

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122724.4 - phase: 0
(357 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
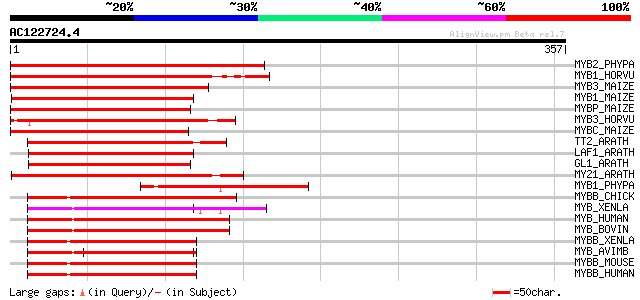
Score E
Sequences producing significant alignments: (bits) Value
MYB2_PHYPA (P80073) Myb-related protein Pp2 202 8e-52
MYB1_HORVU (P20026) Myb-related protein Hv1 199 1e-50
MYB3_MAIZE (P20025) Myb-related protein Zm38 194 3e-49
MYB1_MAIZE (P20024) Myb-related protein Zm1 185 2e-46
MYBP_MAIZE (P27898) Myb-related protein P 184 2e-46
MYB3_HORVU (P20027) Myb-related protein Hv33 172 2e-42
MYBC_MAIZE (P10290) Anthocyanin regulatory C1 protein 164 2e-40
TT2_ARATH (Q9FJA2) TRANSPARENT TESTA 2 protein (Myb-related prot... 149 1e-35
LAF1_ARATH (Q9M0K4) Transcription factor LAF1 (Long after far-re... 144 3e-34
GL1_ARATH (P27900) Trichome differentiation protein GL1 (GLABROU... 144 4e-34
MY21_ARATH (Q9LK95) Transcription factor MYB21 (Myb-related prot... 139 1e-32
MYB1_PHYPA (P80074) Myb-related protein Pp1 (Fragment) 108 2e-23
MYBB_CHICK (Q03237) Myb-related protein B (B-Myb) 107 4e-23
MYB_XENLA (Q08759) Myb protein 106 1e-22
MYB_HUMAN (P10242) Myb proto-oncogene protein (C-myb) 105 1e-22
MYB_BOVIN (P46200) Myb proto-oncogene protein (C-myb) 105 1e-22
MYBB_XENLA (P52551) Myb-related protein B (B-Myb) (Myb-related p... 105 2e-22
MYB_AVIMB (P01104) Transforming protein Myb 104 3e-22
MYBB_MOUSE (P48972) Myb-related protein B (B-Myb) 104 3e-22
MYBB_HUMAN (P10244) Myb-related protein B (B-Myb) 104 3e-22
>MYB2_PHYPA (P80073) Myb-related protein Pp2
Length = 421
Score = 202 bits (515), Expect = 8e-52
Identities = 93/164 (56%), Positives = 116/164 (70%)
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR PCC+KVGL++GPWT EEDQKL+++I +G WRA+P AGL RCGKSCRLRWTNY
Sbjct: 1 MGRKPCCEKVGLRRGPWTSEEDQKLVSHITNNGLSCWRAIPKLAGLLRCGKSCRLRWTNY 60
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG FS EE I+ LHA LGNRWS IA L RTDNEIKNYWNT LKKRL G
Sbjct: 61 LRPDLKRGIFSEAEENLILDLHATLGNRWSRIAAQLPGRTDNEIKNYWNTRLKKRLRSQG 120
Query: 121 IDPVTHKPKNDALLSSSDNAHSKTASNLSHMAQWESARLEAEAR 164
+DP TH P D+ L +++ + S + ++++ E ++
Sbjct: 121 LDPNTHLPLEDSKLDDTEDDTDDEGGDSSDVTMSDASKSEKRSK 164
>MYB1_HORVU (P20026) Myb-related protein Hv1
Length = 267
Score = 199 bits (505), Expect = 1e-50
Identities = 97/167 (58%), Positives = 120/167 (71%), Gaps = 12/167 (7%)
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGRSPCC+K KG WT EED +L AYI HG G WR+LP AGL RCGKSCRLRW NY
Sbjct: 1 MGRSPCCEKAHTNKGAWTKEEDDRLTAYIKAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 60
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG FS +E++ II+LH+LLGN+WS IA L RTDNEIKNYWNTH++++LT G
Sbjct: 61 LRPDLKRGNFSHEEDELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRKLTSRG 120
Query: 121 IDPVTHKPKNDALLSSSDNAHSKTASNLSHMAQWESARLEAEARLVR 167
IDPVTH+ N SD+A ASN++ +ESA+ + + + R
Sbjct: 121 IDPVTHRAIN------SDHA----ASNIT--ISFESAQRDDKGAVFR 155
>MYB3_MAIZE (P20025) Myb-related protein Zm38
Length = 255
Score = 194 bits (493), Expect = 3e-49
Identities = 83/128 (64%), Positives = 101/128 (78%)
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGRSPCC+K +G WT EED++L+AYI HG G WR+LP AGL RCGKSCRLRW NY
Sbjct: 1 MGRSPCCEKAHTNRGAWTKEEDERLVAYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 60
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG F+ E+ I++LH+LLGN+WS IA L RTDNEIKNYWNTH++++L G
Sbjct: 61 LRPDLKRGNFTADEDDLIVKLHSLLGNKWSLIAARLPGRTDNEIKNYWNTHVRRKLLGRG 120
Query: 121 IDPVTHKP 128
IDPVTH+P
Sbjct: 121 IDPVTHRP 128
>MYB1_MAIZE (P20024) Myb-related protein Zm1
Length = 340
Score = 185 bits (469), Expect = 2e-46
Identities = 80/117 (68%), Positives = 97/117 (82%)
Query: 2 GRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYL 61
GR+PCC KVGL +G WTP+ED +L+AYI +HGH +WRALP +AGL RCGKSCRLRW NYL
Sbjct: 4 GRAPCCAKVGLNRGSWTPQEDMRLIAYIQKHGHTNWRALPKQAGLLRCGKSCRLRWINYL 63
Query: 62 RPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTK 118
RPD+KRG F+ +EE+ II+LH LLGN+WS IA L RTDNEIKN WNTHLKK++ +
Sbjct: 64 RPDLKRGNFTDEEEEAIIRLHGLLGNKWSKIAACLPGRTDNEIKNVWNTHLKKKVAQ 120
>MYBP_MAIZE (P27898) Myb-related protein P
Length = 399
Score = 184 bits (468), Expect = 2e-46
Identities = 82/116 (70%), Positives = 94/116 (80%)
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR+PCC+KVGLK+G WT EEDQ L YI EHG GSWR+LP AGL RCGKSCRLRW NY
Sbjct: 1 MGRTPCCEKVGLKRGRWTAEEDQLLANYIAEHGEGSWRSLPKNAGLLRCGKSCRLRWINY 60
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRL 116
LR D+KRG S +EE II+LHA LGNRWS IA+HL RTDNEIKNYWN+HL +++
Sbjct: 61 LRADVKRGNISKEEEDIIIKLHATLGNRWSLIASHLPGRTDNEIKNYWNSHLSRQI 116
>MYB3_HORVU (P20027) Myb-related protein Hv33
Length = 302
Score = 172 bits (435), Expect = 2e-42
Identities = 85/148 (57%), Positives = 105/148 (70%), Gaps = 9/148 (6%)
Query: 1 MGRSPCCDKVG---LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRW 57
MGR P VG ++KG W+PEED+KL +I HG G W ++P A L RCGKSCRLRW
Sbjct: 1 MGR-PSSGAVGQPKVRKGLWSPEEDEKLYNHIIRHGVGCWSSVPRLAALNRCGKSCRLRW 59
Query: 58 TNYLRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLT 117
NYLRPD+KRG FS QEE I+ LH +LGNRWS IA+HL RTDNEIKN+WN+ +KK+L
Sbjct: 60 INYLRPDLKRGCFSQQEEDHIVALHQILGNRWSQIASHLPGRTDNEIKNFWNSCIKKKLR 119
Query: 118 KMGIDPVTHKPKNDALLSSSDNAHSKTA 145
+ GIDP THKP ++S+D A + A
Sbjct: 120 QQGIDPATHKP-----MASADTATAAAA 142
>MYBC_MAIZE (P10290) Anthocyanin regulatory C1 protein
Length = 273
Score = 164 bits (416), Expect = 2e-40
Identities = 74/115 (64%), Positives = 85/115 (73%)
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR CC K G+K+G WT +ED L AY+ HG G WR +P KAGL+RCGKSCRLRW NY
Sbjct: 1 MGRRACCAKEGVKRGAWTSKEDDALAAYVKAHGEGKWREVPQKAGLRRCGKSCRLRWLNY 60
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKR 115
LRP+I+RG S EE II+LH LLGNRWS IA L RTDNEIKNYWN+ L +R
Sbjct: 61 LRPNIRRGNISYDEEDLIIRLHRLLGNRWSLIAGRLPGRTDNEIKNYWNSTLGRR 115
>TT2_ARATH (Q9FJA2) TRANSPARENT TESTA 2 protein (Myb-related protein
123) (AtMYB123) (Myb-related transcription factor
LBM2-like)
Length = 258
Score = 149 bits (375), Expect = 1e-35
Identities = 73/128 (57%), Positives = 88/128 (68%), Gaps = 4/128 (3%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L +G WT ED+ L YI HG G W LP +AGL+RCGKSCRLRW NYLRP IKRG S
Sbjct: 14 LNRGAWTDHEDKILRDYITTHGEGKWSTLPNQAGLKRCGKSCRLRWKNYLRPGIKRGNIS 73
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKND 131
EE+ II+LH LLGNRWS IA L RTDNEIKN+WN++L+KRL K T +PK
Sbjct: 74 SDEEELIIRLHNLLGNRWSLIAGRLPGRTDNEIKNHWNSNLRKRLPK----TQTKQPKRI 129
Query: 132 ALLSSSDN 139
++++N
Sbjct: 130 KHSTNNEN 137
>LAF1_ARATH (Q9M0K4) Transcription factor LAF1 (Long after far-red
light protein 1) (Myb-related protein 18) (AtMYB18)
Length = 283
Score = 144 bits (363), Expect = 3e-34
Identities = 64/106 (60%), Positives = 80/106 (75%)
Query: 13 KKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFSL 72
+KG W+PEED+KL ++I +GH W +P KAGLQR GKSCRLRW NYLRP +KR S
Sbjct: 11 RKGLWSPEEDEKLRSFILSYGHSCWTTVPIKAGLQRNGKSCRLRWINYLRPGLKRDMISA 70
Query: 73 QEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTK 118
+EE+TI+ H+ LGN+WS IA L RTDNEIKNYW++HLKK+ K
Sbjct: 71 EEEETILTFHSSLGNKWSQIAKFLPGRTDNEIKNYWHSHLKKKWLK 116
>GL1_ARATH (P27900) Trichome differentiation protein GL1 (GLABROUS1
protein)
Length = 228
Score = 144 bits (362), Expect = 4e-34
Identities = 63/104 (60%), Positives = 76/104 (72%)
Query: 13 KKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFSL 72
KKG WT EED L+ Y+ HG G W + K GL+RCGKSCRLRW NYL P++ +G F+
Sbjct: 15 KKGLWTVEEDNILMDYVLNHGTGQWNRIVRKTGLKRCGKSCRLRWMNYLSPNVNKGNFTE 74
Query: 73 QEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRL 116
QEE II+LH LLGNRWS IA + RTDN++KNYWNTHL K+L
Sbjct: 75 QEEDLIIRLHKLLGNRWSLIAKRVPGRTDNQVKNYWNTHLSKKL 118
>MY21_ARATH (Q9LK95) Transcription factor MYB21 (Myb-related protein
21) (AtMYB21) (Myb homolog 3) (AtMyb3)
Length = 226
Score = 139 bits (350), Expect = 1e-32
Identities = 68/149 (45%), Positives = 91/149 (60%), Gaps = 5/149 (3%)
Query: 2 GRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYL 61
G S + ++KGPWT EED L+ YI HG G W +L AGL+R GKSCRLRW NYL
Sbjct: 10 GGSGSSAEAEVRKGPWTMEEDLILINYIANHGDGVWNSLAKSAGLKRTGKSCRLRWLNYL 69
Query: 62 RPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGI 121
RPD++RG + +E+ I++LHA GNRWS IA HL RTDNEIKN+W T ++K + + +
Sbjct: 70 RPDVRRGNITPEEQLIIMELHAKWGNRWSKIAKHLPGRTDNEIKNFWRTRIQKYIKQSDV 129
Query: 122 DPVTHKPKNDALLSSSDNAHSKTASNLSH 150
+ + S + + AS SH
Sbjct: 130 TTTSSVGSH-----HSSEINDQAASTSSH 153
>MYB1_PHYPA (P80074) Myb-related protein Pp1 (Fragment)
Length = 449
Score = 108 bits (270), Expect = 2e-23
Identities = 61/111 (54%), Positives = 73/111 (64%), Gaps = 5/111 (4%)
Query: 85 LGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKNDALL---SSSDNAH 141
LGNRWSAIA + +RTDNEIKNYWNTHLKKRL +MGIDPVTHK L S
Sbjct: 1 LGNRWSAIA--IPRRTDNEIKNYWNTHLKKRLMQMGIDPVTHKSTAAEELVHYSIIPGLR 58
Query: 142 SKTASNLSHMAQWESARLEAEARLVRESKIRSHNSLLHHNHLKAWNINLES 192
++NL+HM+QW+SAR EAEARL R+S + S S L L+ NL +
Sbjct: 59 PVVSTNLTHMSQWDSARAEAEARLSRQSSLTSPASDLAQTSLEHQKSNLST 109
>MYBB_CHICK (Q03237) Myb-related protein B (B-Myb)
Length = 686
Score = 107 bits (268), Expect = 4e-23
Identities = 53/135 (39%), Positives = 83/135 (61%), Gaps = 1/135 (0%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L KGPWT EEDQK++ + ++G W L AK R GK CR RW N+L P++K+ ++
Sbjct: 81 LVKGPWTKEEDQKVIELVKKYGTKQW-TLIAKHLKGRLGKQCRERWHNHLNPEVKKSSWT 139
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKND 131
+E++ I + H +LGNRW+ IA L RTDN +KN+WN+ +K+++ G T + +
Sbjct: 140 EEEDRIIFEAHKVLGNRWAEIAKLLPGRTDNAVKNHWNSTIKRKVDTGGFLNETKESQPL 199
Query: 132 ALLSSSDNAHSKTAS 146
LL D+ S++ +
Sbjct: 200 YLLVEVDDNESQSGT 214
>MYB_XENLA (Q08759) Myb protein
Length = 624
Score = 106 bits (264), Expect = 1e-22
Identities = 59/171 (34%), Positives = 100/171 (57%), Gaps = 18/171 (10%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L KGPWT EEDQ+++ + ++G W + AK R GK CR RW N+L P++K+ ++
Sbjct: 87 LIKGPWTKEEDQRVIELVHKYGPKRWSVI-AKHLKGRIGKQCRERWHNHLNPEVKKSSWT 145
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGI---------- 121
+E++TI + H LGNRW+ IA L RTDN IKN+WN+ ++++ + G
Sbjct: 146 EEEDRTIYEAHKRLGNRWAEIAKLLPGRTDNAIKNHWNSTMRRKEEQEGYLQNSSKTNQH 205
Query: 122 DPVTHKPKNDALL------SSSDNAHSKTAS-NLSHMAQWESARLEAEARL 165
VT+ PK++ L+ +S++++ + T+S H+A+ ++A R+
Sbjct: 206 TIVTNFPKSNHLMTFTHTRASAEHSQASTSSFPYYHIAEHQNASYPVALRV 256
Score = 70.5 bits (171), Expect = 6e-12
Identities = 36/108 (33%), Positives = 57/108 (52%), Gaps = 2/108 (1%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L K WT EED+KL ++++G W+ + A R C+ RW L P++ +G ++
Sbjct: 35 LGKTRWTREEDEKLKKLVEQNGTEEWKVI-ASFLPNRTDVQCQHRWQKVLNPELIKGPWT 93
Query: 72 LQEEQTIIQL-HALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTK 118
+E+Q +I+L H RWS IA HL R + + W+ HL + K
Sbjct: 94 KEEDQRVIELVHKYGPKRWSVIAKHLKGRIGKQCRERWHNHLNPEVKK 141
>MYB_HUMAN (P10242) Myb proto-oncogene protein (C-myb)
Length = 640
Score = 105 bits (263), Expect = 1e-22
Identities = 52/131 (39%), Positives = 81/131 (61%), Gaps = 2/131 (1%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L KGPWT EEDQ+++ + ++G W + AK R GK CR RW N+L P++K+ ++
Sbjct: 90 LIKGPWTKEEDQRVIELVQKYGPKRWSVI-AKHLKGRIGKQCRERWHNHLNPEVKKTSWT 148
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKND 131
+E++ I Q H LGNRW+ IA L RTDN IKN+WN+ +++++ + G + K
Sbjct: 149 EEEDRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHWNSTMRRKVEQEGYLQESSKASQP 208
Query: 132 ALLSS-SDNAH 141
A+ +S N+H
Sbjct: 209 AVATSFQKNSH 219
>MYB_BOVIN (P46200) Myb proto-oncogene protein (C-myb)
Length = 640
Score = 105 bits (263), Expect = 1e-22
Identities = 52/131 (39%), Positives = 81/131 (61%), Gaps = 2/131 (1%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L KGPWT EEDQ+++ + ++G W + AK R GK CR RW N+L P++K+ ++
Sbjct: 90 LIKGPWTKEEDQRVIELVQKYGPKRWSVI-AKHLKGRIGKQCRERWHNHLNPEVKKTSWT 148
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKND 131
+E++ I Q H LGNRW+ IA L RTDN IKN+WN+ +++++ + G + K
Sbjct: 149 EEEDRIIYQAHKRLGNRWAEIAKLLPGRTDNAIKNHWNSTMRRKVEQEGYLQESSKASQP 208
Query: 132 ALLSS-SDNAH 141
A+ +S N+H
Sbjct: 209 AVTTSFQKNSH 219
>MYBB_XENLA (P52551) Myb-related protein B (B-Myb) (Myb-related
protein 1) (XMYB1)
Length = 743
Score = 105 bits (262), Expect = 2e-22
Identities = 48/109 (44%), Positives = 72/109 (66%), Gaps = 1/109 (0%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L KGPWT EED+K++ + ++G W L AK R GK CR RW N+L P++K+ ++
Sbjct: 81 LVKGPWTKEEDEKVIELVKKYGTKHW-TLIAKQLRGRMGKQCRERWHNHLNPEVKKSSWT 139
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
+E++ I Q H +LGNRW+ IA L RTDN +KN+WN+ +K+++ G
Sbjct: 140 EEEDRIICQAHKVLGNRWAEIAKLLPGRTDNAVKNHWNSTIKRKVETGG 188
>MYB_AVIMB (P01104) Transforming protein Myb
Length = 382
Score = 104 bits (260), Expect = 3e-22
Identities = 46/109 (42%), Positives = 73/109 (66%), Gaps = 1/109 (0%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L KGPWT EEDQ+++ ++ ++G W + AK R GK CR RW N+L P++K+ ++
Sbjct: 19 LNKGPWTKEEDQRVIEHVQKYGPKRWSDI-AKHLKGRIGKQCRERWHNHLNPEVKKTSWT 77
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
+E++ I Q H LGNRW+ IA L RTDN +KN+WN+ +++++ + G
Sbjct: 78 EEEDRIIYQAHKRLGNRWAEIAKLLPGRTDNAVKNHWNSTMRRKVEQEG 126
Score = 49.3 bits (116), Expect = 1e-05
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 1/72 (1%)
Query: 48 RCGKSCRLRWTNYLRPDIKRGKFSLQEEQTIIQLHALLG-NRWSAIATHLAKRTDNEIKN 106
R C+ RW L P++ +G ++ +E+Q +I+ G RWS IA HL R + +
Sbjct: 2 RTDVQCQHRWQKVLNPELNKGPWTKEEDQRVIEHVQKYGPKRWSDIAKHLKGRIGKQCRE 61
Query: 107 YWNTHLKKRLTK 118
W+ HL + K
Sbjct: 62 RWHNHLNPEVKK 73
>MYBB_MOUSE (P48972) Myb-related protein B (B-Myb)
Length = 704
Score = 104 bits (260), Expect = 3e-22
Identities = 48/109 (44%), Positives = 72/109 (66%), Gaps = 1/109 (0%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L KGPWT EEDQK++ + ++G W L AK R GK CR RW N+L P++K+ ++
Sbjct: 81 LVKGPWTKEEDQKVIELVKKYGTKQW-TLIAKHLKGRLGKQCRERWHNHLNPEVKKSCWT 139
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
+E++ I + H +LGNRW+ IA L RTDN +KN+WN+ +K+++ G
Sbjct: 140 EEEDRIICEAHKVLGNRWAEIAKMLPGRTDNAVKNHWNSTIKRKVDTGG 188
>MYBB_HUMAN (P10244) Myb-related protein B (B-Myb)
Length = 700
Score = 104 bits (260), Expect = 3e-22
Identities = 48/109 (44%), Positives = 72/109 (66%), Gaps = 1/109 (0%)
Query: 12 LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFS 71
L KGPWT EEDQK++ + ++G W L AK R GK CR RW N+L P++K+ ++
Sbjct: 81 LVKGPWTKEEDQKVIELVKKYGTKQW-TLIAKHLKGRLGKQCRERWHNHLNPEVKKSCWT 139
Query: 72 LQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
+E++ I + H +LGNRW+ IA L RTDN +KN+WN+ +K+++ G
Sbjct: 140 EEEDRIICEAHKVLGNRWAEIAKMLPGRTDNAVKNHWNSTIKRKVDTGG 188
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.309 0.128 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,955,540
Number of Sequences: 164201
Number of extensions: 2068865
Number of successful extensions: 11587
Number of sequences better than 10.0: 220
Number of HSP's better than 10.0 without gapping: 96
Number of HSP's successfully gapped in prelim test: 128
Number of HSP's that attempted gapping in prelim test: 8818
Number of HSP's gapped (non-prelim): 908
length of query: 357
length of database: 59,974,054
effective HSP length: 111
effective length of query: 246
effective length of database: 41,747,743
effective search space: 10269944778
effective search space used: 10269944778
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC122724.4


