

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
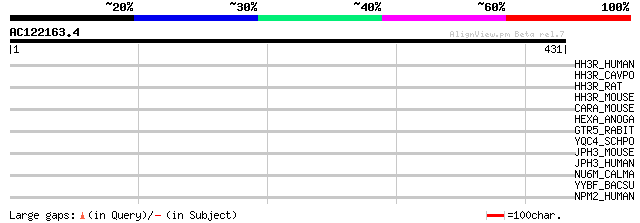
Query= AC122163.4 - phase: 0
(431 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
HH3R_HUMAN (Q9Y5N1) Histamine H3 receptor (HH3R) (G protein-coup... 37 0.13
HH3R_CAVPO (Q9JI35) Histamine H3 receptor (HH3R) 37 0.13
HH3R_RAT (Q9QYN8) Histamine H3 receptor (HH3R) 35 0.29
HH3R_MOUSE (P58406) Histamine H3 receptor (HH3R) 35 0.29
CARA_MOUSE (P58660) Caspase recruitment domain protein 10 (Bcl10... 33 1.1
HEXA_ANOGA (Q17020) Hexamerin 1.1 precursor (HEX-1.1) 32 3.2
GTR5_RABIT (P46408) Solute carrier family 2, facilitated glucose... 32 3.2
YQC4_SCHPO (O74466) Hypothetical protein C1739.04c in chromosome... 31 5.4
JPH3_MOUSE (Q9ET77) Junctophilin 3 (Junctophilin type 3) (JP-3) 31 5.4
JPH3_HUMAN (Q8WXH2) Junctophilin 3 (Junctophilin type 3) (JP-3) 31 5.4
NU6M_CALMA (P43196) NADH-ubiquinone oxidoreductase chain 6 (EC 1... 31 7.0
YYBF_BACSU (P37498) Hypothetical transport protein yybF 30 9.2
NPM2_HUMAN (Q86SE8) Nucleoplasmin 2 30 9.2
>HH3R_HUMAN (Q9Y5N1) Histamine H3 receptor (HH3R) (G protein-coupled
receptor 97)
Length = 445
Score = 36.6 bits (83), Expect = 0.13
Identities = 45/176 (25%), Positives = 71/176 (39%), Gaps = 19/176 (10%)
Query: 40 QSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFATLSFI 99
Q+N ++ S F A+ +P+ ++L T + + V V L + + +
Sbjct: 68 QNNFFLLNLAISDFLVGAFCIPLYVPYVLTGRWTFGRGLCKLWLV-VDYLLCTSSAFNIV 126
Query: 100 SLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVYKIWW 159
+S +DR FL S + Q+G + R K++L+W L + W
Sbjct: 127 LIS-YDR--------FLSVTRAVSYRAQQGDTRRAVR--KMLLVWVLAFLLYGPAILSWE 175
Query: 160 YISGASQIP---YYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQR 212
Y+SG S IP Y E + + L T +F FL F L YL IQR
Sbjct: 176 YLSGGSSIPEGHCYAEFFYNWYFLITASTLEFFTP----FLSVTFFNLSIYLNIQR 227
>HH3R_CAVPO (Q9JI35) Histamine H3 receptor (HH3R)
Length = 445
Score = 36.6 bits (83), Expect = 0.13
Identities = 31/101 (30%), Positives = 42/101 (40%), Gaps = 9/101 (8%)
Query: 115 FLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVYKIWWYISGASQIP---YYG 171
FL S + Q+G + R K++L+W L + W Y+SG S IP Y
Sbjct: 134 FLSVTRAVSYRAQQGDTRRAVR--KMVLVWVLAFLLYGPAILSWEYLSGGSSIPEGHCYA 191
Query: 172 EVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQR 212
E + + L T +F FL F L YL IQR
Sbjct: 192 EFFYNWYFLITASTLEFFTP----FLSVTFFNLSIYLNIQR 228
>HH3R_RAT (Q9QYN8) Histamine H3 receptor (HH3R)
Length = 445
Score = 35.4 bits (80), Expect = 0.29
Identities = 45/176 (25%), Positives = 70/176 (39%), Gaps = 19/176 (10%)
Query: 40 QSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFATLSFI 99
Q+N ++ S F A+ +P+ ++L T + + V V L + + +
Sbjct: 68 QNNFFLLNLAISDFLVGAFCIPLYVPYVLTGRWTFGRGLCKLWLV-VDYLLCASSVFNIV 126
Query: 100 SLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVYKIWW 159
+S +DR FL S + Q+G + R K+ L+W L + W
Sbjct: 127 LIS-YDR--------FLSVTRAVSYRAQQGDTRRAVR--KMALVWVLAFLLYGPAILSWE 175
Query: 160 YISGASQIP---YYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQR 212
Y+SG S IP Y E + + L T +F FL F L YL IQR
Sbjct: 176 YLSGGSSIPEGHCYAEFFYNWYFLITASTLEFFTP----FLSVTFFNLSIYLNIQR 227
>HH3R_MOUSE (P58406) Histamine H3 receptor (HH3R)
Length = 445
Score = 35.4 bits (80), Expect = 0.29
Identities = 45/176 (25%), Positives = 70/176 (39%), Gaps = 19/176 (10%)
Query: 40 QSNLCKAGISWSVFFTLAYVVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFATLSFI 99
Q+N ++ S F A+ +P+ ++L T + + V V L + + +
Sbjct: 68 QNNFFLLNLAISDFLVGAFCIPLYVPYVLTGRWTFGRGLCKLWLV-VDYLLCASSVFNIV 126
Query: 100 SLSKWDRKYGFSKFLFLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVYKIWW 159
+S +DR FL S + Q+G + R K+ L+W L + W
Sbjct: 127 LIS-YDR--------FLSVTRAVSYRAQQGDTRRAVR--KMALVWVLAFLLYGPAILSWE 175
Query: 160 YISGASQIP---YYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQR 212
Y+SG S IP Y E + + L T +F FL F L YL IQR
Sbjct: 176 YLSGGSSIPEGHCYAEFFYNWYFLITASTLEFFTP----FLSVTFFNLSIYLNIQR 227
>CARA_MOUSE (P58660) Caspase recruitment domain protein 10
(Bcl10-interacting MAGUK protein 1) (Bimp1)
Length = 1021
Score = 33.5 bits (75), Expect = 1.1
Identities = 24/100 (24%), Positives = 44/100 (44%), Gaps = 1/100 (1%)
Query: 286 LVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSFDNIDGETPTALIASAQA 345
L ++T + G +L + + T L AS +C+ ++S ++ E P+ L
Sbjct: 428 LTTVTSLEGTKAMLEAQLQRTQGGSCLKACASSHSLCSNLSSTWSLS-EFPSPLGGPEAT 486
Query: 346 MAADISWGSSEDESGDEEDELNNTKMMPIQTRTISFHKRQ 385
A S + +E+ D E E+N ++P S +RQ
Sbjct: 487 GEAGGSEPHTSEEATDSEKEINRLSILPFPPSAGSILRRQ 526
>HEXA_ANOGA (Q17020) Hexamerin 1.1 precursor (HEX-1.1)
Length = 692
Score = 32.0 bits (71), Expect = 3.2
Identities = 16/40 (40%), Positives = 26/40 (65%)
Query: 86 VQISLSVFATLSFISLSKWDRKYGFSKFLFLDKVSDESLK 125
V+ISL+V A+ S++ +K++ KY +FLF K E L+
Sbjct: 8 VEISLAVLASGSYVPSTKFEAKYADKEFLFKQKFFFEVLR 47
>GTR5_RABIT (P46408) Solute carrier family 2, facilitated glucose
transporter, member 5 (Glucose transporter type 5, small
intestine) (Fructose transporter)
Length = 486
Score = 32.0 bits (71), Expect = 3.2
Identities = 19/80 (23%), Positives = 38/80 (46%)
Query: 140 LILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVC 199
L + GL + V +W +SG + I YY ++Y+S + T + ++ VC
Sbjct: 265 LCAMRGLAWQLISVVPLMWQQLSGVNAIYYYDQIYLSPLDTDTQYYTAATGAVNVLMTVC 324
Query: 200 VLFRLIGYLQIQRLDEFAPV 219
+F + + ++ L F+P+
Sbjct: 325 TVFVVESWARLLLLLGFSPL 344
>YQC4_SCHPO (O74466) Hypothetical protein C1739.04c in chromosome
III
Length = 288
Score = 31.2 bits (69), Expect = 5.4
Identities = 25/87 (28%), Positives = 39/87 (44%), Gaps = 7/87 (8%)
Query: 315 LASKWHICATINSFDNIDGETPTALIASAQAMAADISWGSSEDESGDEEDE------LNN 368
L SK C I +F N+ +PT L + + + S+ D S EDE + +
Sbjct: 68 LLSKQDACHLIEAFVNVSITSPTLLTSVDPVLYQQLEASSTNDISTVFEDESSSLPIILH 127
Query: 369 TKMMPIQTRTISFHKRQALVTYMENNR 395
K +Q RT++ K A V+ E N+
Sbjct: 128 PKFSSMQVRTVTSPK-DAFVSAFEENK 153
>JPH3_MOUSE (Q9ET77) Junctophilin 3 (Junctophilin type 3) (JP-3)
Length = 744
Score = 31.2 bits (69), Expect = 5.4
Identities = 29/101 (28%), Positives = 53/101 (51%), Gaps = 14/101 (13%)
Query: 269 VIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSF 328
V+ H+D +ILK + GL +L+SGL + K++S + LAS+ + +SF
Sbjct: 197 VLVAHSDSEILKSKKKGLFRRSLLSGLKL---------RKSESKSSLASQ---RSKQSSF 244
Query: 329 DNIDGETPTALIASAQAMAADISWGSSEDESGDEEDELNNT 369
+ G + + ++A + + IS G +E E ED+++ T
Sbjct: 245 RSEAGMSTVS--STASDIHSTISLGEAEAELAVIEDDIDAT 283
>JPH3_HUMAN (Q8WXH2) Junctophilin 3 (Junctophilin type 3) (JP-3)
Length = 748
Score = 31.2 bits (69), Expect = 5.4
Identities = 29/101 (28%), Positives = 53/101 (51%), Gaps = 14/101 (13%)
Query: 269 VIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATINSF 328
V+ H+D +ILK + GL +L+SGL + K++S + LAS+ + +SF
Sbjct: 197 VLVAHSDSEILKSKKKGLFRRSLLSGLKL---------RKSESKSSLASQ---RSKQSSF 244
Query: 329 DNIDGETPTALIASAQAMAADISWGSSEDESGDEEDELNNT 369
+ G + + ++A + + IS G +E E ED+++ T
Sbjct: 245 RSEAGMSTVS--STASDIHSTISLGEAEAELAVIEDDIDAT 283
>NU6M_CALMA (P43196) NADH-ubiquinone oxidoreductase chain 6 (EC
1.6.5.3)
Length = 173
Score = 30.8 bits (68), Expect = 7.0
Identities = 20/63 (31%), Positives = 31/63 (48%), Gaps = 10/63 (15%)
Query: 140 LILLWGLPCFICQCVYKIWWYISGASQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVC 199
L+LL GL CF+ + ++ PYYG V G++L ++ C W + F+
Sbjct: 4 LVLLLGL-CFV------LGGLAVASNPSPYYGVV---GLVLASVAGCGWLLSLGVSFVSL 53
Query: 200 VLF 202
VLF
Sbjct: 54 VLF 56
>YYBF_BACSU (P37498) Hypothetical transport protein yybF
Length = 404
Score = 30.4 bits (67), Expect = 9.2
Identities = 52/213 (24%), Positives = 88/213 (40%), Gaps = 34/213 (15%)
Query: 67 LLDCSTTCDADHRRPYHV-PVQISLSVFAT--------LSFISLSK-WDRK--YGFSKFL 114
+L C + R +HV P SLS+ T L F SLS+ W RK G S L
Sbjct: 28 ILYCVQPLMEEFTREFHVTPTAASLSLSVTTMLLAVSMLVFGSLSEVWGRKPIMGIS-ML 86
Query: 115 FLDKVSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQCVYK----------IWWYISGA 164
+ S + + + RT++ + L GLP + + + YISG
Sbjct: 87 AASVLCLASAFSPSFHTLLVLRTIQGVALAGLPSIAMAYLGEEIEPGSLGSAMGLYISGN 146
Query: 165 SQIPYYGEVYVSGIILCTLELCSWFYRTSIFFLVCVLFRLIGYLQIQRLDEFAPVFQRET 224
+ +G + VSG++ E +W ++ ++ +I ++ + F P +
Sbjct: 147 AIGAVFGRI-VSGLLS---EYLNWHMAMGTIGVISLIASVIFFINLPPSRHFTPRKLKLG 202
Query: 225 EVGTILLEHLRIRRNLRVISHRFRAFILSSLLL 257
++G L+ HLR R+ F F++ LLL
Sbjct: 203 KLGMSLIGHLRDRK-------LFSLFLIGFLLL 228
>NPM2_HUMAN (Q86SE8) Nucleoplasmin 2
Length = 214
Score = 30.4 bits (67), Expect = 9.2
Identities = 12/33 (36%), Positives = 19/33 (57%)
Query: 333 GETPTALIASAQAMAADISWGSSEDESGDEEDE 365
G P L + A+D++W E+E G+EE+E
Sbjct: 109 GSGPVFLSGQERYEASDLTWEEEEEEEGEEEEE 141
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,218,724
Number of Sequences: 164201
Number of extensions: 1917028
Number of successful extensions: 5709
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 5707
Number of HSP's gapped (non-prelim): 14
length of query: 431
length of database: 59,974,054
effective HSP length: 113
effective length of query: 318
effective length of database: 41,419,341
effective search space: 13171350438
effective search space used: 13171350438
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC122163.4


