

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123570.11 + phase: 0
(398 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
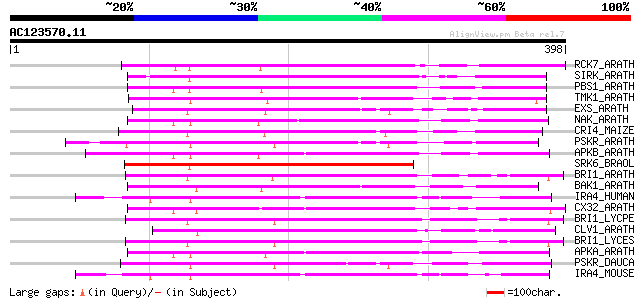
Sequences producing significant alignments: (bits) Value
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 201 4e-51
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 200 5e-51
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 195 2e-49
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 189 8e-48
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 187 3e-47
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 176 7e-44
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 175 2e-43
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 169 1e-41
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 166 1e-40
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 165 2e-40
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 165 2e-40
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 165 2e-40
IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (... 164 3e-40
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 162 1e-39
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 162 1e-39
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 162 2e-39
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 162 2e-39
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 159 9e-39
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 159 1e-38
IRA4_MOUSE (Q8R4K2) Interleukin-1 receptor-associated kinase-4 (... 157 3e-38
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 201 bits (510), Expect = 4e-51
Identities = 128/327 (39%), Positives = 184/327 (56%), Gaps = 29/327 (8%)
Query: 81 KSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELR--GQKIAIKKMK---MQATREFL 135
K+ F+++ELA AT NF +G+GGFG+V+ G + Q +AIK++ +Q REF+
Sbjct: 87 KAQTFTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIREFV 146
Query: 136 SELKVLTSVHHWNLVHLIGYCVEGFL-FLVYEYMENGNLSQHLH--NSEKEPMTLSTRMK 192
E+ L+ H NLV LIG+C EG LVYEYM G+L HLH S K+P+ +TRMK
Sbjct: 147 VEVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMK 206
Query: 193 IALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVH 252
IA ARGLEY+HD P I+RD+K NILL E++ K++DFGL K+ + + +
Sbjct: 207 IAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVSTR 266
Query: 253 VAGTFGYMPPENAY-GRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNE 311
V GT+GY P+ A G+++ K D+Y+FGVVL ELI+ + A ID T KT +
Sbjct: 267 VMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKA---IDNT---------KTRK 314
Query: 312 SIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQ 371
+ LF + N K+VDP L Y + + + ++ C+ P
Sbjct: 315 DQNLVGWARPLFKDRRN--------FPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTM 366
Query: 372 RPTMRDVVVSLMELNSSIDNKNSTGSA 398
RP + DVV++L L SS + NS S+
Sbjct: 367 RPVVSDVVLALNFLASSKYDPNSPSSS 393
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 200 bits (509), Expect = 5e-51
Identities = 120/306 (39%), Positives = 175/306 (56%), Gaps = 29/306 (9%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMK---MQATREFLSELKVL 141
F Y E+ N T+NF + IG+GGFG+VY+G + G+++A+K + Q +EF +E+ +L
Sbjct: 564 FKYSEVVNITNNFE--RVIGKGGFGKVYHGVINGEQVAVKVLSEESAQGYKEFRAEVDLL 621
Query: 142 TSVHHWNLVHLIGYCVE-GFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARG 200
VHH NL L+GYC E + L+YEYM N NL +L ++ R+KI+LD A+G
Sbjct: 622 MRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSFILSWEERLKISLDAAQG 681
Query: 201 LEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYM 260
LEY+H+ P +HRD+K NILLNE K+ADFGL++ S + VAG+ GY+
Sbjct: 682 LEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQISTVVAGSIGYL 741
Query: 261 PPENAYGR-ISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSL 319
PE R ++ K DVY+ GVVL E+I+ + A+ KTE K + S D +S+
Sbjct: 742 DPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIAS-SKTE--------KVHIS-DHVRSI 791
Query: 320 VALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV 379
+A D+R +VD RL Y + S KM+++A AC QRPTM VV
Sbjct: 792 LA------------NGDIRGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVV 839
Query: 380 VSLMEL 385
+ L ++
Sbjct: 840 MELKQI 845
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 195 bits (495), Expect = 2e-49
Identities = 124/310 (40%), Positives = 175/310 (56%), Gaps = 29/310 (9%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGEL--RGQKIAIKKMK---MQATREFLSELK 139
F++ ELA AT NF +G+GGFG VY G L GQ +A+K++ +Q REFL E+
Sbjct: 74 FAFRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVL 133
Query: 140 VLTSVHHWNLVHLIGYCVEGFL-FLVYEYMENGNLSQHLHN--SEKEPMTLSTRMKIALD 196
+L+ +HH NLV+LIGYC +G LVYE+M G+L HLH+ +KE + + RMKIA
Sbjct: 134 MLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAG 193
Query: 197 VARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGT 256
A+GLE++HD + P I+RD KS NILL+E F K++DFGL KL + + + V GT
Sbjct: 194 AAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRVMGT 253
Query: 257 FGYMPPENAY-GRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDE 315
+GY PE A G+++ K DVY+FGVV ELI+ + A+ +E
Sbjct: 254 YGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAI----------------DSEMPHG 297
Query: 316 YKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTM 375
++LVA + N I KL DPRL + ++ + +A CI RP +
Sbjct: 298 EQNLVAWARPLFNDRRKFI----KLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLI 353
Query: 376 RDVVVSLMEL 385
DVV +L L
Sbjct: 354 ADVVTALSYL 363
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 189 bits (481), Expect = 8e-48
Identities = 130/317 (41%), Positives = 185/317 (58%), Gaps = 39/317 (12%)
Query: 86 SYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKMKM-----QATREFLSELK 139
S + L + T+NFS +G GGFG VY GEL G KIA+K+M+ + EF SE+
Sbjct: 577 SIQVLRSVTNNFSSDNILGSGGFGVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIA 636
Query: 140 VLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNSEKE---PMTLSTRMKIAL 195
VLT V H +LV L+GYC++G LVYEYM G LS+HL +E P+ R+ +AL
Sbjct: 637 VLTKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRLTLAL 696
Query: 196 DVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAG 255
DVARG+EY+H + +IHRD+K NILL ++ KVADFGL +L + T +AG
Sbjct: 697 DVARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIET-RIAG 755
Query: 256 TFGYMPPENAY-GRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSL-EIKTNESI 313
TFGY+ PE A GR++ K+DVY+FGV+L ELI+ + KSL E + ESI
Sbjct: 756 TFGYLAPEYAVTGRVTTKVDVYSFGVILMELITGR-------------KSLDESQPEESI 802
Query: 314 DEYKSLVALFDEV-MNQTGDCIDDLRKLVDPRLGYN-YSIDSISKMAKLAKACINRDPKQ 371
LV+ F + +N+ +K +D + + ++ S+ +A+LA C R+P Q
Sbjct: 803 ----HLVSWFKRMYINKEA----SFKKAIDTTIDLDEETLASVHTVAELAGHCCAREPYQ 854
Query: 372 RPTMR---DVVVSLMEL 385
RP M +++ SL+EL
Sbjct: 855 RPDMGHAVNILSSLVEL 871
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 187 bits (476), Expect = 3e-47
Identities = 128/307 (41%), Positives = 178/307 (57%), Gaps = 33/307 (10%)
Query: 89 ELANATDNFSLAKKIGQGGFGEVYYGELRGQK-IAIKKM---KMQATREFLSELKVLTSV 144
++ ATD+FS IG GGFG VY L G+K +A+KK+ K Q REF++E++ L V
Sbjct: 909 DIVEATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQGNREFMAEMETLGKV 968
Query: 145 HHWNLVHLIGYC-VEGFLFLVYEYMENGNLSQHLHNSEK--EPMTLSTRMKIALDVARGL 201
H NLV L+GYC LVYEYM NG+L L N E + S R+KIA+ ARGL
Sbjct: 969 KHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGAARGL 1028
Query: 202 EYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMP 261
++H +P IHRDIK+ NILL+ +F KVADFGL +L A S +TV +AGTFGY+P
Sbjct: 1029 AFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTV-IAGTFGYIP 1087
Query: 262 PENAYGRISR---KIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKS 318
PE YG+ +R K DVY+FGV+L EL++ K + T +FK E +
Sbjct: 1088 PE--YGQSARATTKGDVYSFGVILLELVTGK------EPTGPDFKE---------SEGGN 1130
Query: 319 LVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDV 378
LV + +NQ G +D ++DP L +S ++ ++A C+ P +RP M DV
Sbjct: 1131 LVGWAIQKINQ-GKAVD----VIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDV 1185
Query: 379 VVSLMEL 385
+ +L E+
Sbjct: 1186 LKALKEI 1192
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 176 bits (447), Expect = 7e-44
Identities = 118/320 (36%), Positives = 178/320 (54%), Gaps = 39/320 (12%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGEL-----------RGQKIAIKKMKM---QA 130
FS EL +AT NF +G+GGFG V+ G + G IA+K++ Q
Sbjct: 56 FSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLNQEGFQG 115
Query: 131 TREFLSELKVLTSVHHWNLVHLIGYCVEG-FLFLVYEYMENGNLSQHL--HNSEKEPMTL 187
RE+L+E+ L + H NLV LIGYC+E LVYE+M G+L HL + +P++
Sbjct: 116 HREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFYQPLSW 175
Query: 188 STRMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSA 247
+TR+++AL ARGL ++H+ + P I+RD K+ NILL+ N+ K++DFGL + +++
Sbjct: 176 NTRVRMALGAARGLAFLHN-AQPQVIYRDFKASNILLDSNYNAKLSDFGLARDGPMGDNS 234
Query: 248 DNTVHVAGTFGYMPPEN-AYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLE 306
+ V GT GY PE A G +S K DVY+FGVVL EL+S + A+ K
Sbjct: 235 HVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDK------------ 282
Query: 307 IKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACIN 366
N+ + E+ + + N+ L +++DPRL YS+ K+A LA CI+
Sbjct: 283 ---NQPVGEHNLVDWARPYLTNKR-----RLLRVMDPRLQGQYSLTRALKIAVLALDCIS 334
Query: 367 RDPKQRPTMRDVVVSLMELN 386
D K RPTM ++V ++ EL+
Sbjct: 335 IDAKSRPTMNEIVKTMEELH 354
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 175 bits (444), Expect = 2e-43
Identities = 110/317 (34%), Positives = 180/317 (56%), Gaps = 38/317 (11%)
Query: 79 VDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKM-----KMQATR 132
+ ++ EFSYEEL AT FS ++G+G F V+ G LR G +A+K+ ++++
Sbjct: 487 IRRAQEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASDVKKSSK 546
Query: 133 EFLSELKVLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNSE---KEPMTLS 188
EF +EL +L+ ++H +L++L+GYC +G LVYE+M +G+L QHLH + K+ + +
Sbjct: 547 EFHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKKRLNWA 606
Query: 189 TRMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSAD 248
R+ IA+ ARG+EY+H ++ P IHRDIKS NIL++E+ +VADFGL+ L A +
Sbjct: 607 RRVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPADSGTP 666
Query: 249 NTVHVAGTFGYMPPENAYGR---ISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSL 305
+ AGT GY+ PE Y R ++ K DVY+FGVVL E++S + A+ +
Sbjct: 667 LSELPAGTLGYLDPE--YYRLHYLTTKSDVYSFGVVLLEILSGRKAI-----------DM 713
Query: 306 EIKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACI 365
+ + ++ L+ D+ ++DP L ++++ K+A +A C+
Sbjct: 714 QFEEGNIVEWAVPLIK------------AGDIFAILDPVLSPPSDLEALKKIASVACKCV 761
Query: 366 NRDPKQRPTMRDVVVSL 382
K RP+M V +L
Sbjct: 762 RMRGKDRPSMDKVTTAL 778
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 169 bits (427), Expect = 1e-41
Identities = 123/351 (35%), Positives = 187/351 (53%), Gaps = 43/351 (12%)
Query: 41 RKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKS--PEFSYEELANATDNFS 98
R+++GE + + +SM ++ + GSK + + +S E SY++L ++T++F
Sbjct: 683 RRRSGEVDPEIEESESM-------NRKELGEIGSKLVVLFQSNDKELSYDDLLDSTNSFD 735
Query: 99 LAKKIGQGGFGEVYYGELR-GQKIAIKKMKM---QATREFLSELKVLTSVHHWNLVHLIG 154
A IG GGFG VY L G+K+AIKK+ Q REF +E++ L+ H NLV L G
Sbjct: 736 QANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQIEREFEAEVETLSRAQHPNLVLLRG 795
Query: 155 YCV-EGFLFLVYEYMENGNLSQHLHNSEKEPMTLS--TRMKIALDVARGLEYIHDHSVPV 211
+C + L+Y YMENG+L LH P L TR++IA A+GL Y+H+ P
Sbjct: 796 FCFYKNDRLLIYSYMENGSLDYWLHERNDGPALLKWKTRLRIAQGAAKGLLYLHEGCDPH 855
Query: 212 YIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYGRIS- 270
+HRDIKS NILL+ENF +ADFGL +L + +T + GT GY+PPE YG+ S
Sbjct: 856 ILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVST-DLVGTLGYIPPE--YGQASV 912
Query: 271 --RKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMN 328
K DVY+FGVVL EL++ K V + + K L V+
Sbjct: 913 ATYKGDVYSFGVVLLELLTDKRPV-------------------DMCKPKGCRDLISWVVK 953
Query: 329 QTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV 379
+ ++ DP + + + ++ ++A C++ +PKQRPT + +V
Sbjct: 954 MKHE--SRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLV 1002
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 166 bits (419), Expect = 1e-40
Identities = 116/351 (33%), Positives = 177/351 (50%), Gaps = 39/351 (11%)
Query: 55 DSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYG 114
DS+ + V TN + + F++ EL AT NF +G+GGFG V+ G
Sbjct: 27 DSLGSKSSSVSIRTNPRTEGEILQSPNLKSFTFAELKAATRNFRPDSVLGEGGFGSVFKG 86
Query: 115 EL-----------RGQKIAIKKMKM---QATREFLSELKVLTSVHHWNLVHLIGYCVEG- 159
+ G IA+KK+ Q +E+L+E+ L H NLV LIGYC+E
Sbjct: 87 WIDEQTLTASKPGTGVVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDE 146
Query: 160 FLFLVYEYMENGNLSQHL--HNSEKEPMTLSTRMKIALDVARGLEYIHDHSVPVYIHRDI 217
LVYE+M G+L HL S +P++ + R+K+AL A+GL ++H+ V I+RD
Sbjct: 147 HRLLVYEFMPRGSLENHLFRRGSYFQPLSWTLRLKVALGAAKGLAFLHNAETSV-IYRDF 205
Query: 218 KSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPEN-AYGRISRKIDVY 276
K+ NILL+ + K++DFGL K + + + + GT+GY PE A G ++ K DVY
Sbjct: 206 KTSNILLDSEYNAKLSDFGLAKDGPTGDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVY 265
Query: 277 AFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGDCIDD 336
++GVVL E++S + AV K N E K + + N+
Sbjct: 266 SYGVVLLEVLSGRRAVDK---------------NRPPGEQKLVEWARPLLANKR-----K 305
Query: 337 LRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNS 387
L +++D RL YS++ K+A LA C+ + K RP M +VV L + +
Sbjct: 306 LFRVIDNRLQDQYSMEEACKVATLALRCLTFEIKLRPNMNEVVSHLEHIQT 356
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 165 bits (418), Expect = 2e-40
Identities = 90/214 (42%), Positives = 135/214 (63%), Gaps = 7/214 (3%)
Query: 83 PEFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKMK---MQATREFLSEL 138
P E + AT+NFS K+GQGGFG VY G L G++IA+K++ +Q T EF++E+
Sbjct: 514 PLIEMETVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKRLSKTSVQGTDEFMNEV 573
Query: 139 KVLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHN-SEKEPMTLSTRMKIALD 196
++ + H NLV ++G C+EG L+YEY+EN +L +L + + + + R I
Sbjct: 574 TLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKTRRSKLNWNERFDITNG 633
Query: 197 VARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGT 256
VARGL Y+H S IHRD+K NILL++N K++DFG+ ++ + + NT+ V GT
Sbjct: 634 VARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFERDETEANTMKVVGT 693
Query: 257 FGYMPPENA-YGRISRKIDVYAFGVVLYELISAK 289
+GYM PE A YG S K DV++FGV++ E++S K
Sbjct: 694 YGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGK 727
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 165 bits (418), Expect = 2e-40
Identities = 117/324 (36%), Positives = 175/324 (53%), Gaps = 31/324 (9%)
Query: 84 EFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKM---KMQATREFLSELK 139
+ ++ +L AT+ F IG GGFG+VY L+ G +AIKK+ Q REF++E++
Sbjct: 870 KLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQGDREFMAEME 929
Query: 140 VLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNSEKEPMTL--STRMKIALD 196
+ + H NLV L+GYC G LVYE+M+ G+L LH+ +K + L STR KIA+
Sbjct: 930 TIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTRRKIAIG 989
Query: 197 VARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGT 256
ARGL ++H + P IHRD+KS N+LL+EN +V+DFG+ +L A ++ + +AGT
Sbjct: 990 SARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGT 1049
Query: 257 FGYMPPENAYG-RISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDE 315
GY+PPE R S K DVY++GVVL EL++ K S + N +
Sbjct: 1050 PGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPT----------DSPDFGDNNLVGW 1099
Query: 316 YKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTM 375
K L + D D DP L I+ + + K+A AC++ +RPTM
Sbjct: 1100 VKQHAKL------RISDVFDPELMKEDPAL----EIELLQHL-KVAVACLDDRAWRRPTM 1148
Query: 376 RDVVVSLMEL--NSSIDNKNSTGS 397
V+ E+ S ID++++ S
Sbjct: 1149 VQVMAMFKEIQAGSGIDSQSTIRS 1172
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 165 bits (418), Expect = 2e-40
Identities = 109/304 (35%), Positives = 163/304 (52%), Gaps = 29/304 (9%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKMKMQATR----EFLSELK 139
FS EL A+DNFS +G+GGFG+VY G L G +A+K++K + T+ +F +E++
Sbjct: 277 FSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGELQFQTEVE 336
Query: 140 VLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHN--SEKEPMTLSTRMKIALD 196
+++ H NL+ L G+C+ LVY YM NG+++ L + P+ R +IAL
Sbjct: 337 MISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWPKRQRIALG 396
Query: 197 VARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGT 256
ARGL Y+HDH P IHRD+K+ NILL+E F V DFGL KL D ++ T V GT
Sbjct: 397 SARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTA-VRGT 455
Query: 257 FGYMPPEN-AYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDE 315
G++ PE + G+ S K DV+ +GV+L ELI+ + A F + ++ +
Sbjct: 456 IGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRA----------FDLARLANDDDVML 505
Query: 316 YKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTM 375
+ L E L LVD L NY + + ++ ++A C P +RP M
Sbjct: 506 LDWVKGLLKE---------KKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKM 556
Query: 376 RDVV 379
+VV
Sbjct: 557 SEVV 560
>IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4) (NY-REN-64 antigen)
Length = 460
Score = 164 bits (416), Expect = 3e-40
Identities = 118/357 (33%), Positives = 175/357 (48%), Gaps = 54/357 (15%)
Query: 48 ESKFPPEDSMTPSTKDVD-KDTNDDNGSKYIWVDKSPEFSYEELANATDNFSL------A 100
E + P DS +P K ++ DT + FS+ EL N T+NF
Sbjct: 142 EQSYMPPDSSSPENKSLEVSDT------------RFHSFSFYELKNVTNNFDERPISVGG 189
Query: 101 KKIGQGGFGEVYYGELRGQKIAIKKMKM-------QATREFLSELKVLTSVHHWNLVHLI 153
K+G+GGFG VY G + +A+KK+ + ++F E+KV+ H NLV L+
Sbjct: 190 NKMGEGGFGVVYKGYVNNTTVAVKKLAAMVDITTEELKQQFDQEIKVMAKCQHENLVELL 249
Query: 154 GYCVEGF-LFLVYEYMENGNLSQHLHNSE-KEPMTLSTRMKIALDVARGLEYIHDHSVPV 211
G+ +G L LVY YM NG+L L + P++ R KIA A G+ ++H++
Sbjct: 250 GFSSDGDDLCLVYVYMPNGSLLDRLSCLDGTPPLSWHMRCKIAQGAANGINFLHENH--- 306
Query: 212 YIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYGRISR 271
+IHRDIKS NILL+E FT K++DFGL + ++ T + GT YM PE G I+
Sbjct: 307 HIHRDIKSANILLDEAFTAKISDFGLARASEKFAQTVMTSRIVGTTAYMAPEALRGEITP 366
Query: 272 KIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTG 331
K D+Y+FGVVL E+I+ AV D+ L+IK E DE K++ D+ MN
Sbjct: 367 KSDIYSFGVVLLEIITGLPAV---DEHREPQLLLDIK-EEIEDEEKTIEDYIDKKMNDAD 422
Query: 332 DCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNSS 388
S+ M +A C++ +RP ++ V L E+ +S
Sbjct: 423 S-------------------TSVEAMYSVASQCLHEKKNKRPDIKKVQQLLQEMTAS 460
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 162 bits (411), Expect = 1e-39
Identities = 112/332 (33%), Positives = 176/332 (52%), Gaps = 40/332 (12%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGEL-----------RGQKIAIKKMKMQATR- 132
+++ +L AT NF +GQGGFG+VY G + G +AIK++ ++ +
Sbjct: 74 YNFLDLKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRLNSESVQG 133
Query: 133 --EFLSELKVLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNSEKEPMTLST 189
E+ SE+ L + H NLV L+GYC E L LVYE+M G+L HL +P
Sbjct: 134 FAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLESHLFR-RNDPFPWDL 192
Query: 190 RMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADN 249
R+KI + ARGL ++H V I+RD K+ NILL+ N+ K++DFGL KL A +
Sbjct: 193 RIKIVIGAARGLAFLHSLQREV-IYRDFKASNILLDSNYDAKLSDFGLAKLGPADEKSHV 251
Query: 250 TVHVAGTFGYMPPE-NAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIK 308
T + GT+GY PE A G + K DV+AFGVVL E+++ A + +
Sbjct: 252 TTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTA----------HNTKRPR 301
Query: 309 TNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRD 368
ES+ ++ L E+ N+ +++++D + Y+ ++MA++ +CI D
Sbjct: 302 GQESLVDW-----LRPELSNK-----HRVKQIMDKGIKGQYTTKVATEMARITLSCIEPD 351
Query: 369 PKQRPTMRDVVVSLMELN--SSIDNKNSTGSA 398
PK RP M++VV L + + + N++ST A
Sbjct: 352 PKNRPHMKEVVEVLEHIQGLNVVPNRSSTKQA 383
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 162 bits (411), Expect = 1e-39
Identities = 110/324 (33%), Positives = 175/324 (53%), Gaps = 31/324 (9%)
Query: 84 EFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKM---KMQATREFLSELK 139
+ ++ +L AT+ F +G GGFG+VY +L+ G +AIKK+ Q REF +E++
Sbjct: 875 KLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEME 934
Query: 140 VLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNSEKEPMTLS--TRMKIALD 196
+ + H NLV L+GYC G LVYEYM+ G+L LH+ +K + L+ R KIA+
Sbjct: 935 TIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIAIG 994
Query: 197 VARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGT 256
ARGL ++H + +P IHRD+KS N+LL+EN +V+DFG+ +L A ++ + +AGT
Sbjct: 995 AARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGT 1054
Query: 257 FGYMPPENAYG-RISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDE 315
GY+PPE R S K DVY++GVVL EL++ K D F + +
Sbjct: 1055 PGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSAD-----FGDNNLVGWVKLHA 1109
Query: 316 YKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTM 375
+ +FD + + I+ I+ + + K+A AC++ +RPTM
Sbjct: 1110 KGKITDVFDRELLKEDASIE---------------IELLQHL-KVACACLDDRHWKRPTM 1153
Query: 376 RDVVVSLMEL--NSSIDNKNSTGS 397
V+ E+ S +D+ ++ G+
Sbjct: 1154 IQVMAMFKEIQAGSGMDSTSTIGA 1177
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 162 bits (409), Expect = 2e-39
Identities = 105/296 (35%), Positives = 163/296 (54%), Gaps = 24/296 (8%)
Query: 103 IGQGGFGEVYYGELRGQ-KIAIKKMKMQATRE----FLSELKVLTSVHHWNLVHLIGYCV 157
IG+GG G VY G + +AIK++ + T F +E++ L + H ++V L+GY
Sbjct: 698 IGKGGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRLLGYVA 757
Query: 158 -EGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHSVPVYIHRD 216
+ L+YEYM NG+L + LH S+ + TR ++A++ A+GL Y+H P+ +HRD
Sbjct: 758 NKDTNLLLYEYMPNGSLGELLHGSKGGHLQWETRHRVAVEAAKGLCYLHHDCSPLILHRD 817
Query: 217 IKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYG-RISRKIDV 275
+KS+NILL+ +F VADFGL K +++ +AG++GY+ PE AY ++ K DV
Sbjct: 818 VKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAYTLKVDEKSDV 877
Query: 276 YAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGDCID 335
Y+FGVVL ELI+ K V EF E +D + + +E+ + I
Sbjct: 878 YSFGVVLLELIAGKKPV-----GEF---------GEGVDIVRWVRNTEEEITQPSDAAI- 922
Query: 336 DLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNSSIDN 391
+ +VDPRL Y + S+ + K+A C+ + RPTMR+VV L S+ N
Sbjct: 923 -VVAIVDPRL-TGYPLTSVIHVFKIAMMCVEEEAAARPTMREVVHMLTNPPKSVAN 976
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 162 bits (409), Expect = 2e-39
Identities = 110/324 (33%), Positives = 175/324 (53%), Gaps = 31/324 (9%)
Query: 84 EFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKM---KMQATREFLSELK 139
+ ++ +L AT+ F +G GGFG+VY +L+ G +AIKK+ Q REF +E++
Sbjct: 875 KLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEME 934
Query: 140 VLTSVHHWNLVHLIGYCVEGF-LFLVYEYMENGNLSQHLHNSEKEPMTLS--TRMKIALD 196
+ + H NLV L+GYC G LVYEYM+ G+L LH+ +K + L+ R KIA+
Sbjct: 935 TIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIAIG 994
Query: 197 VARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGT 256
ARGL ++H + +P IHRD+KS N+LL+EN +V+DFG+ +L A ++ + +AGT
Sbjct: 995 AARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGT 1054
Query: 257 FGYMPPENAYG-RISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDE 315
GY+PPE R S K DVY++GVVL EL++ K D F + +
Sbjct: 1055 PGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSAD-----FGDNNLVGWVKLHA 1109
Query: 316 YKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTM 375
+ +FD + + I+ I+ + + K+A AC++ +RPTM
Sbjct: 1110 KGKITDVFDRELLKEDASIE---------------IELLQHL-KVACACLDDRHWKRPTM 1153
Query: 376 RDVVVSLMEL--NSSIDNKNSTGS 397
V+ E+ S +D+ ++ G+
Sbjct: 1154 IQVMAMFKEIQAGSGMDSTSTIGA 1177
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 159 bits (403), Expect = 9e-39
Identities = 111/321 (34%), Positives = 167/321 (51%), Gaps = 39/321 (12%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGEL-----------RGQKIAIKKMKM---QA 130
FS+ EL +AT NF +G+GGFG V+ G + G IA+KK+ Q
Sbjct: 56 FSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGLVIAVKKLNQDGWQG 115
Query: 131 TREFLSELKVLTSVHHWNLVHLIGYCVEG-FLFLVYEYMENGNLSQHLHNSEK--EPMTL 187
+E+L+E+ L H +LV LIGYC+E LVYE+M G+L HL +P++
Sbjct: 116 HQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGLYFQPLSW 175
Query: 188 STRMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSA 247
R+K+AL A+GL ++H V I+RD K+ NILL+ + K++DFGL K + +
Sbjct: 176 KLRLKVALGAAKGLAFLHSSETRV-IYRDFKTSNILLDSEYNAKLSDFGLAKDGPIGDKS 234
Query: 248 DNTVHVAGTFGYMPPEN-AYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLE 306
+ V GT GY PE A G ++ K DVY+FGVVL EL+S + AV K ++ E +E
Sbjct: 235 HVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDK-NRPSGERNLVE 293
Query: 307 IKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACIN 366
+++ K + +++D RL YS++ K+A L+ C+
Sbjct: 294 WAKPYLVNKRK-------------------IFRVIDNRLQDQYSMEEACKVATLSLRCLT 334
Query: 367 RDPKQRPTMRDVVVSLMELNS 387
+ K RP M +VV L + S
Sbjct: 335 TEIKLRPNMSEVVSHLEHIQS 355
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 159 bits (402), Expect = 1e-38
Identities = 111/320 (34%), Positives = 167/320 (51%), Gaps = 36/320 (11%)
Query: 80 DKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIKKMKM---QATREFL 135
D + E S +++ +T +F+ A IG GGFG VY L G K+AIK++ Q REF
Sbjct: 726 DSNNELSLDDILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQMDREFQ 785
Query: 136 SELKVLTSVHHWNLVHLIGYC-VEGFLFLVYEYMENGNLSQHLHNSEKEPMTLS--TRMK 192
+E++ L+ H NLVHL+GYC + L+Y YM+NG+L LH P +L TR++
Sbjct: 786 AEVETLSRAQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLR 845
Query: 193 IALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVH 252
IA A GL Y+H P +HRDIKS NILL++ F +ADFGL +L T
Sbjct: 846 IARGAAEGLAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLI-LPYDTHVTTD 904
Query: 253 VAGTFGYMPPENAYGRIS---RKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEI-K 308
+ GT GY+PPE YG+ S K DVY+FGVVL EL++ + + +++ K
Sbjct: 905 LVGTLGYIPPE--YGQASVATYKGDVYSFGVVLLELLTGR-------------RPMDVCK 949
Query: 309 TNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRD 368
S D ++ + E ++ DP + + + + ++A C+ +
Sbjct: 950 PRGSRDLISWVLQMKTEKRES---------EIFDPFIYDKDHAEEMLLVLEIACRCLGEN 1000
Query: 369 PKQRPTMRDVVVSLMELNSS 388
PK RPT + +V L ++ S
Sbjct: 1001 PKTRPTTQQLVSWLENIDVS 1020
>IRA4_MOUSE (Q8R4K2) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4)
Length = 459
Score = 157 bits (398), Expect = 3e-38
Identities = 115/355 (32%), Positives = 174/355 (48%), Gaps = 52/355 (14%)
Query: 48 ESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEELANATDNFSL------AK 101
E P DS +P + V+ + FS+ EL + T+NF
Sbjct: 142 EHSCEPPDSSSPDNRSVESSDT-----------RFHSFSFHELKSITNNFDEQPASAGGN 190
Query: 102 KIGQGGFGEVYYGELRGQKIAIKKMKM-------QATREFLSELKVLTSVHHWNLVHLIG 154
++G+GGFG VY G + +A+KK+ + ++F E+KV+ + H NLV L+G
Sbjct: 191 RMGEGGFGVVYKGCVNNTIVAVKKLGAMVEISTEELKQQFDQEIKVMATCQHENLVELLG 250
Query: 155 YCVEGF-LFLVYEYMENGNLSQHLHNSE-KEPMTLSTRMKIALDVARGLEYIHDHSVPVY 212
+ + L LVY YM NG+L L + P++ TR K+A A G+ ++H++ +
Sbjct: 251 FSSDSDNLCLVYAYMPNGSLLDRLSCLDGTPPLSWHTRCKVAQGTANGIRFLHENH---H 307
Query: 213 IHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYGRISRK 272
IHRDIKS NILL+++FT K++DFGL + + T + GT YM PE G I+ K
Sbjct: 308 IHRDIKSANILLDKDFTAKISDFGLARASARLAQTVMTSRIVGTTAYMAPEALRGEITPK 367
Query: 273 IDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGD 332
D+Y+FGVVL ELI+ AAV D+ L+IK E DE K++ DE M+
Sbjct: 368 SDIYSFGVVLLELITGLAAV---DENREPQLLLDIK-EEIEDEEKTIEDYTDEKMSD--- 420
Query: 333 CIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNS 387
DP S+ M A C++ +RP + V L E+++
Sbjct: 421 --------ADPA--------SVEAMYSAASQCLHEKKNRRPDIAKVQQLLQEMSA 459
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.135 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,756,763
Number of Sequences: 164201
Number of extensions: 2209042
Number of successful extensions: 10496
Number of sequences better than 10.0: 1647
Number of HSP's better than 10.0 without gapping: 943
Number of HSP's successfully gapped in prelim test: 704
Number of HSP's that attempted gapping in prelim test: 6607
Number of HSP's gapped (non-prelim): 2035
length of query: 398
length of database: 59,974,054
effective HSP length: 112
effective length of query: 286
effective length of database: 41,583,542
effective search space: 11892893012
effective search space used: 11892893012
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC123570.11


