GL ++H + + I+RD K+ NILLD + AK++DFGL K G S
Sbjct: 177 TLRLKVALGAAKGLAFLH-NAETSVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPTGDKS 235
Query: 454 VPTVNMAGTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLV 512
+ + GT+GY PEY A G +++K DVY++GVVL E++S + AV G
Sbjct: 236 HVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSGRRAVDKNRPPGE------ 289
Query: 513 VLFEEVFDQPHPI----EGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSS 568
+++ + P+ L +++D RL D Y ++ K+A LA C + + RPNM+
Sbjct: 290 ---QKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRCLTFEIKLRPNMNE 346
Query: 569 VVVALTTLTSTTE 581
VV L + + E
Sbjct: 347 VVSHLEHIQTLNE 359
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 179 bits (455), Expect = 1e-44
Identities = 127/356 (35%), Positives = 191/356 (52%), Gaps = 42/356 (11%)
Query: 257 QVKNDEVSGNATYGTSDSAS---PANMIGIRVEKSGE-----------FSYEELANATNN 302
QVK + + Y D S A+ + +R E FS+ EL +AT N
Sbjct: 8 QVKAESSGASTKYDAKDIGSLGSKASSVSVRPSPRTEGEILQSPNLKSFSFAELKSATRN 67
Query: 303 FNMANKIGQGGFGEVYYAELN-----------GEKAAIKKMDM---KATKEFLAELKVLT 348
F + +G+GGFG V+ ++ G A+KK++ + +E+LAE+ L
Sbjct: 68 FRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGLVIAVKKLNQDGWQGHQEWLAEVNYLG 127
Query: 349 RVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHL--RSSDGEPLSWSIRVKIALDSAR 405
+ H +LV+LIGYC+E LVYE++ G+L HL R +PLSW +R+K+AL +A+
Sbjct: 128 QFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGLYFQPLSWKLRLKVALGAAK 187
Query: 406 GLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGY 465
GL ++H I+RD K+ NILLD + AK++DFGL K G S + + GT GY
Sbjct: 188 GLAFLHSSETRV-IYRDFKTSNILLDSEYNAKLSDFGLAKDGPIGDKSHVSTRVMGTHGY 246
Query: 466 MPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHP 524
PEY A G +++K DVY+FGVVL EL+S + AV SG + LV + +P+
Sbjct: 247 AAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRPSGE--RNLV-----EWAKPYL 299
Query: 525 IEGLK--KLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTS 578
+ K +++D RL D Y ++ K+A L+ C ++ + RPNMS VV L + S
Sbjct: 300 VNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSEVVSHLEHIQS 355
>IRA4_MOUSE (Q8R4K2) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4)
Length = 459
Score = 177 bits (448), Expect = 9e-44
Identities = 121/325 (37%), Positives = 176/325 (53%), Gaps = 33/325 (10%)
Query: 273 DSASPANMIGIRVEKSG----EFSYEELANATNNFNM------ANKIGQGGFGEVYYAEL 322
DS+SP N VE S FS+ EL + TNNF+ N++G+GGFG VY +
Sbjct: 149 DSSSPDNR---SVESSDTRFHSFSFHELKSITNNFDEQPASAGGNRMGEGGFGVVYKGCV 205
Query: 323 NGEKAAIKKMDM-------KATKEFLAELKVLTRVHHVNLVRLIGYCVEG-SLFLVYEYI 374
N A+KK+ + ++F E+KV+ H NLV L+G+ + +L LVY Y+
Sbjct: 206 NNTIVAVKKLGAMVEISTEELKQQFDQEIKVMATCQHENLVELLGFSSDSDNLCLVYAYM 265
Query: 375 DNGNLGQHLRSSDGEP-LSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKN 433
NG+L L DG P LSW R K+A +A G+ ++HE+ +IHRDIKS NILLDK+
Sbjct: 266 PNGSLLDRLSCLDGTPPLSWHTRCKVAQGTANGIRFLHENH---HIHRDIKSANILLDKD 322
Query: 434 FCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAYGSVSSKIDVYAFGVVLYELIS 493
F AK++DFGL + +V T + GT YM PE G ++ K D+Y+FGVVL ELI+
Sbjct: 323 FTAKISDFGLARASARLAQTVMTSRIVGTTAYMAPEALRGEITPKSDIYSFGVVLLELIT 382
Query: 494 AKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAK 553
AAV D + + L+ + EE+ D+ IE D ++ D P V M A
Sbjct: 383 GLAAV----DENREPQLLLDIKEEIEDEEKTIE---DYTDEKMSDADPAS-VEAMYSAAS 434
Query: 554 VCTNSDPQQRPNMSSVVVALTTLTS 578
C + +RP+++ V L +++
Sbjct: 435 QCLHEKKNRRPDIAKVQQLLQEMSA 459
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 173 bits (439), Expect = 1e-42
Identities = 115/315 (36%), Positives = 175/315 (55%), Gaps = 20/315 (6%)
Query: 224 LILLSFCIY----VTYYRKKKIRKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPAN 279
L+ FCI+ V KK+ RK+E E A + + +A TS + +
Sbjct: 808 LLFSLFCIFGLIIVAIETKKRRRKKEAALE---AYMDGHSHSATANSAWKFTSAREALSI 864
Query: 280 MIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKM---DMK 335
+ + + ++ +L ATN F+ + +G GGFG+VY A+L +G AIKK+ +
Sbjct: 865 NLAAFEKPLRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQ 924
Query: 336 ATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQ--HLRSSDGEPLS 392
+EF AE++ + ++ H NLV L+GYC G LVYEY+ G+L H R G L+
Sbjct: 925 GDREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLN 984
Query: 393 WSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGIS 452
W R KIA+ +ARGL ++H + +P IHRD+KS N+LLD+N A+V+DFG+ +L+ A +
Sbjct: 985 WPARRKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDT 1044
Query: 453 SVPTVNMAGTFGYMPPEYAYG-SVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGL 511
+ +AGT GY+PPEY S+K DVY++GVVL EL++ K + +L G
Sbjct: 1045 HLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNLVGW 1104
Query: 512 VVL-----FEEVFDQ 521
V L +VFD+
Sbjct: 1105 VKLHAKGKITDVFDR 1119
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 172 bits (437), Expect = 2e-42
Identities = 115/315 (36%), Positives = 175/315 (55%), Gaps = 20/315 (6%)
Query: 224 LILLSFCIY----VTYYRKKKIRKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPAN 279
L+ FCI+ V KK+ RK+E E A + + +A TS + +
Sbjct: 808 LLFSLFCIFGLIIVAIETKKRRRKKEAALE---AYMDGHSHSATANSAWKFTSAREALSI 864
Query: 280 MIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKM---DMK 335
+ + + ++ +L ATN F+ + +G GGFG+VY A+L +G AIKK+ +
Sbjct: 865 NLAAFEKPLRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQ 924
Query: 336 ATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQ--HLRSSDGEPLS 392
+EF AE++ + ++ H NLV L+GYC G LVYEY+ G+L H R G L+
Sbjct: 925 GDREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLN 984
Query: 393 WSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGIS 452
W R KIA+ +ARGL ++H + +P IHRD+KS N+LLD+N A+V+DFG+ +L+ A +
Sbjct: 985 WPARRKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDT 1044
Query: 453 SVPTVNMAGTFGYMPPEYAYG-SVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGL 511
+ +AGT GY+PPEY S+K DVY++GVVL EL++ K + +L G
Sbjct: 1045 HLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNLVGW 1104
Query: 512 VVL-----FEEVFDQ 521
V L +VFD+
Sbjct: 1105 VKLHAKGKITDVFDR 1119
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 170 bits (431), Expect = 9e-42
Identities = 120/361 (33%), Positives = 189/361 (52%), Gaps = 23/361 (6%)
Query: 225 ILLSF-CIYVTYYRKKKIRKQEFLSEESSAIFGQV---KNDEVSGNATYGTSDSASPANM 280
+L SF CI+ +++RK+ E ++ + D + N + + ++
Sbjct: 800 LLFSFVCIFGLILVGREMRKRRRKKEAELEMYAEGHGNSGDRTANNTNWKLTGVKEALSI 859
Query: 281 IGIRVEKS-GEFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKM---DMK 335
EK + ++ +L ATN F+ + IG GGFG+VY A L +G AIKK+ +
Sbjct: 860 NLAAFEKPLRKLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQ 919
Query: 336 ATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQ--HLRSSDGEPLS 392
+EF+AE++ + ++ H NLV L+GYC G LVYE++ G+L H G L+
Sbjct: 920 GDREFMAEMETIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLN 979
Query: 393 WSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGIS 452
WS R KIA+ SARGL ++H + P IHRD+KS N+LLD+N A+V+DFG+ +L+ A +
Sbjct: 980 WSTRRKIAIGSARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDT 1039
Query: 453 SVPTVNMAGTFGYMPPEYAYG-SVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGL 511
+ +AGT GY+PPEY S+K DVY++GVVL EL++ K + +L G
Sbjct: 1040 HLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSPDFGDNNLVGW 1099
Query: 512 VVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHV--FKMAQLAKVCTNSDPQQRPNMSSV 569
V + H + + DP L P + + ++A C + +RP M V
Sbjct: 1100 V--------KQHAKLRISDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQV 1151
Query: 570 V 570
+
Sbjct: 1152 M 1152
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 166 bits (419), Expect = 2e-40
Identities = 107/299 (35%), Positives = 162/299 (53%), Gaps = 23/299 (7%)
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELN-----------GEKAAIKKMDMKATK- 338
+++ +L AT NF + +GQGGFG+VY ++ G AIK+++ ++ +
Sbjct: 74 YNFLDLKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRLNSESVQG 133
Query: 339 --EFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGEPLSWSI 395
E+ +E+ L + H NLV+L+GYC E L LVYE++ G+L HL + +P W +
Sbjct: 134 FAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLESHLFRRN-DPFPWDL 192
Query: 396 RVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVP 455
R+KI + +ARGL ++H I+RD K+ NILLD N+ AK++DFGL KL A S
Sbjct: 193 RIKIVIGAARGLAFLHS-LQREVIYRDFKASNILLDSNYDAKLSDFGLAKLGPADEKSHV 251
Query: 456 TVNMAGTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVL 514
T + GT+GY PEY A G + K DV+AFGVVL E+++ A G + LV
Sbjct: 252 TTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTAHNTKRPRGQE--SLVDW 309
Query: 515 FEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
H + K+++D + Y +MA++ C DP+ RP+M VV L
Sbjct: 310 LRPELSNKHRV---KQIMDKGIKGQYTTKVATEMARITLSCIEPDPKNRPHMKEVVEVL 365
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,016,764
Number of Sequences: 164201
Number of extensions: 3037502
Number of successful extensions: 11433
Number of sequences better than 10.0: 1641
Number of HSP's better than 10.0 without gapping: 977
Number of HSP's successfully gapped in prelim test: 664
Number of HSP's that attempted gapping in prelim test: 7673
Number of HSP's gapped (non-prelim): 1930
length of query: 601
length of database: 59,974,054
effective HSP length: 116
effective length of query: 485
effective length of database: 40,926,738
effective search space: 19849467930
effective search space used: 19849467930
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC123570.7


