

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
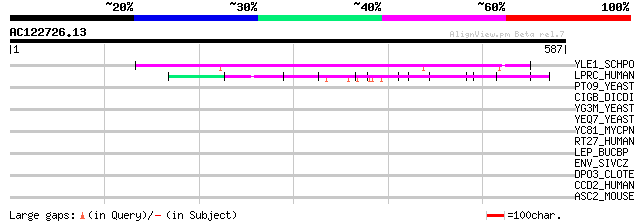
Query= AC122726.13 - phase: 0
(587 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I 76 2e-13
LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP13... 69 3e-11
PT09_YEAST (P32522) PET309 protein, mitochondrial precursor 43 0.003
CIGB_DICDI (Q94481) Protein cigB (Fragment) 43 0.003
YG3M_YEAST (P48237) Hypothetical 101.4 kDa protein in RPL24B-RSR... 42 0.006
YEQ7_YEAST (P40050) Hypothetical 79.5 kDa protein in PTP3-SER3 i... 33 2.7
YC81_MYCPN (P75496) Hypothetical lipoprotein MPN281 precursor (A... 33 2.7
RT27_HUMAN (Q92552) Mitochondrial 28S ribosomal protein S27 (S27... 32 4.6
LEP_BUCBP (Q89AM6) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 32 4.6
ENV_SIVCZ (P17281) Envelope polyprotein GP160 precursor [Contain... 32 4.6
DPO3_CLOTE (Q895K2) DNA polymerase III polC-type (EC 2.7.7.7) (P... 32 6.0
CCD2_HUMAN (Q96LB3) Coiled-coil domain containing protein 2 (Cap... 31 7.9
ASC2_MOUSE (O35885) Achaete-scute homolog 2 (Mash-2) 31 7.9
>YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I
Length = 1261
Score = 76.3 bits (186), Expect = 2e-13
Identities = 91/441 (20%), Positives = 185/441 (41%), Gaps = 25/441 (5%)
Query: 134 ALIIDMLVLAYVKNLELHCAYEAFTRAKDYGFKLSLTSCNPLLSALVKENKIGDVEYVYK 193
+L+ ML A + ++++ + F + + L L+S L K + V+ V++
Sbjct: 772 SLVSKMLTSASPEQVDVNILFFQFGKLIETNKFLHPEVYPTLISVLSKNKRFDAVQRVFE 831
Query: 194 EM--IKRRIHT-NLNTFNIFINGLCRAGKLN-------KAEDAI-EDMKAWGISPNVVTY 242
+ R+I T +L N F+ + A L+ K+ + ++MK G P T+
Sbjct: 832 HSKHLYRKISTKSLEKANWFMALILDAMILSSSFARQFKSSNLFCDNMKMLGYIPRASTF 891
Query: 243 NTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEE 302
L++ +RG A +E + + P+ +N ++ + K F+E
Sbjct: 892 AHLINNSTRRGDTDDATTALNIFEETKRHNVKPSVFLYNAVLSKLGRARRTTECWKLFQE 951
Query: 303 MQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGL-GLKPNIVTYNALINGFCKKK 361
M++ GL P VTY ++IN C G A L+ +M +P + YN +I +
Sbjct: 952 MKESGLLPTSVTYGTVINAACRIGDESLAEKLFAEMENQPNYQPRVAPYNTMIQFEVQTM 1011
Query: 362 MMKE-ATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEG-FSLCSSMLDEGILPNVST 419
+E A ++ + ++ P+ T+ ++DAY + G +++ +P +S
Sbjct: 1012 FNREKALFYYNRLCATDIEPSSHTYKLLMDAYGTLKPVNVGSVKAVLELMERTDVPILSM 1071
Query: 420 YNCLIAGLCRK--QDLQAA----KELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEK 473
+ + D+QAA L + + ++ D + I+ L ND+ +
Sbjct: 1072 HYAAYIHILGNVVSDVQAATSCYMNALAKHDAGEIQLDANLFQSQIESLIANDRIVEGIQ 1131
Query: 474 LLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKE---RKQPNVVTYNVLIK 530
++++M + N N L+ G+ G + A +E E K+P+ TY +++
Sbjct: 1132 IVSDMKRYNVSLNAYIVNALIKGFTKVGMISKARYYFDLLECEGMSGKEPS--TYENMVR 1189
Query: 531 GYCKINKLEAANGLLNEMLEK 551
Y +N A ++ ++ K
Sbjct: 1190 AYLSVNDGRKAMEIVEQLKRK 1210
Score = 35.8 bits (81), Expect = 0.32
Identities = 29/110 (26%), Positives = 43/110 (38%), Gaps = 5/110 (4%)
Query: 457 ILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLK---AALNVRTRM 513
IL + KS N + M LG P T+ L++ G ALN+
Sbjct: 860 ILSSSFARQFKSSNL--FCDNMKMLGYIPRASTFAHLINNSTRRGDTDDATTALNIFEET 917
Query: 514 EKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLEKGLNPNRTTYDIV 563
++ +P+V YN ++ + + L EM E GL P TY V
Sbjct: 918 KRHNVKPSVFLYNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTV 967
>LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP130)
(Leucine-rich PPR-motif containing protein)
Length = 1273
Score = 69.3 bits (168), Expect = 3e-11
Identities = 40/155 (25%), Positives = 72/155 (45%)
Query: 290 DENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIVT 349
+E A + ++ +QK G ++ YN+L+ N D KM ++PN VT
Sbjct: 19 EERTEFAHRIWDTLQKLGAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVT 78
Query: 350 YNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSML 409
Y LI +C ++ A+K+ + ++L F+ ++ + + G ME ++ + M
Sbjct: 79 YQRLIASYCNVGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMR 138
Query: 410 DEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEME 444
D GI P TY L+ K D+ K+ L ++E
Sbjct: 139 DAGIEPGPDTYLALLNAYAEKGDIDHVKQTLEKVE 173
Score = 67.0 bits (162), Expect = 1e-10
Identities = 39/150 (26%), Positives = 69/150 (46%)
Query: 366 ATKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIA 425
A +++D + K V +V +N ++ Y + + M + I PN TY LIA
Sbjct: 25 AHRIWDTLQKLGAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLIA 84
Query: 426 GLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKP 485
C D++ A ++L M+ K L ++ L+ G + NAE +L M + G++P
Sbjct: 85 SYCNVGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDAGIEP 144
Query: 486 NHVTYNTLMDGYCMEGKLKAALNVRTRMEK 515
TY L++ Y +G + ++EK
Sbjct: 145 GPDTYLALLNAYAEKGDIDHVKQTLEKVEK 174
Score = 59.7 bits (143), Expect = 2e-08
Identities = 36/149 (24%), Positives = 67/149 (44%)
Query: 423 LIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLG 482
L+ L ++ + A + + ++ G DV YN L+ +N+ + L +M
Sbjct: 12 LLPELKLEERTEFAHRIWDTLQKLGAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEAN 71
Query: 483 LKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAAN 542
++PN VTY L+ YC G ++ A + M+ + ++ L+ G+ + +E A
Sbjct: 72 IQPNRVTYQRLIASYCNVGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAE 131
Query: 543 GLLNEMLEKGLNPNRTTYDIVRLEMLEKG 571
+L M + G+ P TY + EKG
Sbjct: 132 NILTVMRDAGIEPGPDTYLALLNAYAEKG 160
Score = 58.9 bits (141), Expect = 4e-08
Identities = 38/163 (23%), Positives = 75/163 (45%), Gaps = 6/163 (3%)
Query: 327 KLEEAID----LWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNV 382
KLEE + +WD + LG ++ YNAL+ + + + T + + + PN
Sbjct: 17 KLEERTEFAHRIWDTLQKLGAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNR 76
Query: 383 ITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNE 442
+T+ +I +YC G +E + M + + + ++ L+ G R D++ A+ +L
Sbjct: 77 VTYQRLIASYCNVGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTV 136
Query: 443 MENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLN--EMFNLGL 483
M + G++ TY L++ + + ++ L E F L L
Sbjct: 137 MRDAGIEPGPDTYLALLNAYAEKGDIDHVKQTLEKVEKFELHL 179
Score = 57.0 bits (136), Expect = 1e-07
Identities = 38/151 (25%), Positives = 68/151 (44%), Gaps = 3/151 (1%)
Query: 228 EDMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGF 287
+ ++ G +V YN L+ Y + + F+ +M I PN VT+ LI +
Sbjct: 30 DTLQKLGAVYDVSHYNALLKVYLQNEYK---FSPTDFLAKMEEANIQPNRVTYQRLIASY 86
Query: 288 CKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNI 347
C ++ A K M+ + L +++L+ G G +E A ++ M G++P
Sbjct: 87 CNVGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDAGIEPGP 146
Query: 348 VTYNALINGFCKKKMMKEATKVFDDVSKQEL 378
TY AL+N + +K + + + V K EL
Sbjct: 147 DTYLALLNAYAEKGDIDHVKQTLEKVEKFEL 177
Score = 55.8 bits (133), Expect = 3e-07
Identities = 79/368 (21%), Positives = 137/368 (36%), Gaps = 63/368 (17%)
Query: 169 LTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIE 228
L SC LL L E + ++ + K +++ +N + + D +
Sbjct: 6 LRSCGSLLPELKLEERTEFAHRIWDTLQKLGAVYDVSHYNALLKVYLQNEYKFSPTDFLA 65
Query: 229 DMKAWGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFC 288
M+ I PN VTY L+ YC G K FMK + E F+ L+ G
Sbjct: 66 KMEEANIQPNRVTYQRLIASYCNVGDIEGASKILGFMK---TKDLPVTEAVFSALVTGHA 122
Query: 289 KDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLKPNIV 348
+ ++ A+ M+ G++P TY +L+N G ++ +K+ L
Sbjct: 123 RAGDMENAENILTVMRDAGIEPGPDTYLALLNAYAEKGDIDHVKQTLEKVEKFELHLMDR 182
Query: 349 TYNALINGF-----------------CKKKMMKEA--------TKVFDDVSKQELV---- 379
+I F C+++ + +A T+ +DV+ Q L+
Sbjct: 183 DLLQIIFSFSKAGYLSMSQKFWKKFTCERRYIPDAMNLILLLVTEKLEDVALQILLACPV 242
Query: 380 -----PNV---------ITFNTMIDA---YCKEGMMEEGFSLCSSMLDEGILPNVSTYNC 422
P+V +T NT ++ YCK+ + S P T +C
Sbjct: 243 SKEDGPSVFGSFFLQHCVTMNTPVEKLTDYCKKLKEVQMHS----------FPLQFTLHC 292
Query: 423 LIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLG 482
+ L K DL AK L+ ++ +G + L+ G K + ++L M LG
Sbjct: 293 AL--LANKTDL--AKALMKAVKEEGFPIRPHYFWPLLVGRRKEKNVQGIIEILKGMQELG 348
Query: 483 LKPNHVTY 490
+ P+ TY
Sbjct: 349 VHPDQETY 356
Score = 50.4 bits (119), Expect = 1e-05
Identities = 36/140 (25%), Positives = 61/140 (42%), Gaps = 1/140 (0%)
Query: 412 GILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNA 471
G + +VS YN L+ + + + + L +ME ++ + VTY LI C A
Sbjct: 36 GAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLIASYCNVGDIEGA 95
Query: 472 EKLLNEMFNLGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKERKQPNVVTYNVLIKG 531
K+L M L ++ L+ G+ G ++ A N+ T M +P TY L+
Sbjct: 96 SKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDAGIEPGPDTYLALLNA 155
Query: 532 YCKINKLEAANGLLNEMLEK 551
Y + ++ L E +EK
Sbjct: 156 YAEKGDIDHVKQTL-EKVEK 174
Score = 37.4 bits (85), Expect = 0.11
Identities = 40/165 (24%), Positives = 80/165 (48%), Gaps = 13/165 (7%)
Query: 283 LIDGFCKDENVAAAKKAFEEMQKQGLKPNIVT--YNSLINGLCNNGKLEEAIDLWDKMVG 340
LI C +EN+ +KA E K + ++VT Y +LIN C + K+E+A++L ++
Sbjct: 564 LILVLCSEENM---QKALELKAKY--ESDMVTGGYAALINLCCRHDKVEDALNLKEEFDR 618
Query: 341 LGLKPNIVT--YNALINGFCKKKMMKEATKVFDDVSKQELV---PNVITFNTMIDAYCKE 395
L + T Y L+ K +++A K+ ++ +++++ ++F M++
Sbjct: 619 LDSSAVLDTGNYLGLVRVLAKHGKLQDAIKILKEMKEKDVLIKDTTALSFFHMLNGAALR 678
Query: 396 GMMEEGFSLCSSMLDEGIL-PNVSTYNCLIAGLCRKQDLQAAKEL 439
G +E L +++ G+ P+ + L+ K DL A E+
Sbjct: 679 GEIETVKQLHEAIVTLGLAEPSTNISFPLVTVHLEKGDLSTALEV 723
>PT09_YEAST (P32522) PET309 protein, mitochondrial precursor
Length = 965
Score = 42.7 bits (99), Expect = 0.003
Identities = 57/313 (18%), Positives = 126/313 (40%), Gaps = 5/313 (1%)
Query: 280 FNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMV 339
F+ L+ + N A ++ F + G P+I Y L+ + G+L+ L+ +++
Sbjct: 314 FDYLLVAHSRLHNWDALQQQFNALFGIGKLPSIQHYGILMYTMARIGELDSVNKLYTQLL 373
Query: 340 GLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVITFNTMIDAYCKEGMME 399
G+ P +L+ K F+ K ++ P+ T M+ Y ++
Sbjct: 374 RRGMIPTYAVLQSLLYAHYKVGDFAACFSHFELFKKYDITPSTATHTIMLKVYRGLNDLD 433
Query: 400 EGFSLCSSML-DEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEM-ENKGLKGDVVTYNI 457
F + + D + + LI C+ + A+EL N M E+ ++ + +
Sbjct: 434 GAFRILKRLSEDPSVEITEGHFALLIQMCCKTTNHLIAQELFNLMTEHYNIQHTGKSISA 493
Query: 458 LIDGLCKNDKSRNAEKLLNE-MFNLGLKPNHVT-YNTLMDGYCMEGKLKAALNVRTRMEK 515
L+D ++++ A L + NL + ++ YN + Y + ++
Sbjct: 494 LMDVYIESNRPTEAIALFEKHSKNLSWRDGLISVYNKAIKAYIGLRNANKCEELFDKITT 553
Query: 516 ERKQPNVVTYNVLIKGYCKINK-LEAANGLLNEMLEKGLNPNRTTYDIVRLEMLEKGFSP 574
+ N Y ++IK +N+ E A +++++++ + T+ + +E +K
Sbjct: 554 SKLAVNSEFYKMMIKFLVTLNEDCETALSIIDQLIKHSVIKVDATHFEIIMEAYDKEGYR 613
Query: 575 DIEGHLYNISSMS 587
D +LY S +
Sbjct: 614 DGIINLYKTMSQN 626
>CIGB_DICDI (Q94481) Protein cigB (Fragment)
Length = 735
Score = 42.7 bits (99), Expect = 0.003
Identities = 87/347 (25%), Positives = 147/347 (42%), Gaps = 65/347 (18%)
Query: 163 YGFKLSLTSCNPLLSALVKENKIGDVEYVYKEMIKRRIHTNLNTFNI-------FINGLC 215
YGF LT ++ VKE ++GD++ +E+I I + L T + G+
Sbjct: 372 YGFNQKLTK--GIIPEGVKELRLGDIK---QELIIDSIPSTLTTVILCDGFNQKLTKGII 426
Query: 216 RAG-KLNKAEDAIEDMKAWGISPNVVTYNTLVDGYCKRGSAG------KMYKAEAFMKEM 268
G K+ D +++ I PN VT L+DG+ ++ + G K +E+
Sbjct: 427 PEGVKVLYIGDIKQELIIDSI-PNTVTRVRLLDGFNQKLTKGIIPESVKELHIGNIKQEL 485
Query: 269 LANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLK--------PNIVTYNSLIN 320
+ + I PN VT TL DGF + + +E+ LK PN VT L +
Sbjct: 486 IIDSI-PNTVTTITLCDGFNQKLTKGIIPEGVKELYIGNLKQELIIDSIPNTVTTVRLCD 544
Query: 321 GLCNNGKLEEAI-----------DLWDKMVGLGLKPNIVTYNALINGF---CKKKMMKEA 366
G N KL + I D+ +++ + PN VT L +GF K ++ E
Sbjct: 545 GF--NQKLTKGIIPEGVKELYIRDIKQELI-IDSIPNTVTTVILCDGFNQKLTKGIIPEW 601
Query: 367 TK--VFDDVSKQEL----VPNVITFNTMIDAY---CKEGMMEEGFSLCSSMLDE------ 411
K D+ KQEL +PN +T ++D + +G++ E S+ +D+
Sbjct: 602 VKGLCLGDI-KQELIIDSIPNTVTTVRLLDGFNQKLTKGIIPE--SVKELYIDDIKQELI 658
Query: 412 -GILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNI 457
+PN T L+ G +K E + E+ +K +++ +I
Sbjct: 659 IDSIPNTVTTVRLLDGFNQKLTKGIIPEWVKELHIGDIKQELIIDSI 705
>YG3M_YEAST (P48237) Hypothetical 101.4 kDa protein in RPL24B-RSR1
intergenic region
Length = 864
Score = 41.6 bits (96), Expect = 0.006
Identities = 23/81 (28%), Positives = 38/81 (46%), Gaps = 2/81 (2%)
Query: 342 GLKPNIVTYNALINGFCKKKMMKEATKVFDDVS--KQELVPNVITFNTMIDAYCKEGMME 399
G+ PN +I + +K+M K+A FD + + P++ T+NTM+ KE
Sbjct: 315 GITPNKQNLTTVIQFYSRKEMTKQAWNTFDTMKFLSTKHFPDICTYNTMLRICEKERNFP 374
Query: 400 EGFSLCSSMLDEGILPNVSTY 420
+ L + D I P +TY
Sbjct: 375 KALDLFQEIQDHNIKPTTNTY 395
Score = 41.2 bits (95), Expect = 0.008
Identities = 24/83 (28%), Positives = 41/83 (48%), Gaps = 1/83 (1%)
Query: 233 WGISPNVVTYNTLVDGYCKRGSAGKMYKAEAFMKEMLANKICPNEVTFNTLIDGFCKDEN 292
+GI+PN T++ Y ++ + + MK L+ K P+ T+NT++ K+ N
Sbjct: 314 YGITPNKQNLTTVIQFYSRKEMTKQAWNTFDTMK-FLSTKHFPDICTYNTMLRICEKERN 372
Query: 293 VAAAKKAFEEMQKQGLKPNIVTY 315
A F+E+Q +KP TY
Sbjct: 373 FPKALDLFQEIQDHNIKPTTNTY 395
Score = 37.0 bits (84), Expect = 0.14
Identities = 22/80 (27%), Positives = 38/80 (47%), Gaps = 2/80 (2%)
Query: 273 ICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLK--PNIVTYNSLINGLCNNGKLEE 330
I PN+ T+I + + E A F+ M+ K P+I TYN+++ +
Sbjct: 316 ITPNKQNLTTVIQFYSRKEMTKQAWNTFDTMKFLSTKHFPDICTYNTMLRICEKERNFPK 375
Query: 331 AIDLWDKMVGLGLKPNIVTY 350
A+DL+ ++ +KP TY
Sbjct: 376 ALDLFQEIQDHNIKPTTNTY 395
Score = 34.7 bits (78), Expect = 0.71
Identities = 22/91 (24%), Positives = 44/91 (48%), Gaps = 5/91 (5%)
Query: 367 TKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSM--LDEGILPNVSTYNCLI 424
TK DD + PN T+I Y ++ M ++ ++ +M L P++ TYN ++
Sbjct: 308 TKFRDDYG---ITPNKQNLTTVIQFYSRKEMTKQAWNTFDTMKFLSTKHFPDICTYNTML 364
Query: 425 AGLCRKQDLQAAKELLNEMENKGLKGDVVTY 455
++++ A +L E+++ +K TY
Sbjct: 365 RICEKERNFPKALDLFQEIQDHNIKPTTNTY 395
Score = 32.3 bits (72), Expect = 3.5
Identities = 20/81 (24%), Positives = 34/81 (41%), Gaps = 2/81 (2%)
Query: 482 GLKPNHVTYNTLMDGYCMEGKLKAALNVRTRME--KERKQPNVVTYNVLIKGYCKINKLE 539
G+ PN T++ Y + K A N M+ + P++ TYN +++ K
Sbjct: 315 GITPNKQNLTTVIQFYSRKEMTKQAWNTFDTMKFLSTKHFPDICTYNTMLRICEKERNFP 374
Query: 540 AANGLLNEMLEKGLNPNRTTY 560
A L E+ + + P TY
Sbjct: 375 KALDLFQEIQDHNIKPTTNTY 395
Score = 32.0 bits (71), Expect = 4.6
Identities = 20/84 (23%), Positives = 38/84 (44%), Gaps = 2/84 (2%)
Query: 307 GLKPNIVTYNSLINGLCNNGKLEEAIDLWDKMVGLGLK--PNIVTYNALINGFCKKKMMK 364
G+ PN ++I ++A + +D M L K P+I TYN ++ K++
Sbjct: 315 GITPNKQNLTTVIQFYSRKEMTKQAWNTFDTMKFLSTKHFPDICTYNTMLRICEKERNFP 374
Query: 365 EATKVFDDVSKQELVPNVITFNTM 388
+A +F ++ + P T+ M
Sbjct: 375 KALDLFQEIQDHNIKPTTNTYIMM 398
>YEQ7_YEAST (P40050) Hypothetical 79.5 kDa protein in PTP3-SER3
intergenic region
Length = 688
Score = 32.7 bits (73), Expect = 2.7
Identities = 34/132 (25%), Positives = 60/132 (44%), Gaps = 17/132 (12%)
Query: 367 TKVFDDVSKQELVPNVITFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAG 426
T +F+ +SKQE ++ E +++ SLC D+ L + YN +A
Sbjct: 238 TILFNGLSKQE---------NLVSKKYGELVLKTIDSLC----DKNELTEIE-YNTALAA 283
Query: 427 LCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPN 486
L D +LLN+ + GLK D +TY ++I + + +LN++ N P+
Sbjct: 284 LINCTDETLVFKLLNK-KCPGLKKDSITYTLMIRSCTRIADEKRFMVVLNDLMN--KIPD 340
Query: 487 HVTYNTLMDGYC 498
+ + L+ YC
Sbjct: 341 YCVDSKLLFEYC 352
>YC81_MYCPN (P75496) Hypothetical lipoprotein MPN281 precursor
(A65_orf377)
Length = 377
Score = 32.7 bits (73), Expect = 2.7
Identities = 52/216 (24%), Positives = 82/216 (37%), Gaps = 32/216 (14%)
Query: 274 CPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIVTYN-------SLINGLCNNG 326
C F+ DG K + +K A +Q K NIV + I G + G
Sbjct: 23 CGARGKFDQFDDGKIKLASSLTSKAAANALQTIIEKYNIVKSGKDYPIEITQIAGGYDGG 82
Query: 327 K--LEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQELVPNVIT 384
+ L+ + + DK L I+ Y L++ + M +FD V+ +L PN +
Sbjct: 83 RTDLQTRVSVKDKTNFYNL---ILNYPDLVSVLARSGM----ELLFDKVNVDKLEPNFLK 135
Query: 385 FNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEME 444
FN I K G + GI ++ST ++ G L +AK+ E
Sbjct: 136 FNEQISGVAKSG-------------NYGIPVSLSTDILVLNGPVLHYILNSAKK---EEA 179
Query: 445 NKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFN 480
N +K + T I G D N + L ++ N
Sbjct: 180 NSQVKSQIKTSQIQAKGSLTIDTDENTKSLWEKIQN 215
>RT27_HUMAN (Q92552) Mitochondrial 28S ribosomal protein S27 (S27mt)
(MRP-S27)
Length = 414
Score = 32.0 bits (71), Expect = 4.6
Identities = 31/138 (22%), Positives = 58/138 (41%), Gaps = 10/138 (7%)
Query: 263 AFMKEMLANKICPNEVTFNTLIDGFCKDENVAAAKKAFEEMQKQGLKPNIV-----TYNS 317
+ M + K+ + +T + LID E + A+ + + PN T ++
Sbjct: 55 SLMDKTFERKLPVSSLTISRLIDNISSREEIDHAEYYLYKFRHS---PNCWYLRNWTIHT 111
Query: 318 LINGLCNNGKLEEAIDLWDKMVGLGLKPNIVTYNALINGFCKKKMMKEATKVFDDVSKQE 377
I ++A+ V G+ P+ T+N L++ F KK+ K+A V +V QE
Sbjct: 112 WIRQCLKYDAQDKALYTLVNKVQYGIFPDNFTFNLLMDSFIKKENYKDALSVVFEVMMQE 171
Query: 378 L--VPNVITFNTMIDAYC 393
VP+ + + +C
Sbjct: 172 AFEVPSTQLLSLYVLFHC 189
Score = 31.6 bits (70), Expect = 6.0
Identities = 34/135 (25%), Positives = 60/135 (44%), Gaps = 28/135 (20%)
Query: 428 CRKQDLQ--AAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNA-----EKLLNEMFN 480
C K D Q A L+N+++ G+ D T+N+L+D K + ++A E ++ E F
Sbjct: 116 CLKYDAQDKALYTLVNKVQY-GIFPDNFTFNLLMDSFIKKENYKDALSVVFEVMMQEAFE 174
Query: 481 LGLKPNHVTYNTLMDGYCMEGKLKAALNVRTRMEKER-----------KQPNVVTYNVLI 529
+ P+ + + +C+ K + E+ER KQ N V ++ +
Sbjct: 175 V---PSTQLLSLYVLFHCLAKKTDFS------WEEERNFGASLLLPGLKQKNSVGFSSQL 225
Query: 530 KGYCKINKLEAANGL 544
GY + K+E GL
Sbjct: 226 YGYALLGKVELQQGL 240
>LEP_BUCBP (Q89AM6) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 310
Score = 32.0 bits (71), Expect = 4.6
Identities = 22/87 (25%), Positives = 40/87 (45%), Gaps = 3/87 (3%)
Query: 447 GLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLG-LKPNHVTYNTLMDGYCMEG--KL 503
GL GD + YNIL L + N + N N +KPN T + ++ + L
Sbjct: 141 GLPGDKINYNILTKRLTITPNNINEQHTKNISINYKYIKPNDFTKHFKLNNIILNNVHSL 200
Query: 504 KAALNVRTRMEKERKQPNVVTYNVLIK 530
+++ N ++E +++ + YN+ K
Sbjct: 201 ESSNNNLLQLEMYQEKIEKIAYNIFFK 227
>ENV_SIVCZ (P17281) Envelope polyprotein GP160 precursor [Contains:
Exterior membrane glycoprotein (GP120); Transmembrane
glycoprotein (GP41)]
Length = 854
Score = 32.0 bits (71), Expect = 4.6
Identities = 31/119 (26%), Positives = 54/119 (45%), Gaps = 12/119 (10%)
Query: 372 DVSKQEL-VPNVI-TFNTMIDAYCKEGMMEEGFSLCSSMLDEGILPNVSTYNCLIAGLCR 429
D S QE+ +PNVI +FN K M+++ S+ D+ + P V + C
Sbjct: 77 DPSPQEVFLPNVIESFNMW-----KNNMVDQMHEDIISLWDQSLKPCVKLTPLCVTLQCS 131
Query: 430 KQDLQAAKELLNEMENKGLKGDVVTYNILIDGLCKNDKSRNAEKL--LNEMFNLGLKPN 486
K + AK L N+ + L+ ++N+ + DK + L + ++ NLG + N
Sbjct: 132 KANFSQAKNLTNQTSSPPLEMKNCSFNVTTE---LRDKKKQVYSLFYVEDVVNLGNENN 187
>DPO3_CLOTE (Q895K2) DNA polymerase III polC-type (EC 2.7.7.7)
(PolIII)
Length = 1427
Score = 31.6 bits (70), Expect = 6.0
Identities = 19/81 (23%), Positives = 36/81 (43%), Gaps = 10/81 (12%)
Query: 179 LVKENKIGDVEYVYKEMIKRRIHTNLNTFNIFINGLCRAGKLNKAEDAIEDMKAWGISPN 238
+VK NKI ++ ++ ++ +HT ++ + +N AE I+ WG
Sbjct: 311 IVKLNKIEKMDIAEEKRVELHLHTKMSAMD----------GMNSAESLIKRAAKWGHKAV 360
Query: 239 VVTYNTLVDGYCKRGSAGKMY 259
+T + +V Y + A K Y
Sbjct: 361 AITDHGVVQAYPEAMEAAKKY 381
>CCD2_HUMAN (Q96LB3) Coiled-coil domain containing protein 2
(Capillary morphogenesis protein-1) (CMG-1)
Length = 600
Score = 31.2 bits (69), Expect = 7.9
Identities = 24/108 (22%), Positives = 53/108 (48%), Gaps = 3/108 (2%)
Query: 443 MENKGLKGDVVTYNILIDGLCKNDKSRNAEKLLNEMFNLGLKPNHVTYNTLMDGYCMEGK 502
+E K L+G + YN+L+D L N + E+++N+ L + + T + + + K
Sbjct: 141 VEIKELQGQLADYNMLVDKLNTNTE---MEEVMNDYNMLKAQNDRETQSLDVIFTERQAK 197
Query: 503 LKAALNVRTRMEKERKQPNVVTYNVLIKGYCKINKLEAANGLLNEMLE 550
K +V +E+E++ + + N+ + K +++ N L + L+
Sbjct: 198 EKQIRSVEEEIEQEKQATDDIIKNMSFENQVKYLEMKTTNEKLLQELD 245
>ASC2_MOUSE (O35885) Achaete-scute homolog 2 (Mash-2)
Length = 263
Score = 31.2 bits (69), Expect = 7.9
Identities = 21/73 (28%), Positives = 35/73 (47%), Gaps = 1/73 (1%)
Query: 404 LCSSMLDEGILPNVSTYNCLIAGLCRKQDLQAAKELLNEMENKGLKGDVVTYNILIDGLC 463
L +S++D G LP + + +AG C + QA+ ELL + G +
Sbjct: 64 LNASLMDGGALPRLMPTSSGVAGACAARRRQASPELL-RCSRRRRSGATEASSSSAAVAR 122
Query: 464 KNDKSRNAEKLLN 476
+N++ RN KL+N
Sbjct: 123 RNERERNRVKLVN 135
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,670,454
Number of Sequences: 164201
Number of extensions: 2872000
Number of successful extensions: 7470
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 7403
Number of HSP's gapped (non-prelim): 46
length of query: 587
length of database: 59,974,054
effective HSP length: 116
effective length of query: 471
effective length of database: 40,926,738
effective search space: 19276493598
effective search space used: 19276493598
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC122726.13


