

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.5 + phase: 1 /partial
(95 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
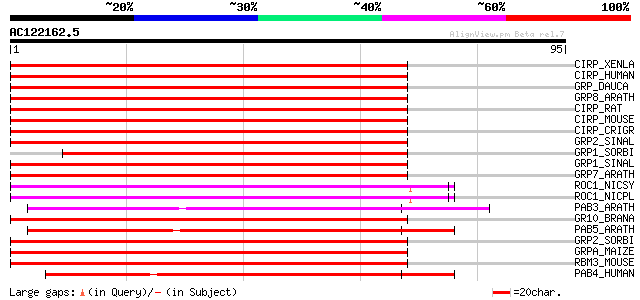
Score E
Sequences producing significant alignments: (bits) Value
CIRP_XENLA (O93235) Cold-inducible RNA-binding protein (Glycine-... 62 3e-10
CIRP_HUMAN (Q14011) Cold-inducible RNA-binding protein (Glycine-... 62 3e-10
GRP_DAUCA (Q03878) Glycine-rich RNA-binding protein 62 3e-10
GRP8_ARATH (Q03251) Glycine-rich RNA-binding protein 8 (CCR1 pro... 62 3e-10
CIRP_RAT (P60825) Cold-inducible RNA-binding protein (Glycine-ri... 62 3e-10
CIRP_MOUSE (P60824) Cold-inducible RNA-binding protein (Glycine-... 62 3e-10
CIRP_CRIGR (P60826) Cold-inducible RNA-binding protein (Glycine-... 62 3e-10
GRP2_SINAL (P49311) Glycine-rich RNA-binding protein GRP2A 59 2e-09
GRP1_SORBI (Q99069) Glycine-rich RNA-binding protein 1 (Fragment) 59 2e-09
GRP1_SINAL (P49310) Glycine-rich RNA-binding protein GRP1A 59 2e-09
GRP7_ARATH (Q03250) Glycine-rich RNA-binding protein 7 59 3e-09
ROC1_NICSY (Q08935) 29 kDa ribonucleoprotein A, chloroplast prec... 58 5e-09
ROC1_NICPL (P49313) 30 kDa ribonucleoprotein, chloroplast precur... 58 5e-09
PAB3_ARATH (O64380) Polyadenylate-binding protein 3 (Poly(A)-bin... 57 6e-09
GR10_BRANA (Q05966) Glycine-rich RNA-binding protein 10 56 1e-08
PAB5_ARATH (Q05196) Polyadenylate-binding protein 5 (Poly(A)-bin... 56 2e-08
GRP2_SORBI (Q99070) Glycine-rich RNA-binding protein 2 56 2e-08
GRPA_MAIZE (P10979) Glycine-rich RNA-binding, abscisic acid-indu... 55 2e-08
RBM3_MOUSE (O89086) Putative RNA-binding protein 3 (RNA binding ... 55 3e-08
PAB4_HUMAN (Q13310) Polyadenylate-binding protein 4 (Poly(A)-bin... 55 3e-08
>CIRP_XENLA (O93235) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (XCIRP)
Length = 163
Score = 62.0 bits (149), Expect = 3e-10
Identities = 28/68 (41%), Positives = 45/68 (66%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L F T E+ L +AF++YG + + +V ++ KR +GFG+VTF ++A+ A + MNGK +
Sbjct: 12 LNFETNEDCLEQAFTKYGRISEVVVVKDRETKRSRGFGFVTFENVDDAKDAMMAMNGKSV 71
Query: 61 HGRVLYVD 68
GR + VD
Sbjct: 72 DGRQIRVD 79
>CIRP_HUMAN (Q14011) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 62.0 bits (149), Expect = 3e-10
Identities = 27/68 (39%), Positives = 45/68 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F T E+ L + FS+YG + + +V ++ +R +GFG+VTF ++A+ A + MNGK +
Sbjct: 13 LSFDTNEQSLEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMAMNGKSV 72
Query: 61 HGRVLYVD 68
GR + VD
Sbjct: 73 DGRQIRVD 80
>GRP_DAUCA (Q03878) Glycine-rich RNA-binding protein
Length = 157
Score = 61.6 bits (148), Expect = 3e-10
Identities = 29/68 (42%), Positives = 47/68 (68%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T +E L +AFSQ+G++ + I+ ++ R +GFG+VTF +E+ R A GMNG+ L
Sbjct: 13 LAWATNDESLEQAFSQFGDITDSKIINDRETGRSRGFGFVTFKDEKSMRDAIEGMNGQEL 72
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 73 DGRNITVN 80
>GRP8_ARATH (Q03251) Glycine-rich RNA-binding protein 8 (CCR1
protein)
Length = 169
Score = 61.6 bits (148), Expect = 3e-10
Identities = 30/68 (44%), Positives = 46/68 (67%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T +E L FSQ+G+V+ + I+ ++ R +GFG+VTF +E+ R A MNGK L
Sbjct: 13 LAWATNDEDLQRTFSQFGDVIDSKIINDRESGRSRGFGFVTFKDEKAMRDAIEEMNGKEL 72
Query: 61 HGRVLYVD 68
GRV+ V+
Sbjct: 73 DGRVITVN 80
>CIRP_RAT (P60825) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 61.6 bits (148), Expect = 3e-10
Identities = 27/68 (39%), Positives = 45/68 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F T E+ L + FS+YG + + +V ++ +R +GFG+VTF ++A+ A + MNGK +
Sbjct: 13 LSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMAMNGKSV 72
Query: 61 HGRVLYVD 68
GR + VD
Sbjct: 73 DGRQIRVD 80
>CIRP_MOUSE (P60824) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 61.6 bits (148), Expect = 3e-10
Identities = 27/68 (39%), Positives = 45/68 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F T E+ L + FS+YG + + +V ++ +R +GFG+VTF ++A+ A + MNGK +
Sbjct: 13 LSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMAMNGKSV 72
Query: 61 HGRVLYVD 68
GR + VD
Sbjct: 73 DGRQIRVD 80
>CIRP_CRIGR (P60826) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 61.6 bits (148), Expect = 3e-10
Identities = 27/68 (39%), Positives = 45/68 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L+F T E+ L + FS+YG + + +V ++ +R +GFG+VTF ++A+ A + MNGK +
Sbjct: 13 LSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMAMNGKSV 72
Query: 61 HGRVLYVD 68
GR + VD
Sbjct: 73 DGRQIRVD 80
>GRP2_SINAL (P49311) Glycine-rich RNA-binding protein GRP2A
Length = 169
Score = 58.9 bits (141), Expect = 2e-09
Identities = 29/68 (42%), Positives = 45/68 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T E L AFSQ+G +V + I+ ++ R +GFG+VTF +E+ + A GMNG+ L
Sbjct: 15 LAWATDERSLETAFSQFGELVDSKIINDRETGRSRGFGFVTFKDEKSMKDAIEGMNGQDL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRSITVN 82
>GRP1_SORBI (Q99069) Glycine-rich RNA-binding protein 1 (Fragment)
Length = 142
Score = 58.9 bits (141), Expect = 2e-09
Identities = 29/59 (49%), Positives = 41/59 (69%)
Query: 10 LAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRVLYVD 68
L AFS YG V+++ I+L++ +R +GFG+VTF+ EE R A GMNGK L GR + V+
Sbjct: 3 LHSAFSTYGEVLESKIILDRETQRSRGFGFVTFSTEEAMRSAIEGMNGKELDGRNITVN 61
>GRP1_SINAL (P49310) Glycine-rich RNA-binding protein GRP1A
Length = 166
Score = 58.9 bits (141), Expect = 2e-09
Identities = 29/68 (42%), Positives = 45/68 (65%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T + L AFSQYG V+ + I+ ++ R +GFG+VTF +E+ + A GMNG+ L
Sbjct: 15 LAWATDDRALETAFSQYGEVLDSKIINDRETGRSRGFGFVTFKDEKSMKDAIEGMNGQDL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRSITVN 82
>GRP7_ARATH (Q03250) Glycine-rich RNA-binding protein 7
Length = 176
Score = 58.5 bits (140), Expect = 3e-09
Identities = 28/68 (41%), Positives = 46/68 (67%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T + L AF+QYG+V+ + I+ ++ R +GFG+VTF +E+ + A GMNG+ L
Sbjct: 15 LAWATDDRALETAFAQYGDVIDSKIINDRETGRSRGFGFVTFKDEKAMKDAIEGMNGQDL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRSITVN 82
>ROC1_NICSY (Q08935) 29 kDa ribonucleoprotein A, chloroplast
precursor (CP29A)
Length = 273
Score = 57.8 bits (138), Expect = 5e-09
Identities = 30/76 (39%), Positives = 42/76 (54%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L FS LAE F + GNV +++ +K R +GFG+VT + +EE A NG L
Sbjct: 94 LPFSADSAALAELFERAGNVEMVEVIYDKLTGRSRGFGFVTMSSKEEVEAACQQFNGYEL 153
Query: 61 HGRVLYVDMDPPNEQK 76
GR L V+ PP E++
Sbjct: 154 DGRALRVNSGPPPEKR 169
Score = 50.4 bits (119), Expect = 8e-07
Identities = 27/78 (34%), Positives = 43/78 (54%), Gaps = 3/78 (3%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA+ ++ L FS+ G VV A +V ++ R +GFG+VT++ EE A ++G L
Sbjct: 195 LAWGVDQDALETLFSEQGKVVDAKVVYDRDSGRSRGFGFVTYSSAEEVNNAIESLDGVDL 254
Query: 61 HGRVLYV---DMDPPNEQ 75
+GR + V + PP Q
Sbjct: 255 NGRAIRVSPAEARPPRRQ 272
>ROC1_NICPL (P49313) 30 kDa ribonucleoprotein, chloroplast precursor
(CP-RBP30)
Length = 279
Score = 57.8 bits (138), Expect = 5e-09
Identities = 30/76 (39%), Positives = 42/76 (54%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L FS LAE F + GNV +++ +K R +GFG+VT + +EE A NG L
Sbjct: 94 LLFSADSAALAELFERAGNVEMVEVIYDKLTGRSRGFGFVTMSSKEEVEAACQQFNGYEL 153
Query: 61 HGRVLYVDMDPPNEQK 76
GR L V+ PP E++
Sbjct: 154 DGRALRVNSGPPPEKR 169
Score = 50.4 bits (119), Expect = 8e-07
Identities = 27/78 (34%), Positives = 43/78 (54%), Gaps = 3/78 (3%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA+ ++ L FS+ G VV A +V ++ R +GFG+VT++ EE A ++G L
Sbjct: 201 LAWGVDQDALETLFSEQGKVVDAKVVYDRDSGRSRGFGFVTYSSAEEVNNAIESLDGVDL 260
Query: 61 HGRVLYV---DMDPPNEQ 75
+GR + V + PP Q
Sbjct: 261 NGRAIRVSPAEARPPRRQ 278
>PAB3_ARATH (O64380) Polyadenylate-binding protein 3
(Poly(A)-binding protein 3) (PABP 3)
Length = 660
Score = 57.4 bits (137), Expect = 6e-09
Identities = 30/79 (37%), Positives = 47/79 (58%), Gaps = 1/79 (1%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S +EKL E FS+YGNV + ++LN + +GFG+V ++ EEA +A MNGK++ +
Sbjct: 342 SVDDEKLKEMFSEYGNVTSSKVMLNP-QGMSRGFGFVAYSNPEEALRALSEMNGKMIGRK 400
Query: 64 VLYVDMDPPNEQKNKIKQA 82
LY+ + E + QA
Sbjct: 401 PLYIALAQRKEDRRAHLQA 419
Score = 40.4 bits (93), Expect = 8e-04
Identities = 19/64 (29%), Positives = 37/64 (57%), Gaps = 1/64 (1%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S + L E FS +G ++ + ++ R KG+G+V F +EE A+ A +NG +++ +
Sbjct: 146 SIDNKALFETFSSFGTILSCKVAMD-VTGRSKGYGFVQFEKEESAQAAIDKLNGMLMNDK 204
Query: 64 VLYV 67
++V
Sbjct: 205 QVFV 208
Score = 34.7 bits (78), Expect = 0.043
Identities = 21/61 (34%), Positives = 32/61 (52%), Gaps = 1/61 (1%)
Query: 7 EEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRVLY 66
E++L + F ++G V+ + +V+ + FG+V F E A A MNG L VLY
Sbjct: 242 EDELRKTFGKFG-VISSAVVMRDQSGNSRCFGFVNFECTEAAASAVEKMNGISLGDDVLY 300
Query: 67 V 67
V
Sbjct: 301 V 301
>GR10_BRANA (Q05966) Glycine-rich RNA-binding protein 10
Length = 169
Score = 56.2 bits (134), Expect = 1e-08
Identities = 27/68 (39%), Positives = 44/68 (64%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T + +L FSQ+G V+ + I+ ++ R +GFG+VTF +E+ + A MNGK L
Sbjct: 13 LAWATGDAELERTFSQFGEVIDSKIINDRETGRSRGFGFVTFKDEKSMKDAIDEMNGKEL 72
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 73 DGRTITVN 80
>PAB5_ARATH (Q05196) Polyadenylate-binding protein 5
(Poly(A)-binding protein 5) (PABP 5)
Length = 668
Score = 55.8 bits (133), Expect = 2e-08
Identities = 28/73 (38%), Positives = 45/73 (61%), Gaps = 1/73 (1%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S +EKL E FS+YGNV +++N ++ +GFG+V ++ EEA A MNGK++ +
Sbjct: 338 SVNDEKLKEMFSEYGNVTSCKVMMN-SQGLSRGFGFVAYSNPEEALLAMKEMNGKMIGRK 396
Query: 64 VLYVDMDPPNEQK 76
LYV + E++
Sbjct: 397 PLYVALAQRKEER 409
Score = 40.4 bits (93), Expect = 8e-04
Identities = 20/64 (31%), Positives = 37/64 (57%), Gaps = 1/64 (1%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S + L E FS +G ++ + ++ R KG+G+V F +EE A+ A +NG +L+ +
Sbjct: 142 SIDNKALYETFSSFGTILSCKVAMDVVG-RSKGYGFVQFEKEETAQAAIDKLNGMLLNDK 200
Query: 64 VLYV 67
++V
Sbjct: 201 QVFV 204
Score = 39.3 bits (90), Expect = 0.002
Identities = 26/81 (32%), Positives = 39/81 (48%), Gaps = 6/81 (7%)
Query: 6 TEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRVL 65
T+++L + F +YG++ A +V+ + FG+V F E A A MNG L VL
Sbjct: 237 TDDELKKTFGKYGDISSA-VVMKDQSGNSRSFGFVNFVSPEAAAVAVEKMNGISLGEDVL 295
Query: 66 YV-----DMDPPNEQKNKIKQ 81
YV D E + K +Q
Sbjct: 296 YVGRAQKKSDREEELRRKFEQ 316
Score = 30.4 bits (67), Expect = 0.81
Identities = 25/96 (26%), Positives = 40/96 (41%), Gaps = 6/96 (6%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHG- 62
S E L + F+Q V V R G+ YV FA E+A +A +N +
Sbjct: 55 SVNESHLLDLFNQVAPVHNLR-VCRDLTHRSLGYAYVNFANPEDASRAMESLNYAPIRDR 113
Query: 63 --RVLYVDMDPPNEQKNKIKQATKNTE--VDNEDVH 94
R++ + DP K KN + +DN+ ++
Sbjct: 114 PIRIMLSNRDPSTRLSGKGNVFIKNLDASIDNKALY 149
>GRP2_SORBI (Q99070) Glycine-rich RNA-binding protein 2
Length = 168
Score = 55.8 bits (133), Expect = 2e-08
Identities = 26/68 (38%), Positives = 43/68 (63%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T E L +AF+ +G V+ + ++ ++ R +GFG+VTF+ E+ A MNGK L
Sbjct: 15 LAWATNNETLEQAFANFGQVIDSKVITDRETGRSRGFGFVTFSSEQSMLDAIENMNGKEL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRNITVN 82
>GRPA_MAIZE (P10979) Glycine-rich RNA-binding, abscisic
acid-inducible protein
Length = 157
Score = 55.5 bits (132), Expect = 2e-08
Identities = 26/68 (38%), Positives = 42/68 (61%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
LA++T+ E L AF+ YG ++ + ++ ++ R +GFG+VTF+ E A MNGK L
Sbjct: 15 LAWATSNESLENAFASYGEILDSKVITDRETGRSRGFGFVTFSSENSMLDAIENMNGKEL 74
Query: 61 HGRVLYVD 68
GR + V+
Sbjct: 75 DGRNITVN 82
>RBM3_MOUSE (O89086) Putative RNA-binding protein 3 (RNA binding
motif protein 3)
Length = 153
Score = 55.1 bits (131), Expect = 3e-08
Identities = 26/68 (38%), Positives = 41/68 (60%)
Query: 1 LAFSTTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKIL 60
L F+T E+ L + FS +G + + +V ++ +R +GFG++TF E A A MNG+ L
Sbjct: 13 LNFNTDEQALEDHFSSFGPISEVVVVKDRETQRSRGFGFITFTNPEHASDAMRAMNGESL 72
Query: 61 HGRVLYVD 68
GR + VD
Sbjct: 73 DGRQIRVD 80
>PAB4_HUMAN (Q13310) Polyadenylate-binding protein 4
(Poly(A)-binding protein 4) (PABP 4) (Inducible
poly(A)-binding protein) (iPABP) (Activated-platelet
protein-1) (APP-1)
Length = 644
Score = 55.1 bits (131), Expect = 3e-08
Identities = 28/70 (40%), Positives = 43/70 (61%), Gaps = 2/70 (2%)
Query: 7 EEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRVLY 66
+EKL + FS +G++ A ++L R KGFG+V F+ EEA KA MNG+I+ + LY
Sbjct: 307 DEKLRKEFSPFGSITSAKVMLEDG--RSKGFGFVCFSSPEEATKAVTEMNGRIVGSKPLY 364
Query: 67 VDMDPPNEQK 76
V + E++
Sbjct: 365 VALAQRKEER 374
Score = 49.3 bits (116), Expect = 2e-06
Identities = 23/61 (37%), Positives = 38/61 (61%), Gaps = 1/61 (1%)
Query: 7 EEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGRVLY 66
+E L E FSQ+G + V+ + KGFG+V++ + E+A KA MNGK + G++++
Sbjct: 204 DESLKELFSQFGKTLSVK-VMRDPNGKSKGFGFVSYEKHEDANKAVEEMNGKEISGKIIF 262
Query: 67 V 67
V
Sbjct: 263 V 263
Score = 39.3 bits (90), Expect = 0.002
Identities = 21/64 (32%), Positives = 36/64 (55%), Gaps = 2/64 (3%)
Query: 4 STTEEKLAEAFSQYGNVVKADIVLNKAKKRCKGFGYVTFAEEEEARKAQIGMNGKILHGR 63
S + L + FS +GN++ +V ++ KG+ +V F +E A KA MNG +L+ R
Sbjct: 109 SIDNKALYDTFSAFGNILSCKVVCDE--NGSKGYAFVHFETQEAADKAIEKMNGMLLNDR 166
Query: 64 VLYV 67
++V
Sbjct: 167 KVFV 170
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.130 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,930,728
Number of Sequences: 164201
Number of extensions: 400568
Number of successful extensions: 1222
Number of sequences better than 10.0: 288
Number of HSP's better than 10.0 without gapping: 244
Number of HSP's successfully gapped in prelim test: 44
Number of HSP's that attempted gapping in prelim test: 764
Number of HSP's gapped (non-prelim): 467
length of query: 95
length of database: 59,974,054
effective HSP length: 71
effective length of query: 24
effective length of database: 48,315,783
effective search space: 1159578792
effective search space used: 1159578792
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC122162.5


